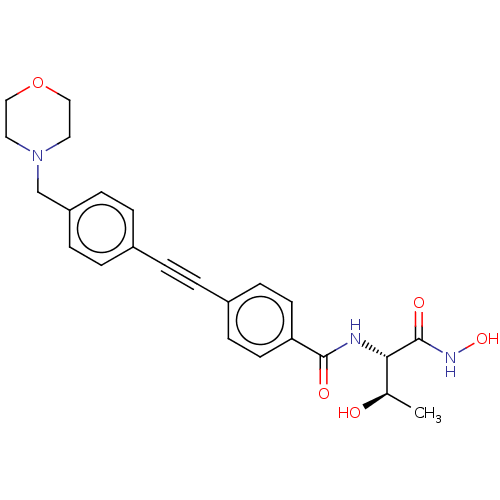
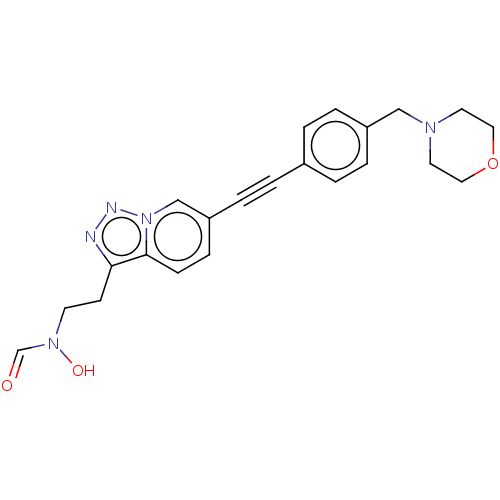
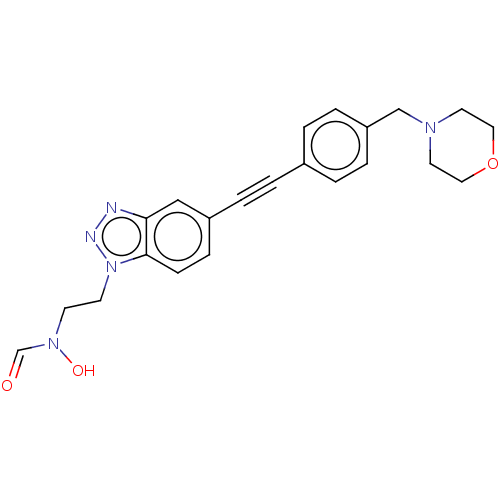
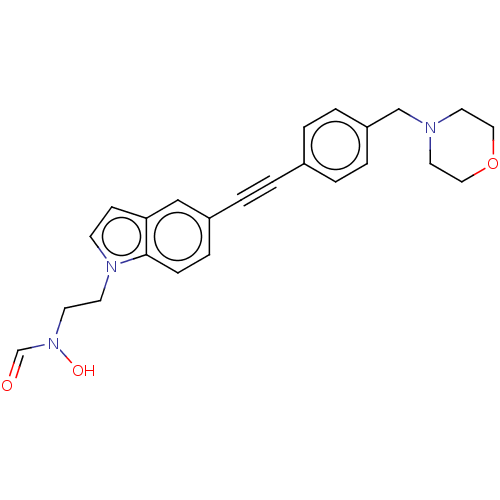
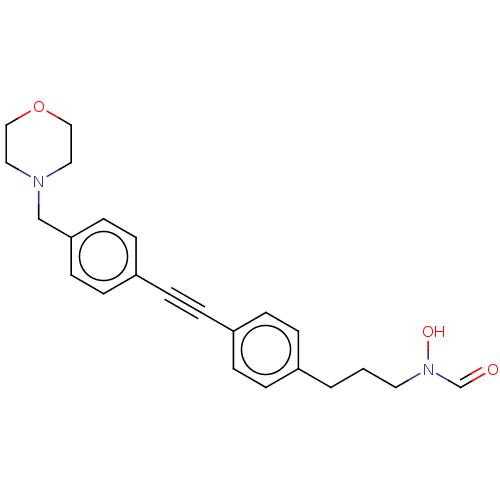
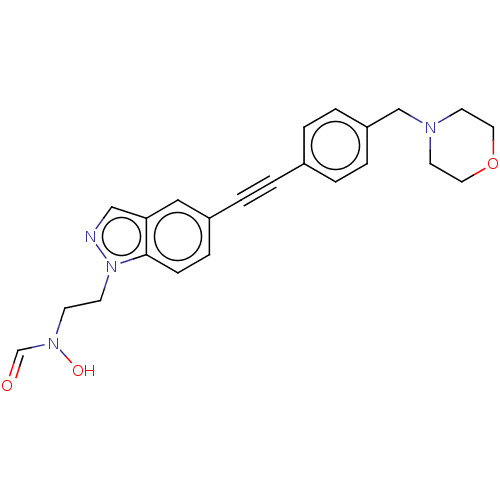
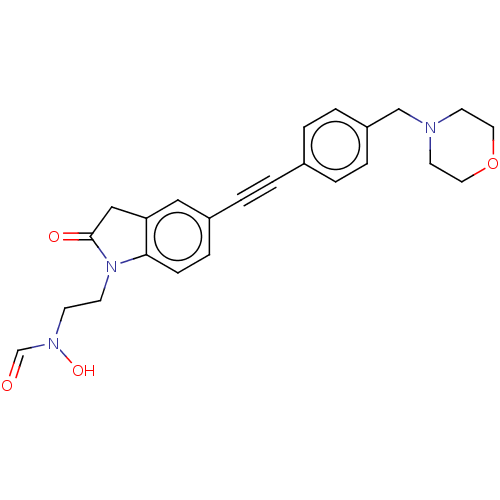
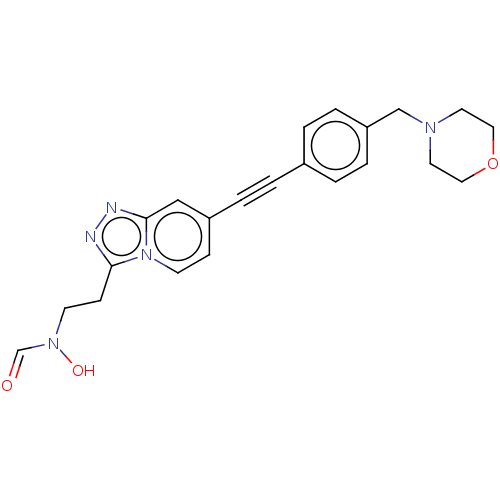
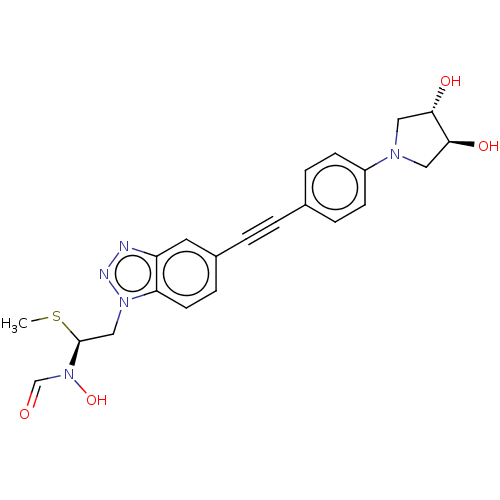
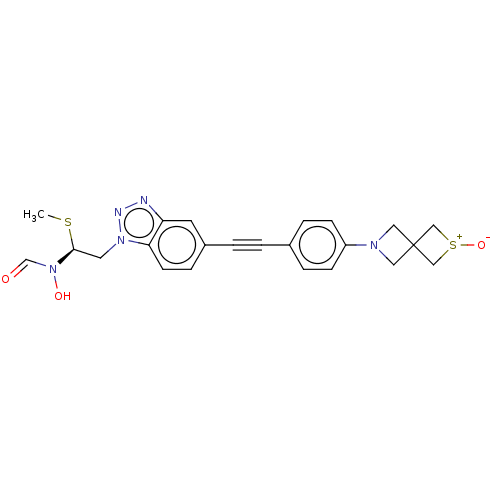
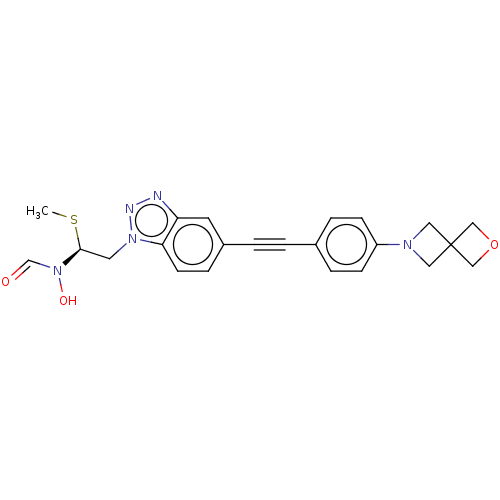
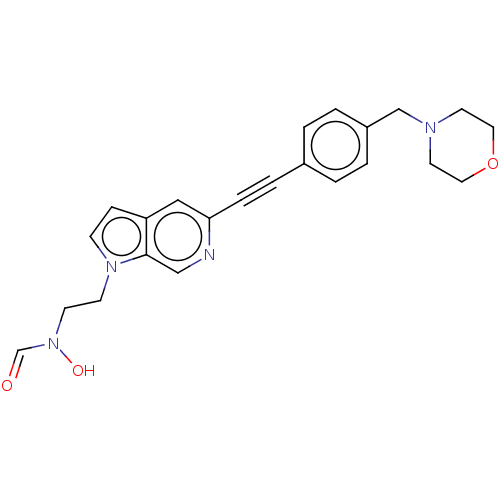
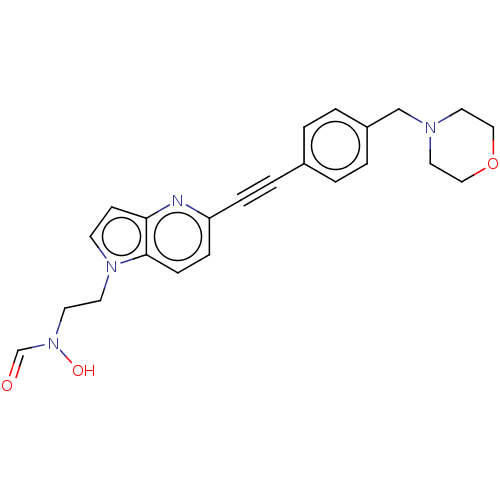
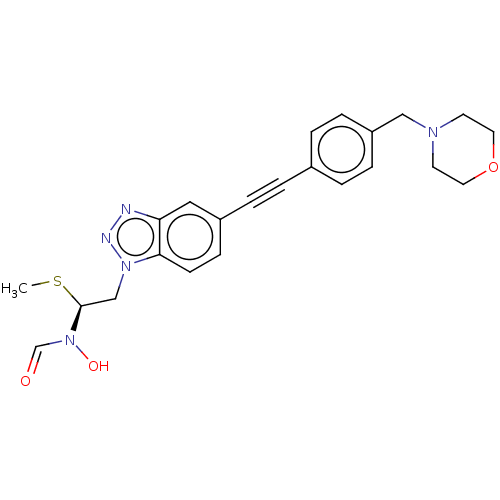
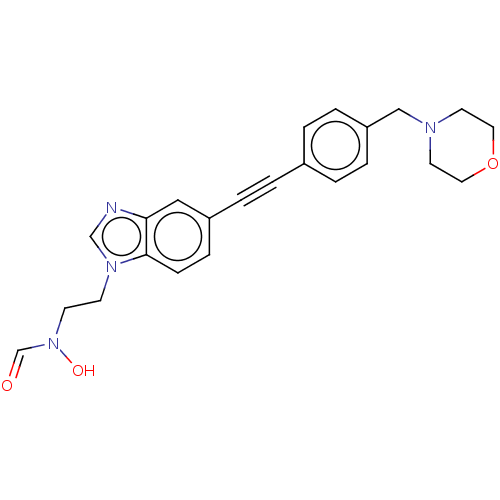
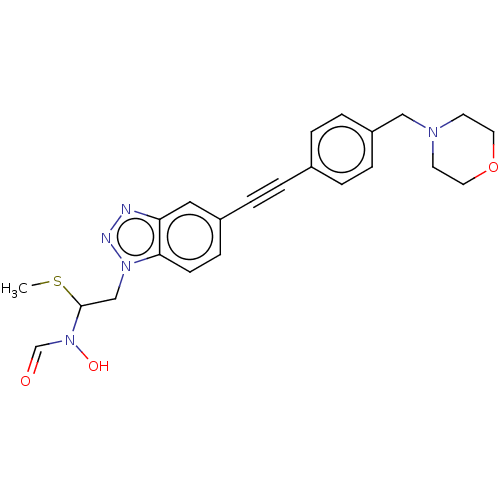
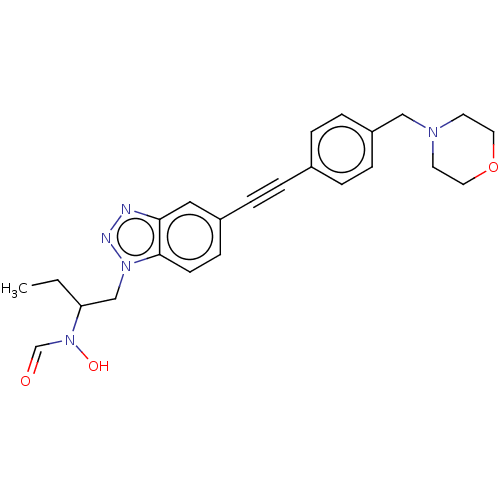
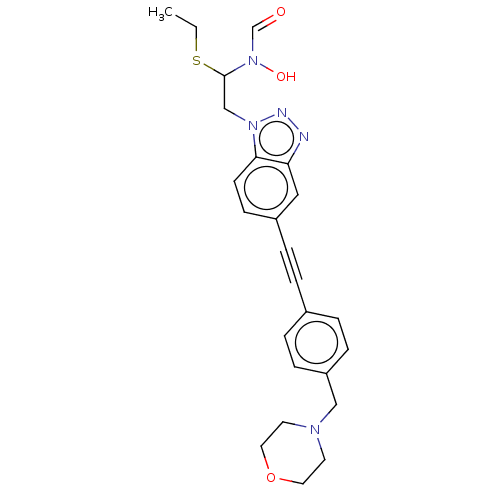
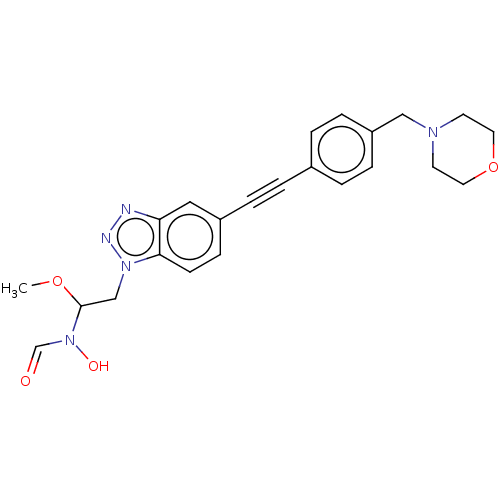
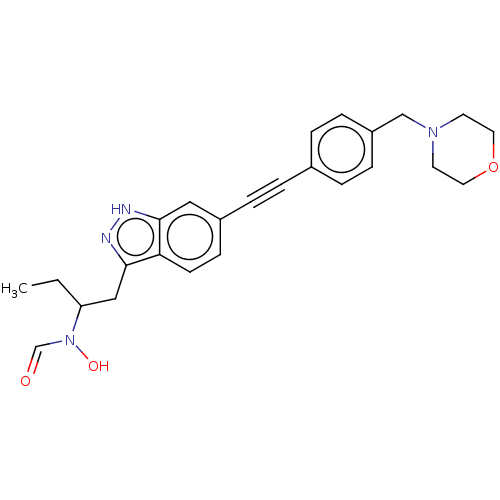
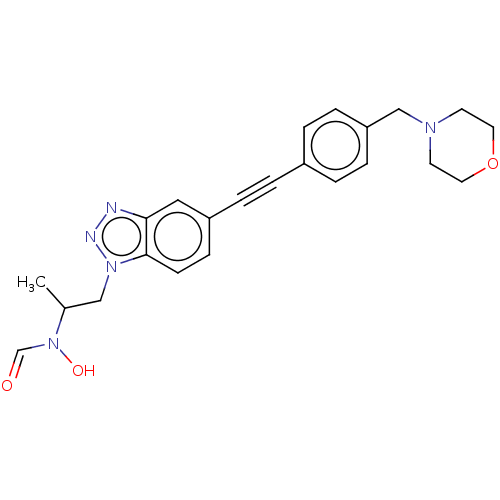
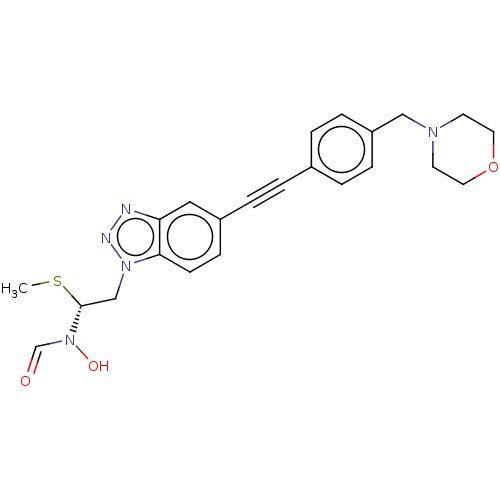
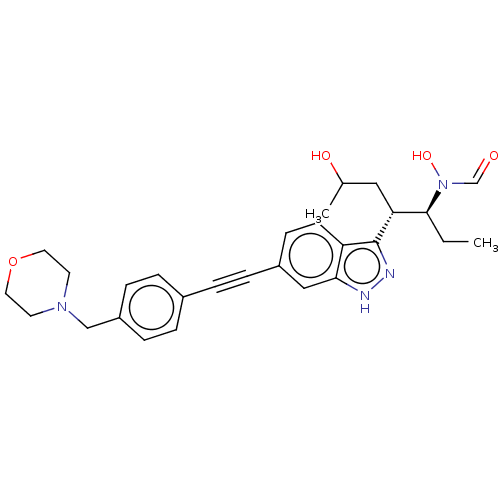
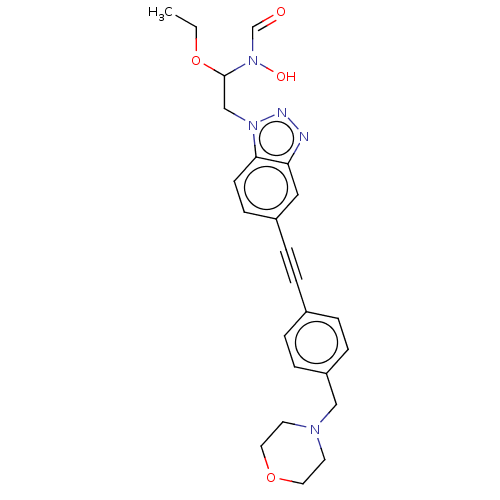
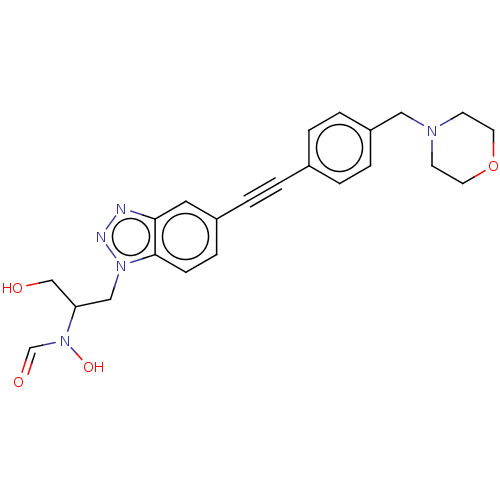
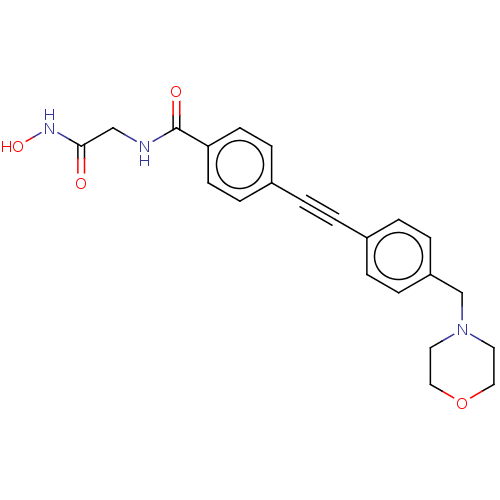
Report error Found 32 Enz. Inhib. hit(s) with all data for entry = 50012156
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 37nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 45nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 63nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 64nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 2.40E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 5.40E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 7.30E+3nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 2.10E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 2.50E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 4.10E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 6.10E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Pseudomonas aeruginosa (strain ATCC 15692 / PAO1 /...)
Entasis Therapeutics
Curated by ChEMBL
Entasis Therapeutics
Curated by ChEMBL
Affinity DataIC50: 6.30E+4nMAssay Description:Displacement of fluorescent ligand from Pseudomonas aeruginosa LpxC measured after 30 mins by fluorescence anisotrophy assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of human AchE expressed in HEK293 cells using acetylthiocholine as substrate by Ellman's methodMore data for this Ligand-Target Pair