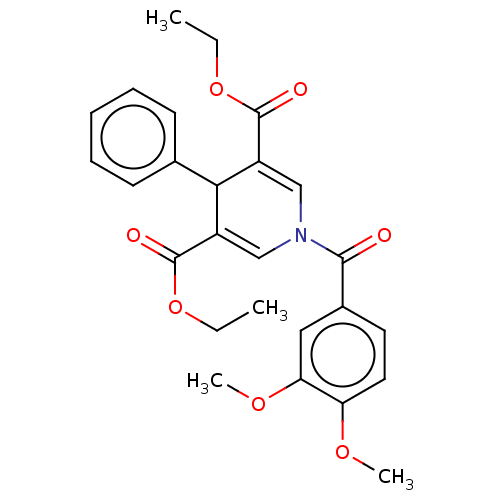
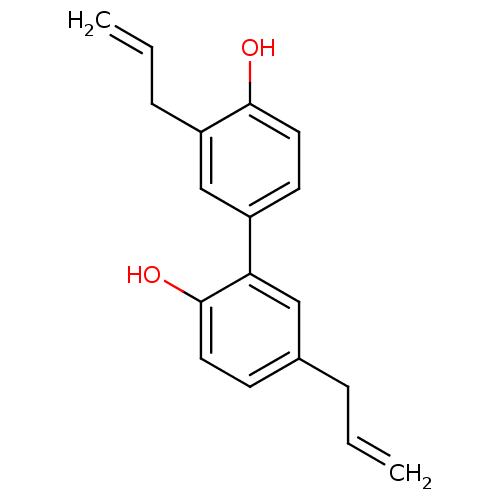
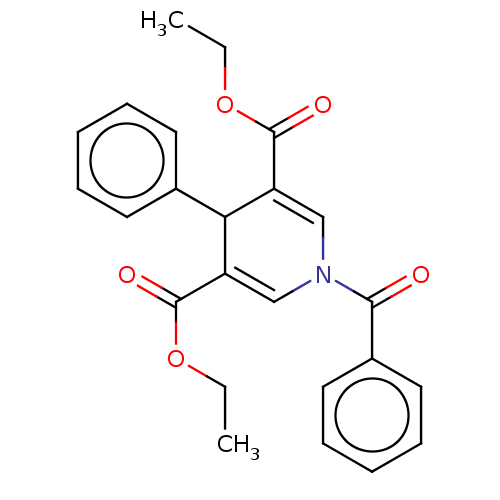
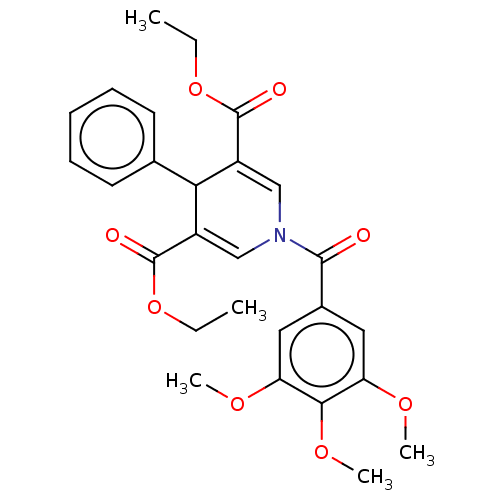
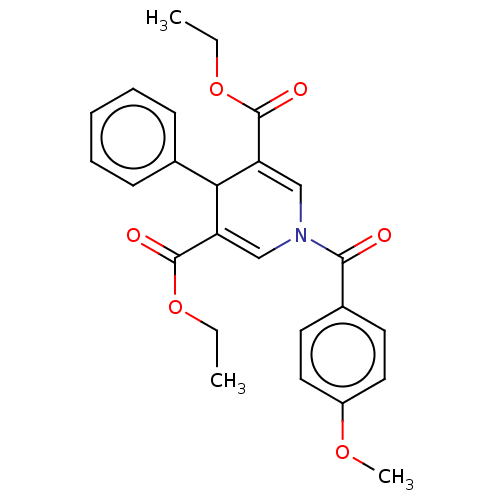
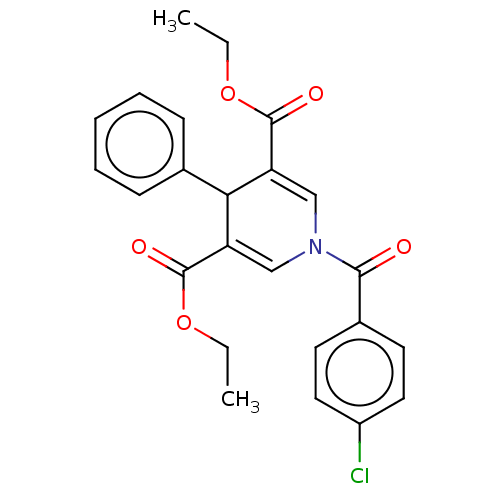
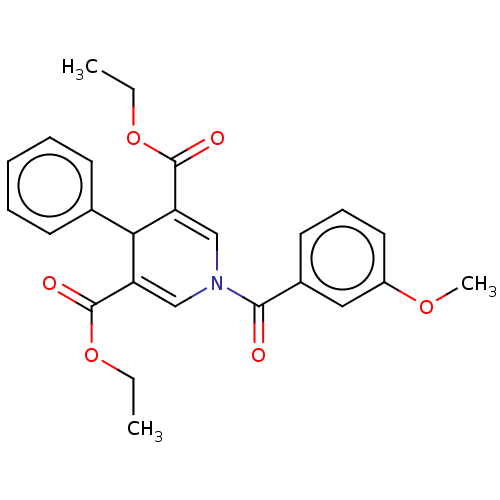
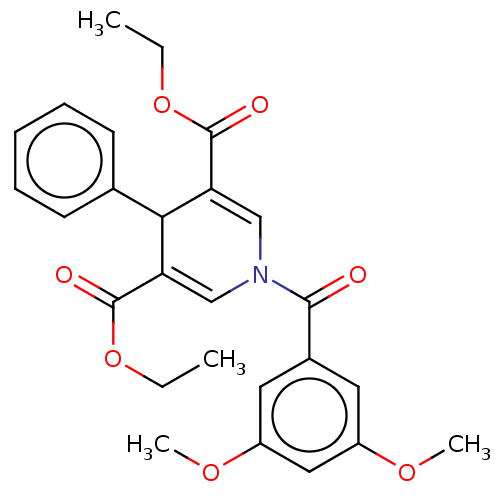
Report error Found 8 Enz. Inhib. hit(s) with all data for entry = 50019569
TargetNAD-dependent protein deacetylase sirtuin-3, mitochondrial(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataKd: 2.90E+4nMAssay Description:Binding affinity to N-terminal his-tagged human SIRT3 (114 to 380 residues) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase sirtuin-3, mitochondrial(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataKd: 4.00E+4nMAssay Description:Binding affinity to N-terminal his-tagged human SIRT3 (114 to 380 residues) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase sirtuin-3, mitochondrial(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataKd: 4.10E+4nMAssay Description:Binding affinity to N-terminal his-tagged human SIRT3 (114 to 380 residues) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase sirtuin-3, mitochondrial(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataKd: 4.70E+4nMAssay Description:Binding affinity to N-terminal his-tagged human SIRT3 (114 to 380 residues) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase sirtuin-3, mitochondrial(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataKd: 7.20E+4nMAssay Description:Binding affinity to N-terminal his-tagged human SIRT3 (114 to 380 residues) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase sirtuin-3, mitochondrial(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataKd: 7.40E+4nMAssay Description:Binding affinity to N-terminal his-tagged human SIRT3 (114 to 380 residues) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase sirtuin-3, mitochondrial(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataKd: 7.80E+4nMAssay Description:Binding affinity to N-terminal his-tagged human SIRT3 (114 to 380 residues) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacetylase sirtuin-3, mitochondrial(Human)
Sapienza University of Rome
Curated by ChEMBL
Sapienza University of Rome
Curated by ChEMBL
Affinity DataKd: 7.90E+4nMAssay Description:Binding affinity to N-terminal his-tagged human SIRT3 (114 to 380 residues) assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair