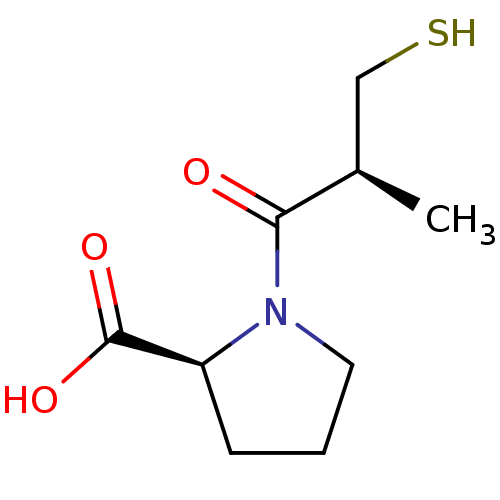
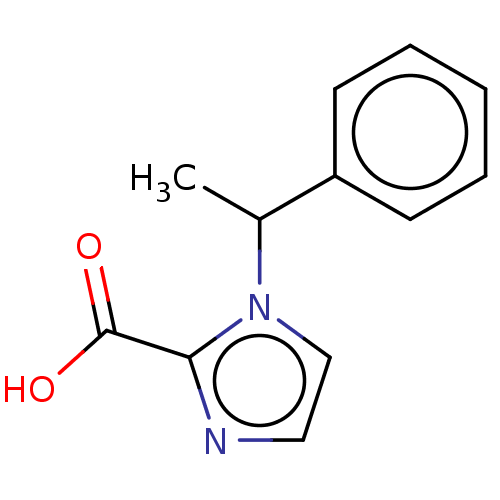
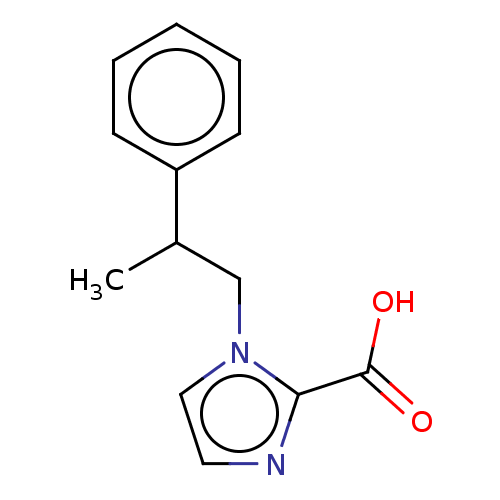
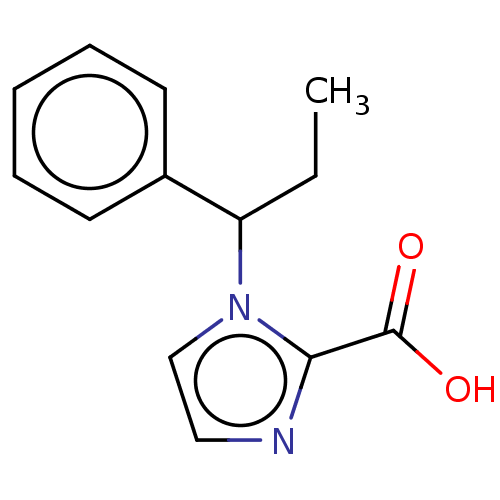
Report error Found 50 Enz. Inhib. hit(s) with all data for entry = 50020115
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
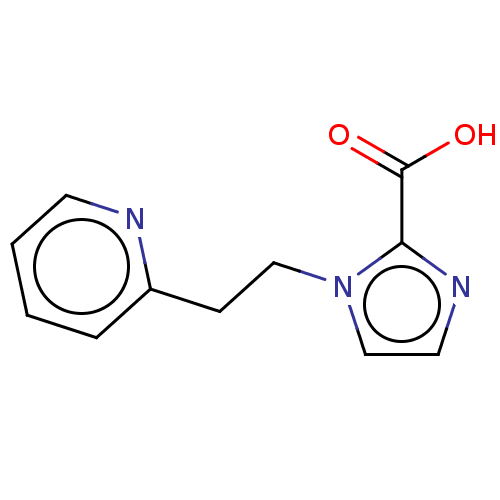
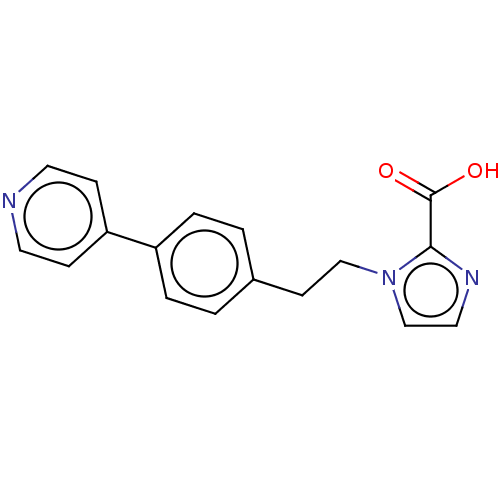
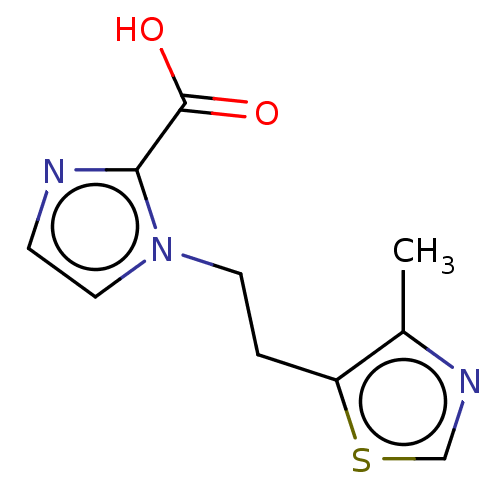
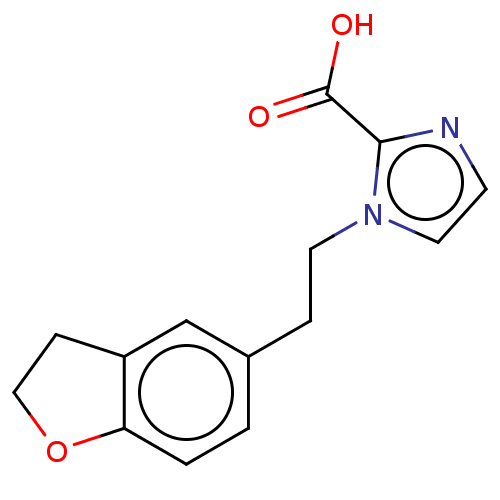
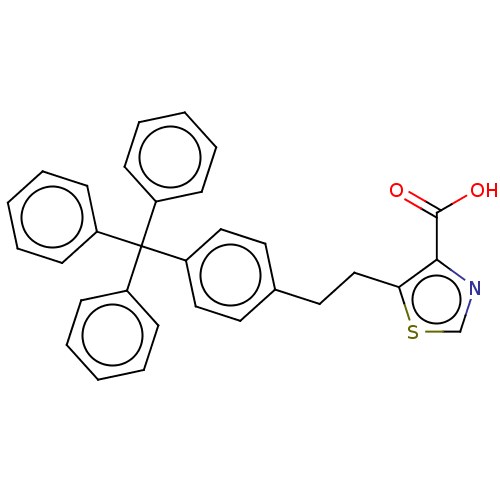
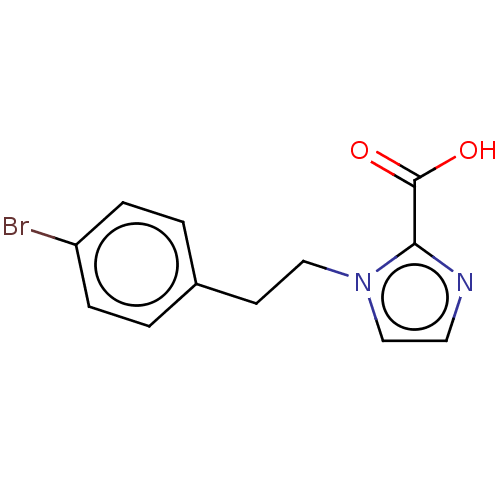
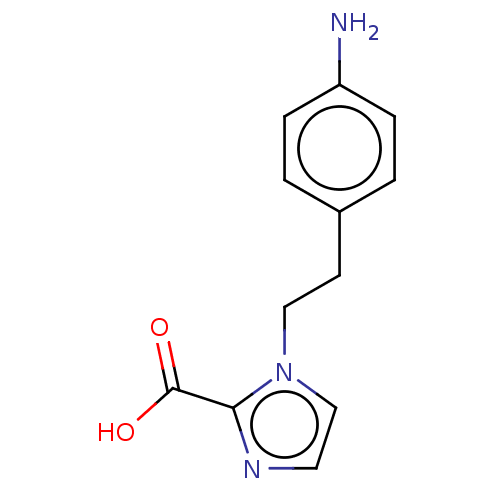
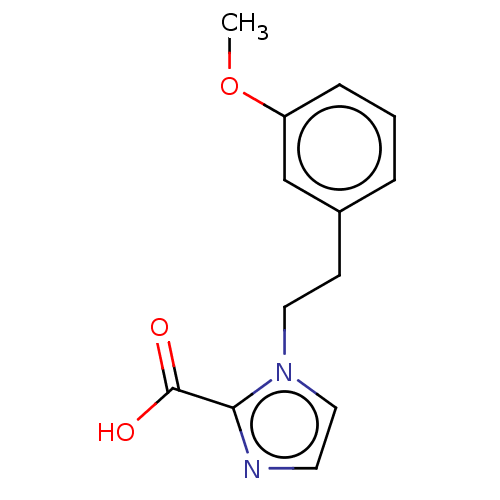
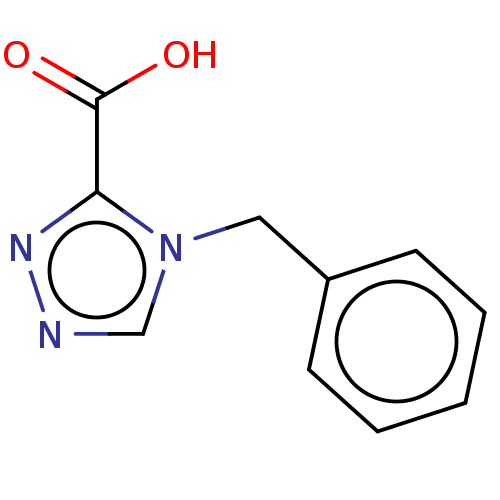
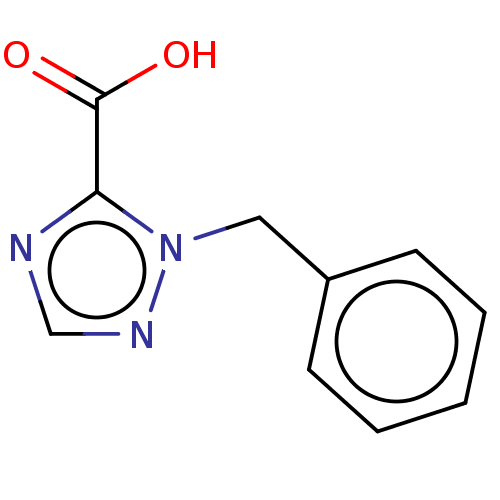
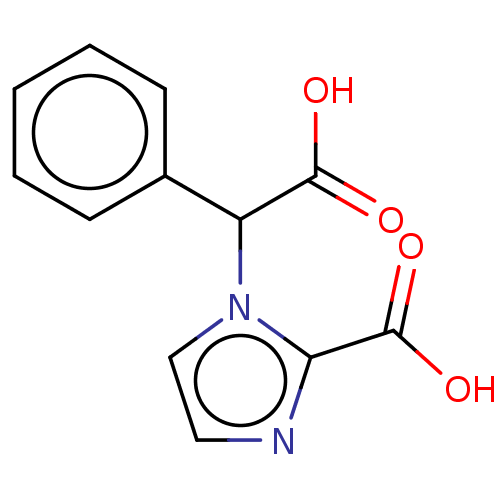
Affinity DataIC50: 1.89E+4nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
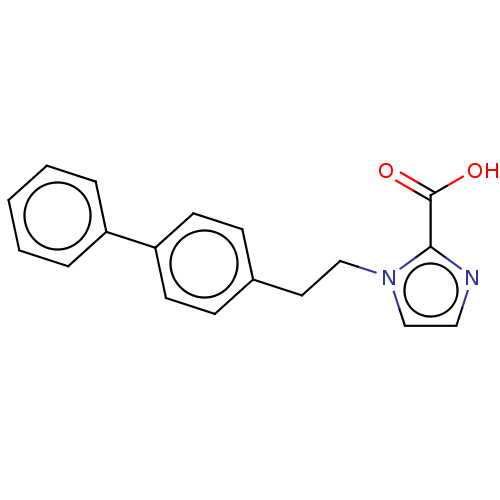
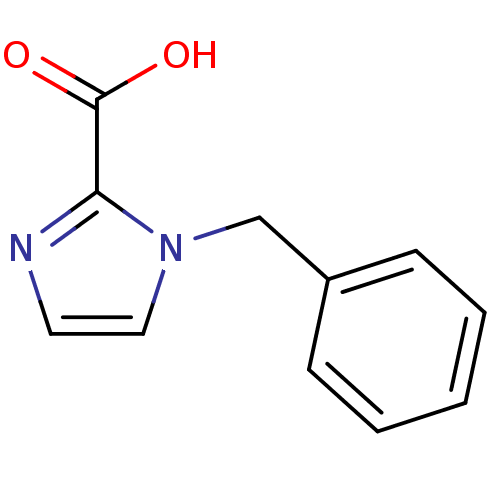
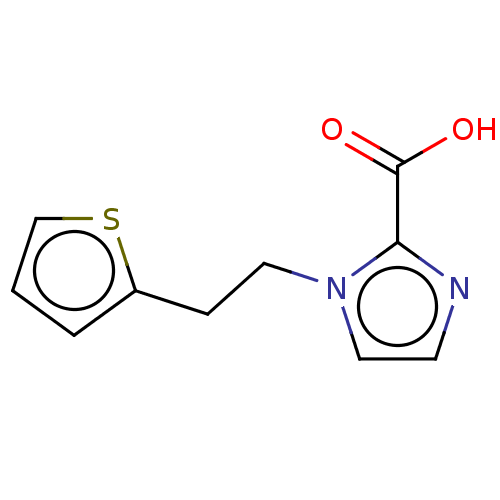
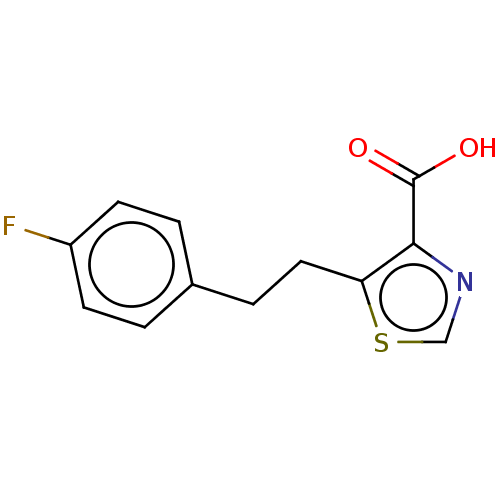
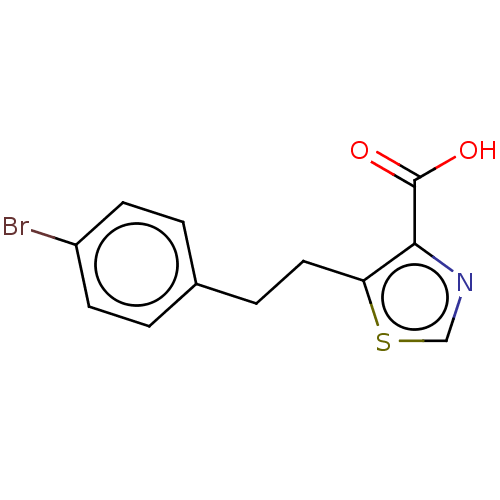
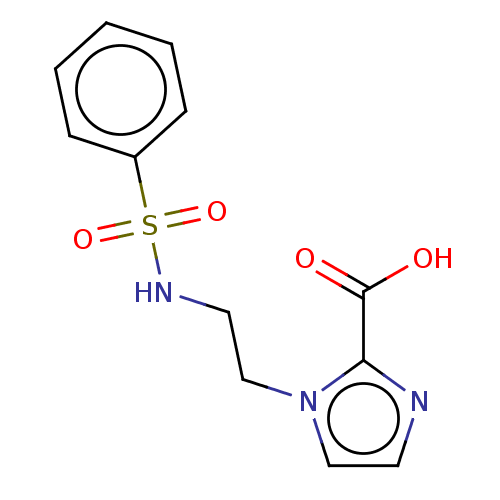
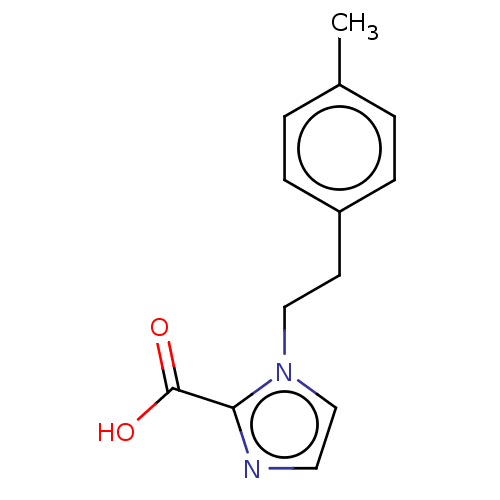
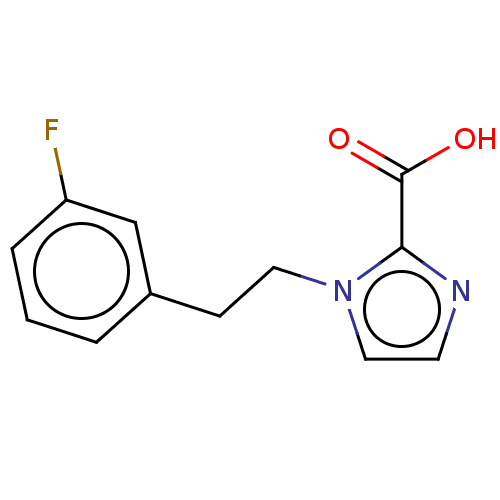
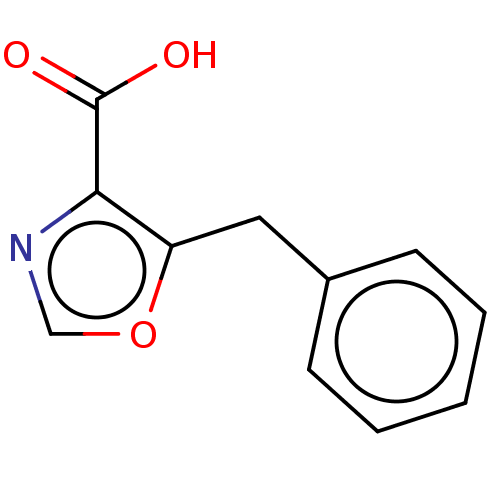
