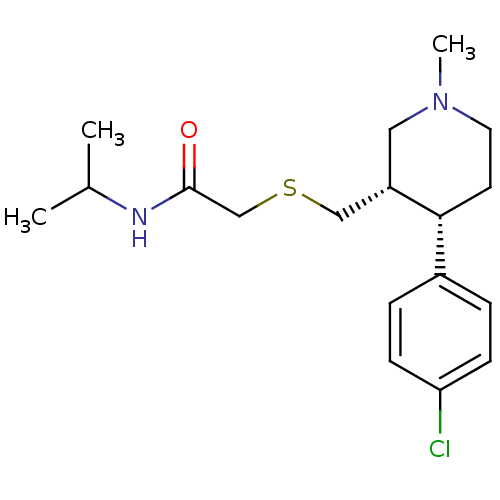
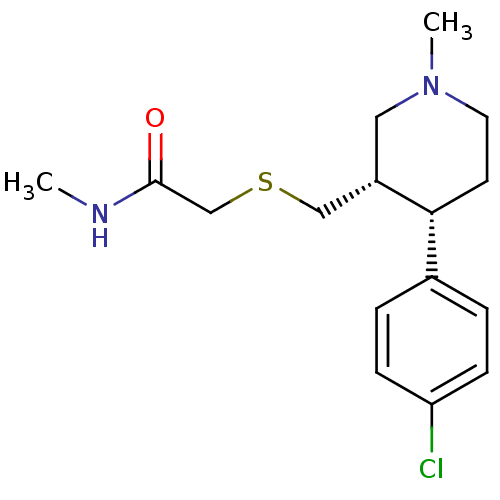
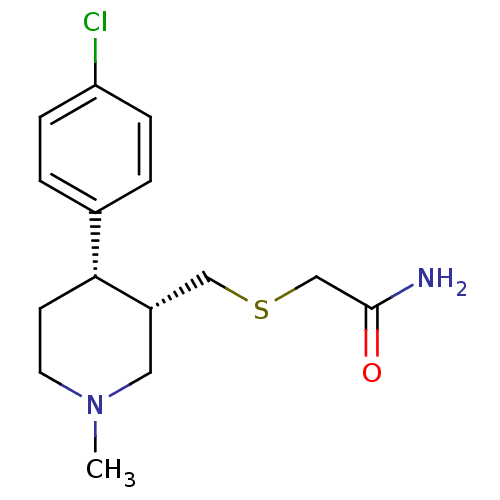
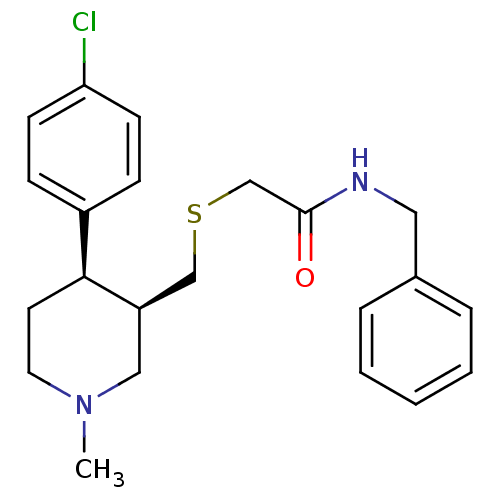
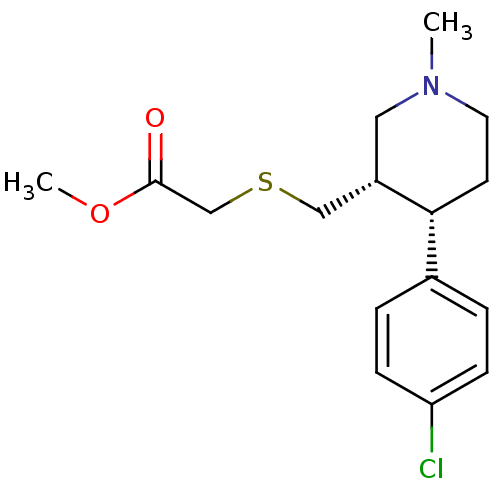
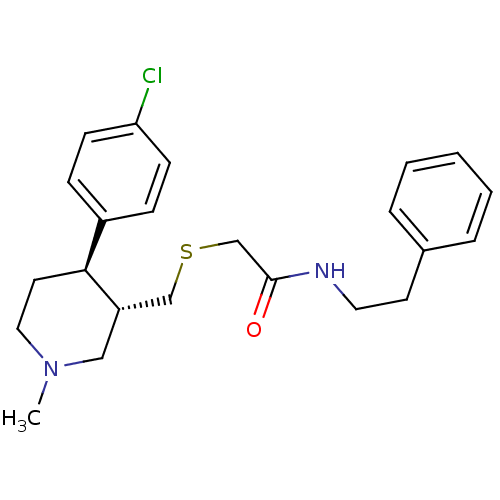
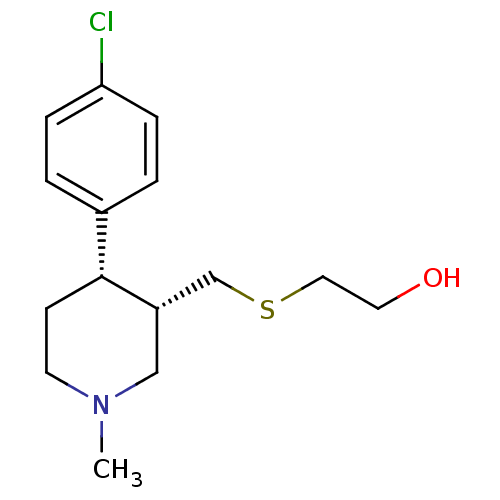
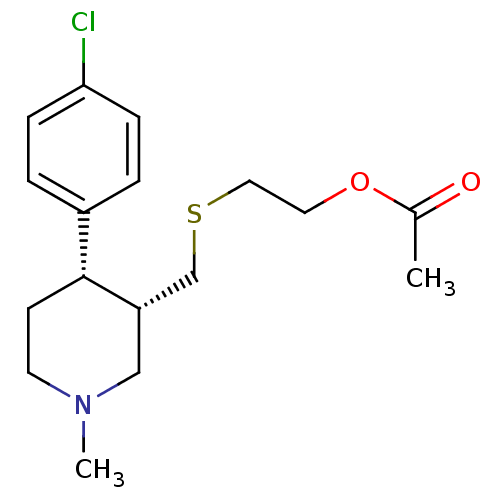
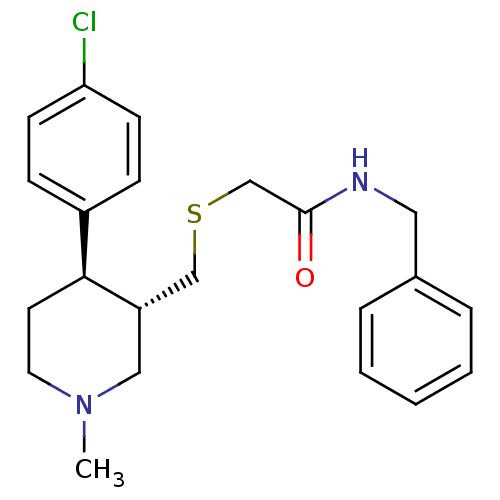
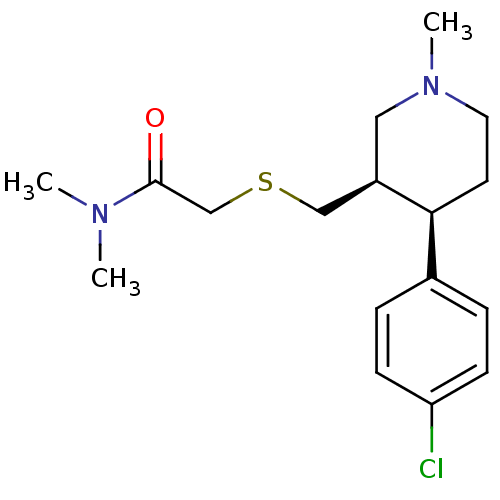
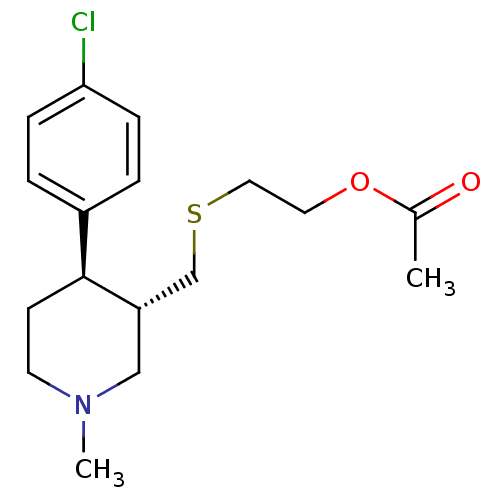
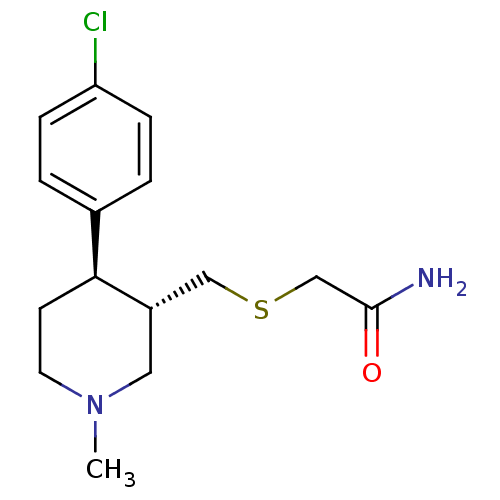
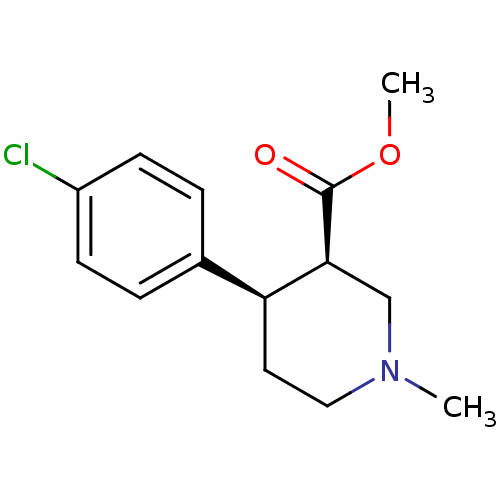
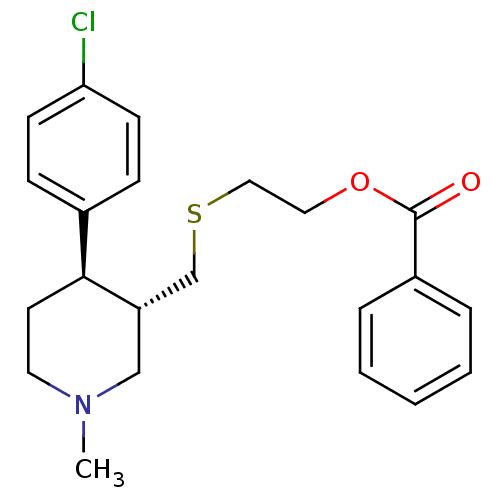
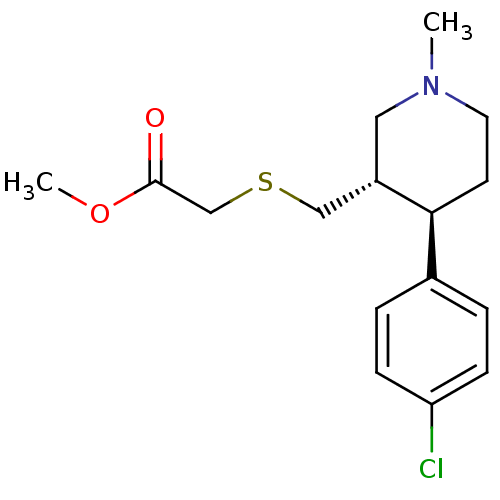
Report error Found 99 Enz. Inhib. hit(s) with all data for entry = 50017180
Affinity DataKi: 3.90nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 5.5nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 8.80nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 22nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 22nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 36nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 44nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 46nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 47nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 52nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 53nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 54nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 54nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 57nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 63nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 67nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 68nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 69nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 69nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 71nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 82nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 85nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 91nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 98nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 101nMAssay Description:Ability to inhibit [3H]NE uptake through NET in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 102nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 108nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 112nMAssay Description:Ability to inhibit [3H]DA uptake through DAT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 116nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair
Affinity DataKi: 121nMAssay Description:Ability to inhibit [3H]5-HT uptake through SERT in rat synaptosomesMore data for this Ligand-Target Pair