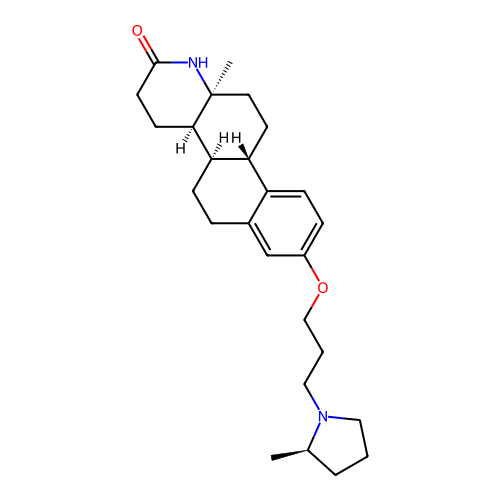
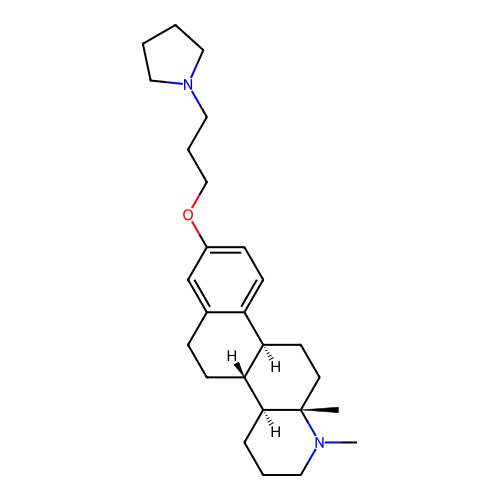
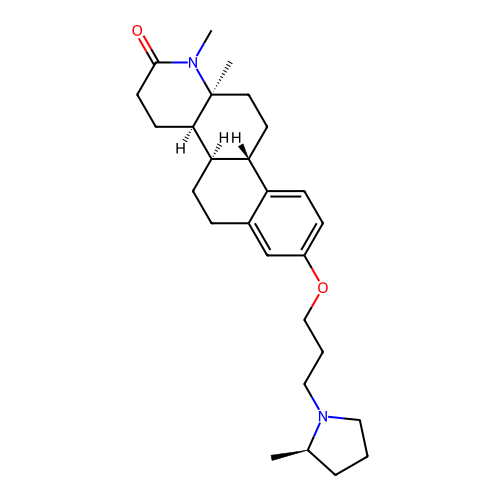
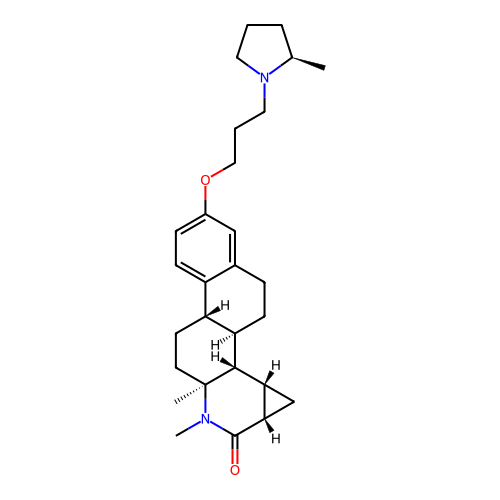
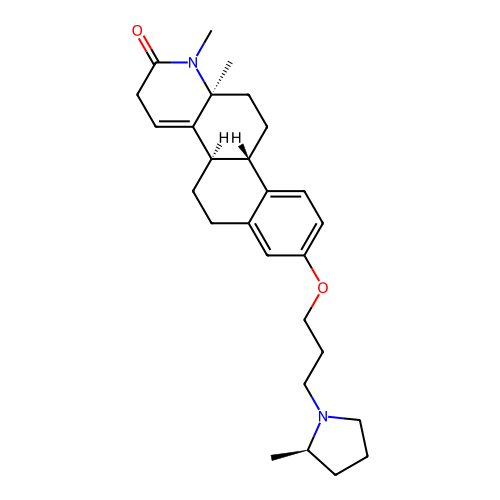
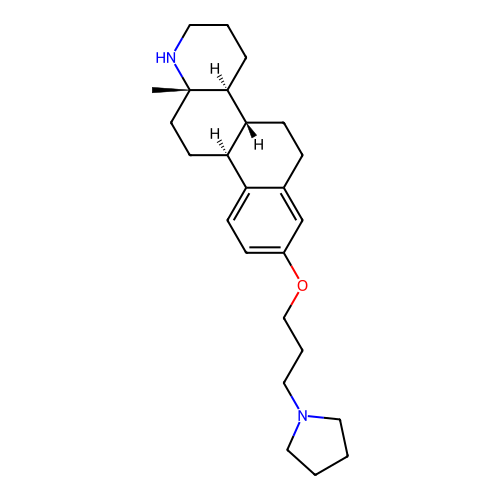
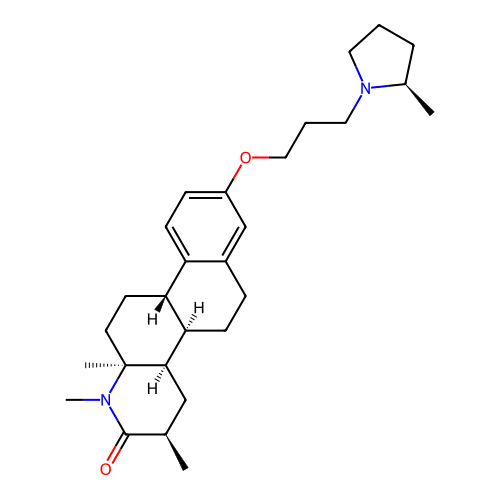
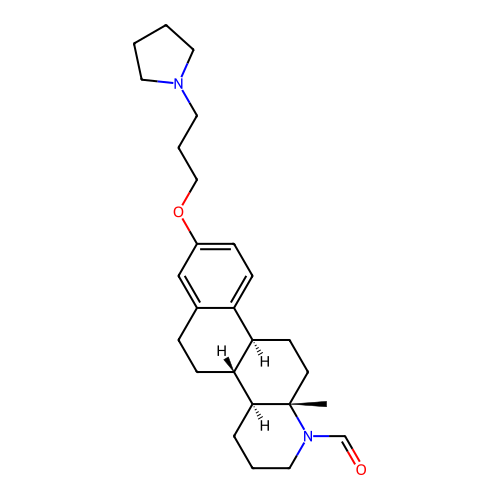
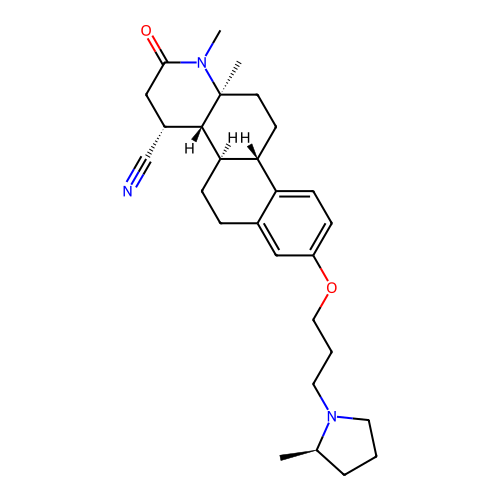
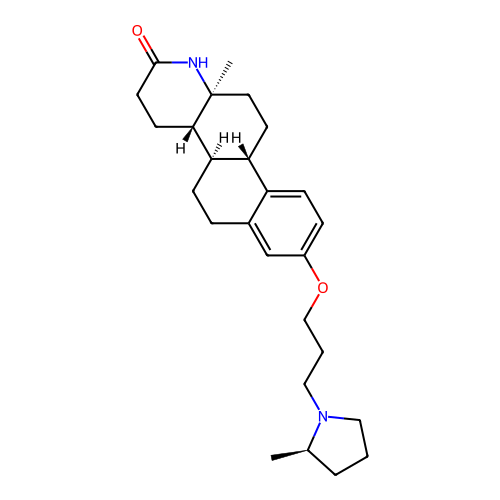
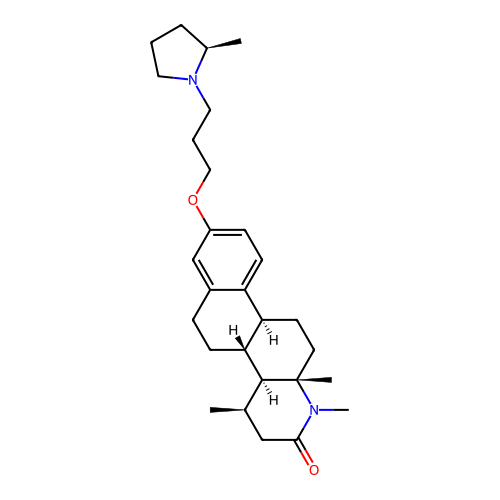
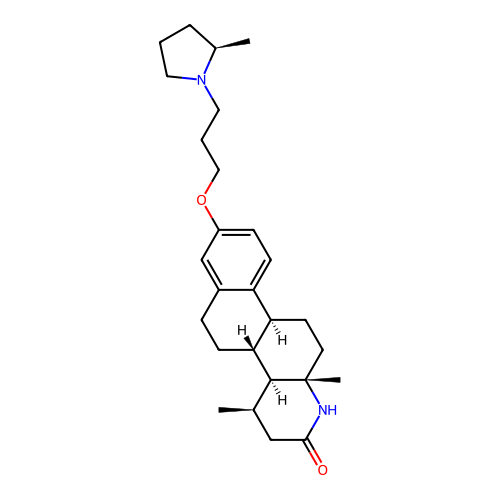
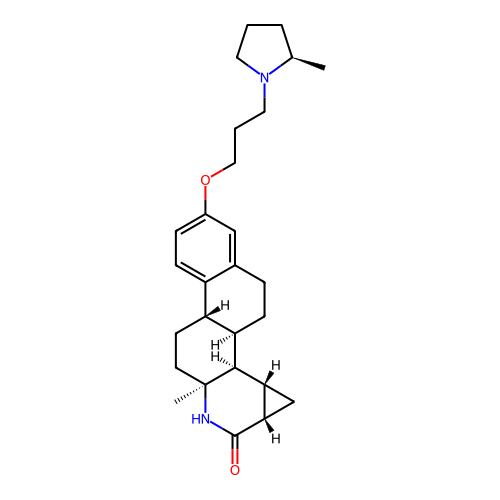
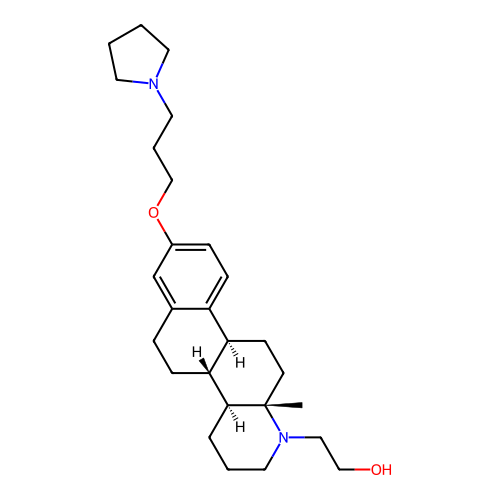
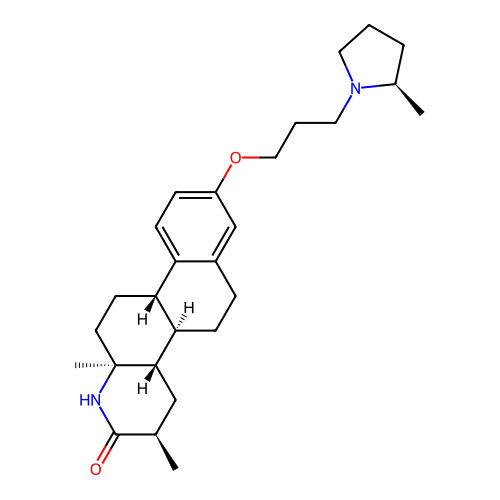
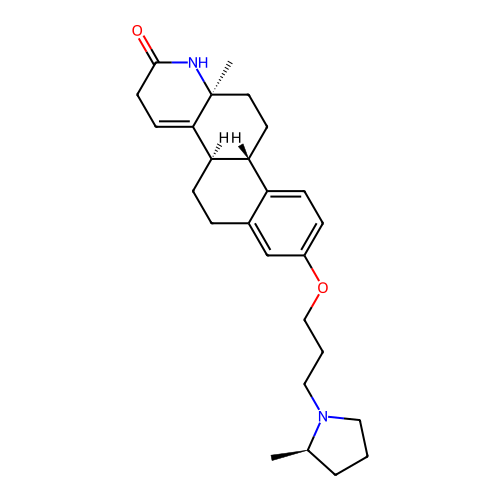
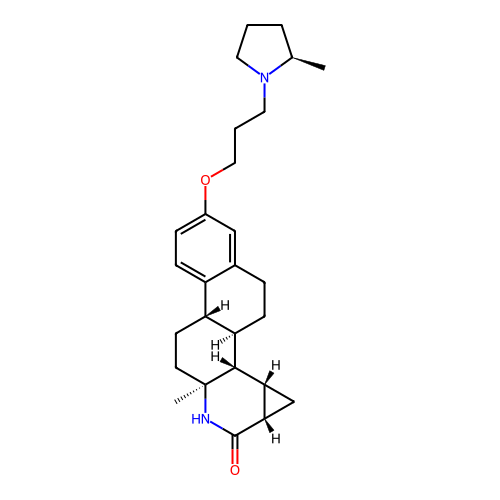
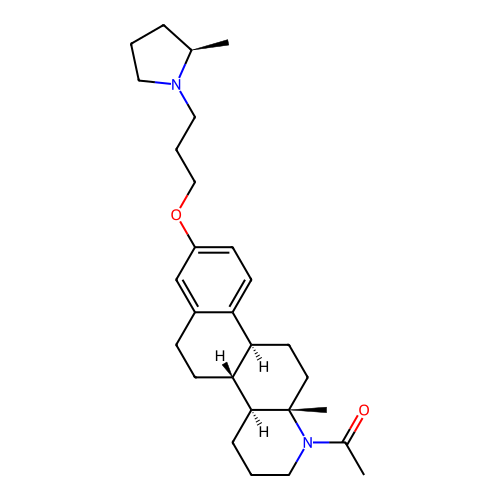
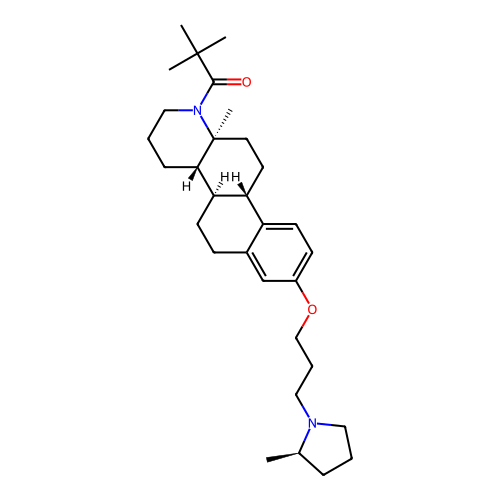
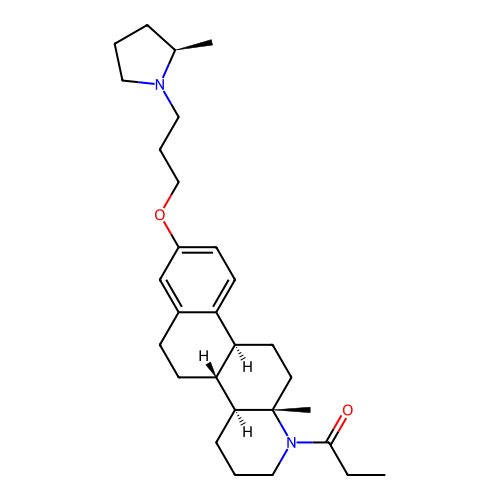
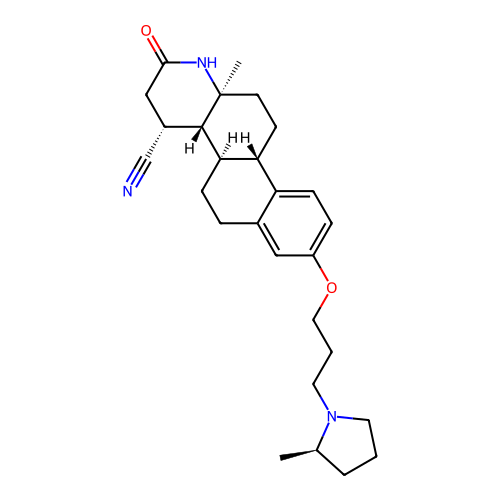
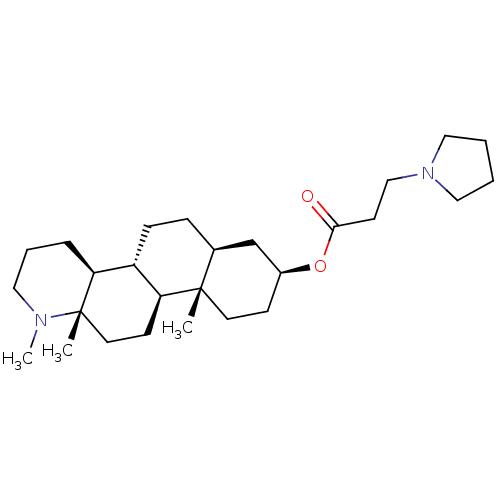
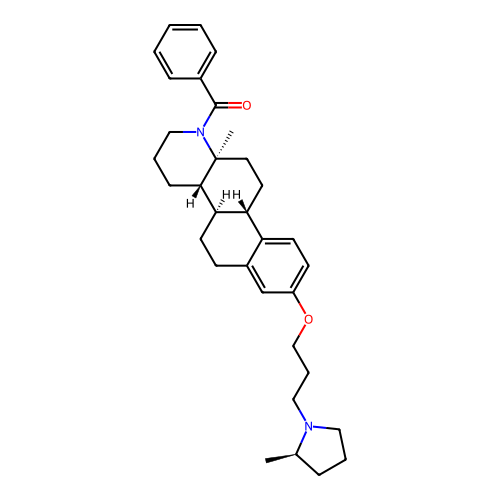
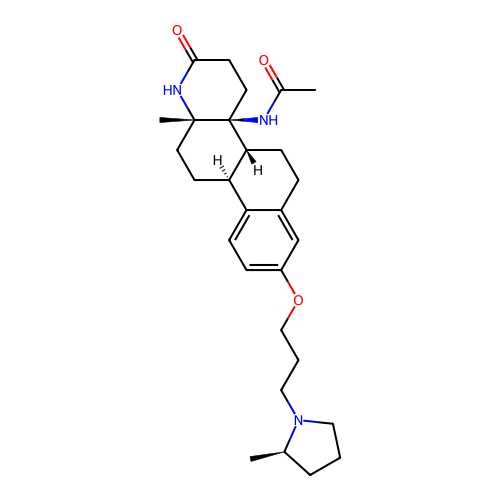
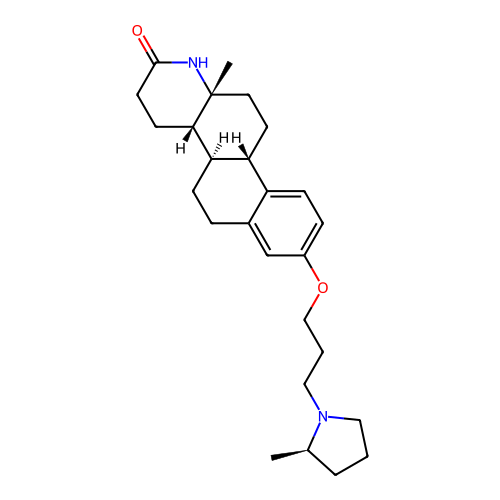
Report error Found 53 Enz. Inhib. hit(s) with all data for entry = 50021475
Affinity DataKi: 0.700nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 1.90nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 2.10nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 2.20nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 3.20nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 3.30nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 3.80nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 4.60nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 4.90nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 5.10nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 5.70nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 6.80nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 7.20nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 7.80nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 8.60nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 9.40nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 9.90nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of N-alpha-[methyl-3H]methylhistamine dihydrochloride from human Histamine H3 receptor expressed in CHO-K1 cell membrane assessed as inh...More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 22nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair
Affinity DataKi: 55nMAssay Description:Displacement of N-alpha-[methyl-3H]-methylhistamine dihydrochloride from Sprague-Dawley rat cerebral cortex membrane Histamine H3 receptor assessed a...More data for this Ligand-Target Pair