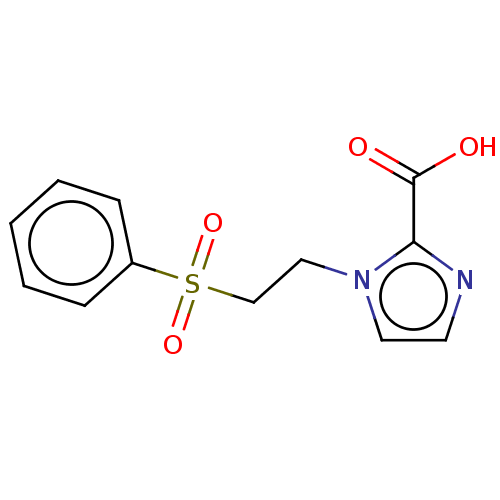
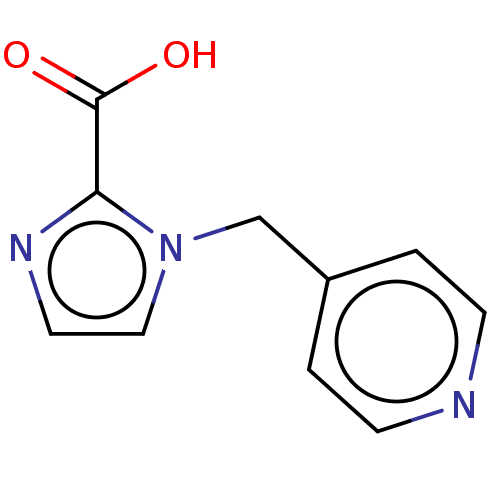
Affinity DataIC50: 2.54E+4nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
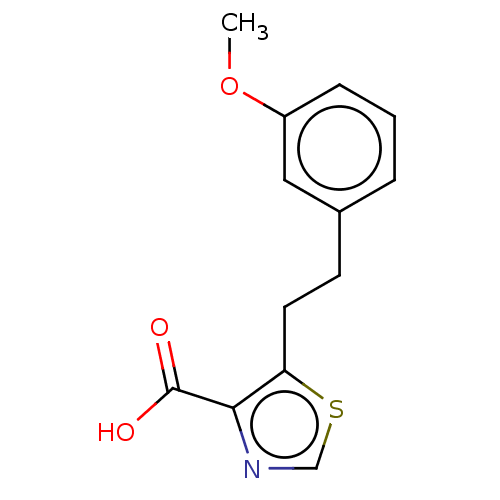
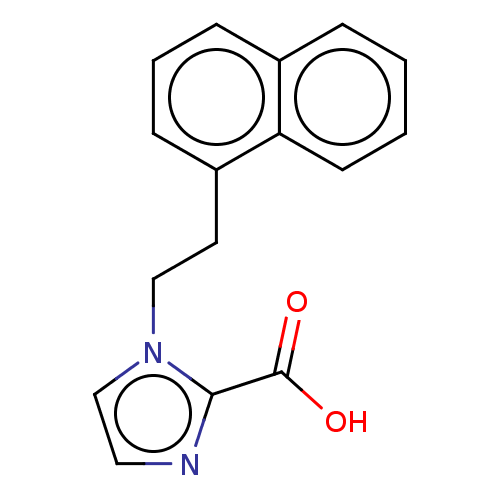
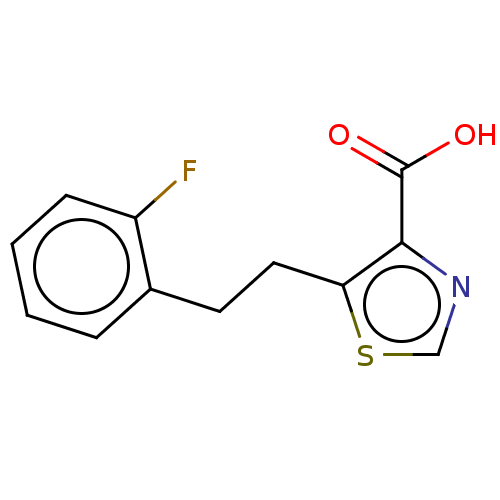
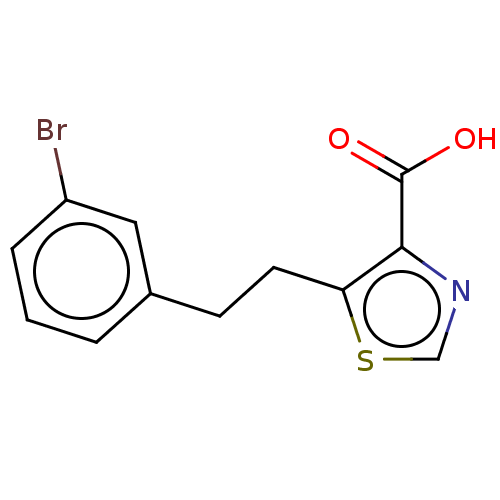
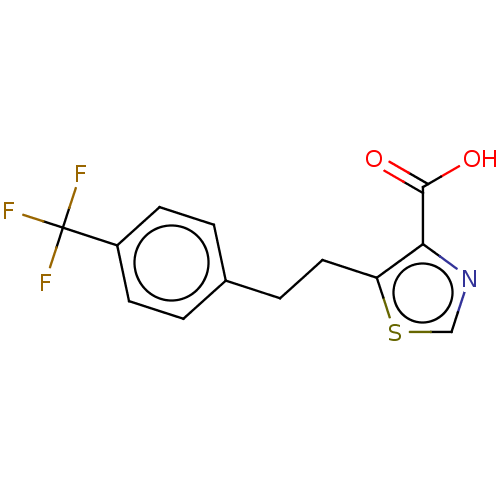
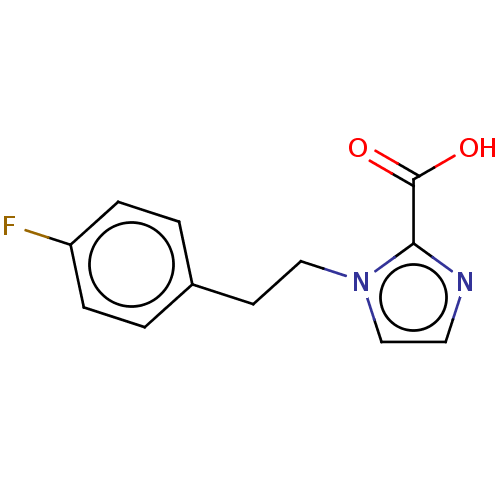
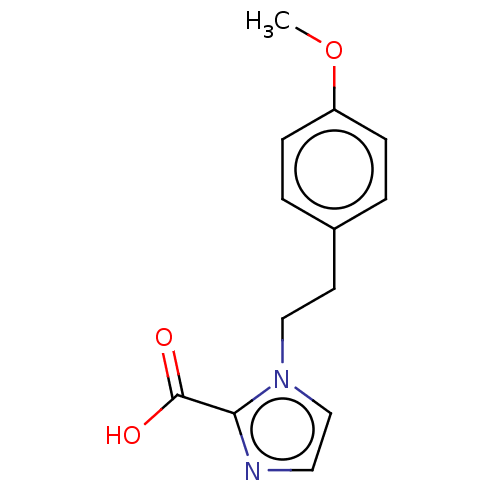
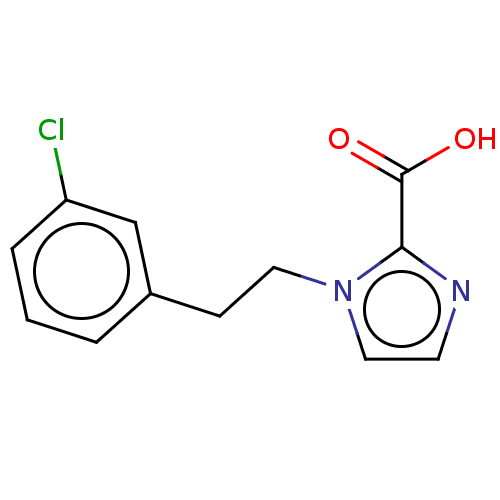
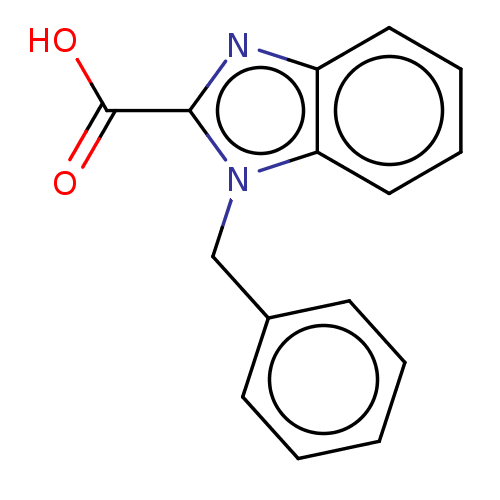
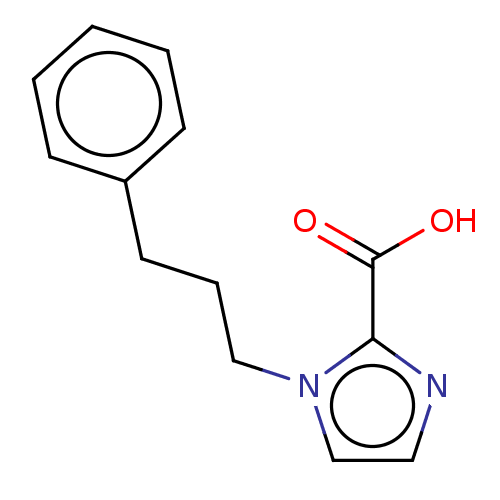
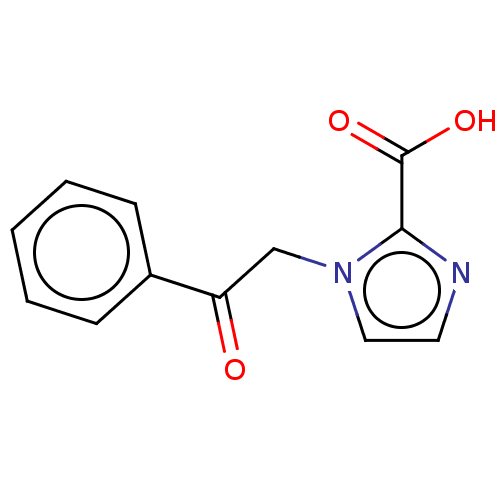
Affinity DataIC50: 2.99E+4nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
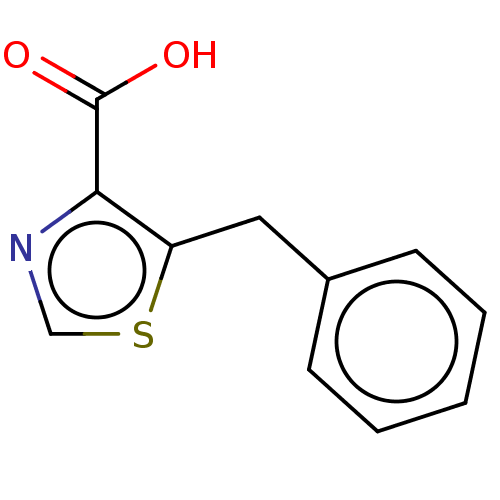
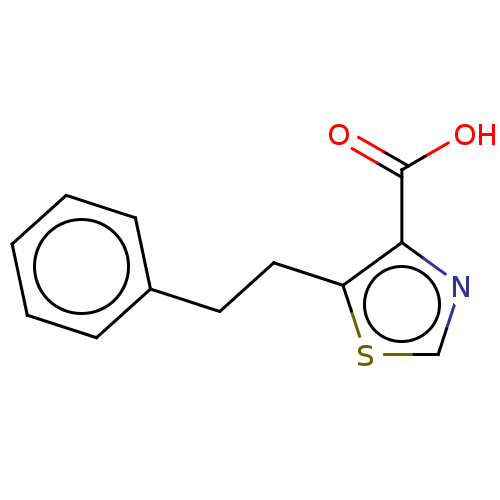
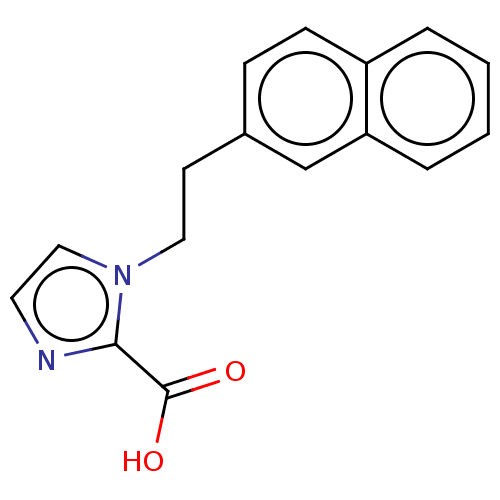
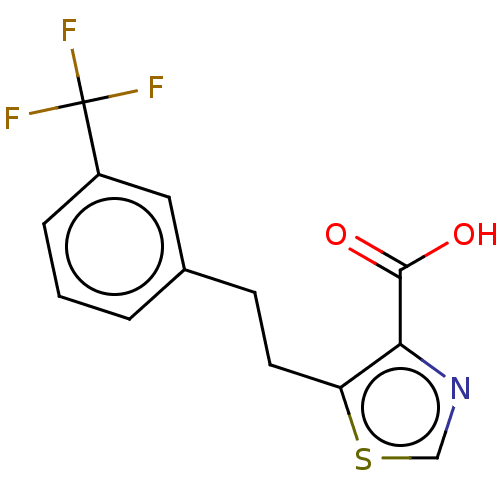
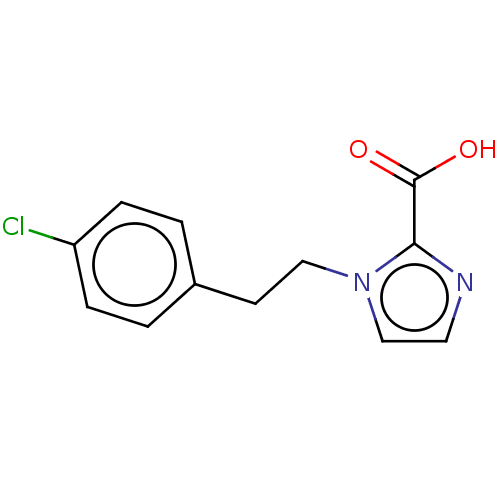
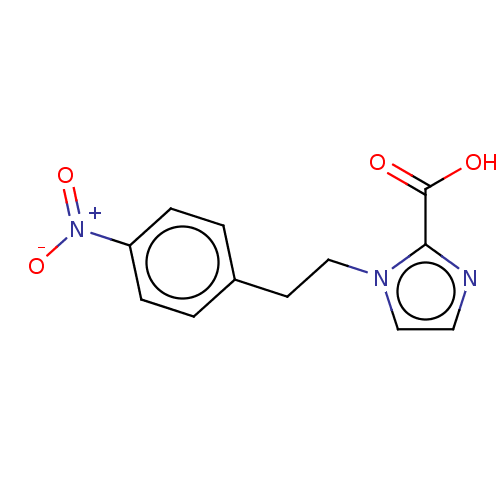
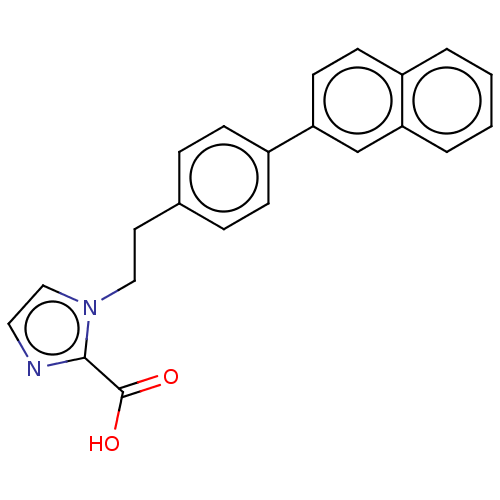
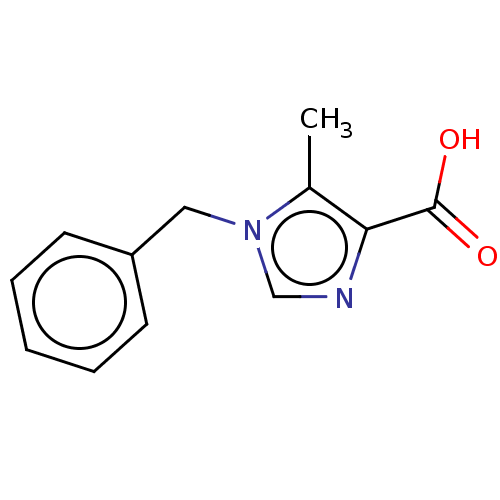
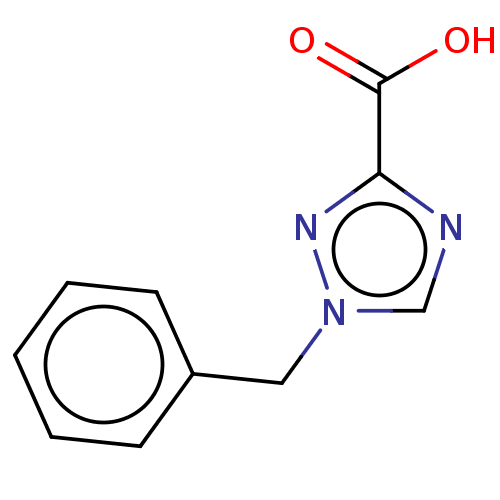
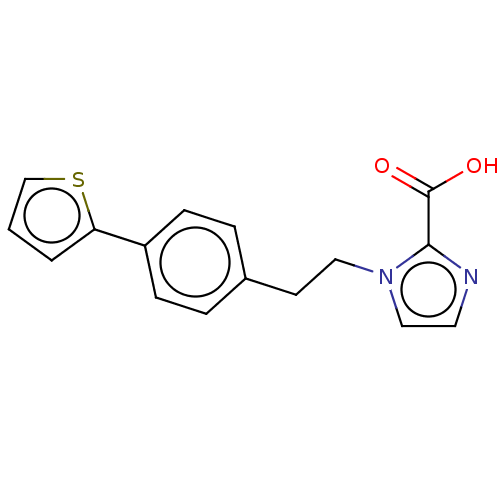
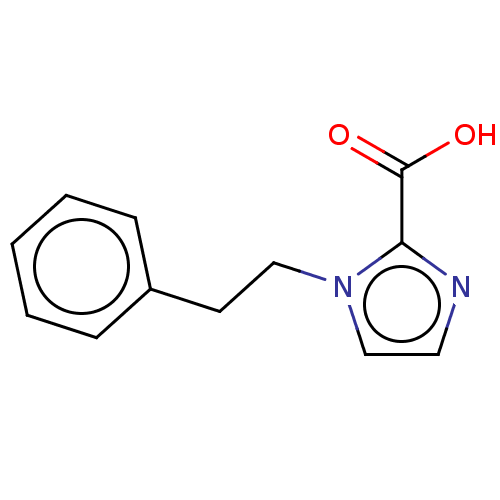
Affinity DataIC50: 4.37E+4nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 5.46E+4nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 6.35E+4nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 7.48E+4nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 8.24E+4nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.26E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 1.97E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 2.03E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair
TargetMetallo-beta-lactamase type 2(Pseudomonas aeruginosa (g-Proteobacteria))
Xihua University
Curated by ChEMBL
Xihua University
Curated by ChEMBL
Affinity DataIC50: 3.01E+5nMAssay Description:Inhibition of NDM-1 (1 to 270 residues) (unknown origin) expressed in Escherichia coli Transetta (DE3) using FC5 as fluorescence substrate preincubat...More data for this Ligand-Target Pair