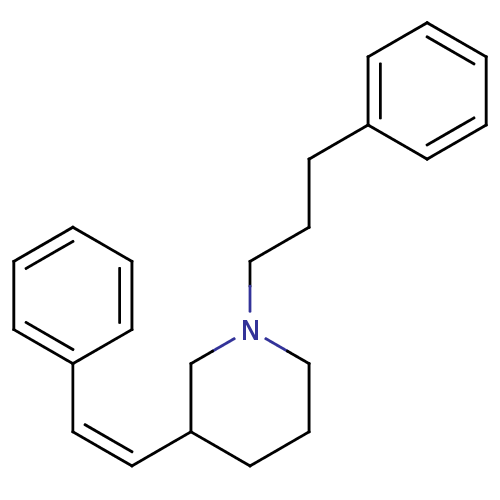
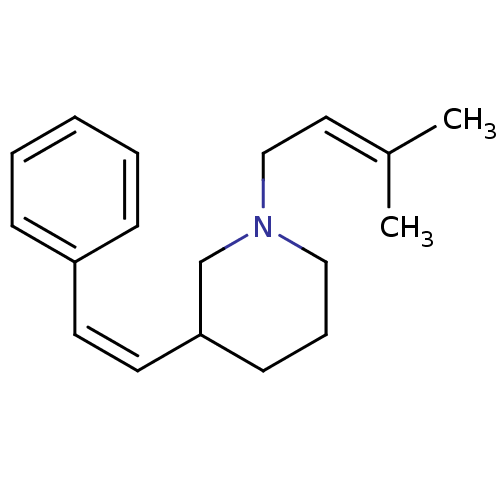
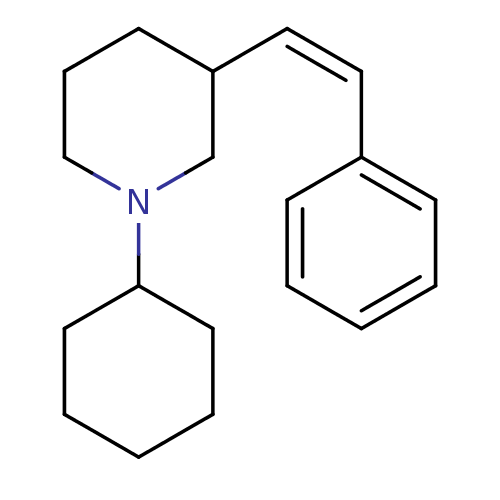
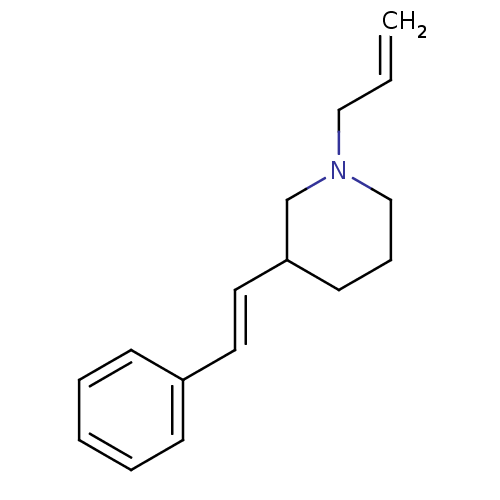
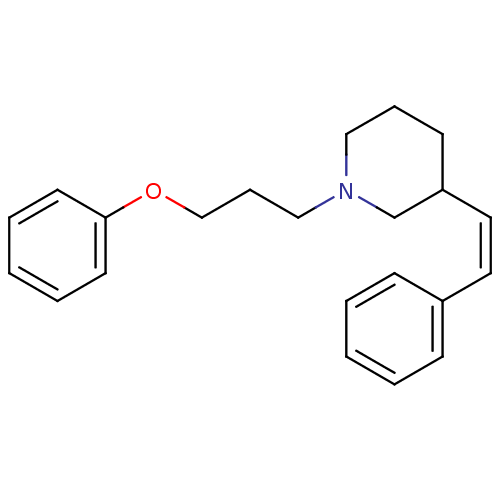
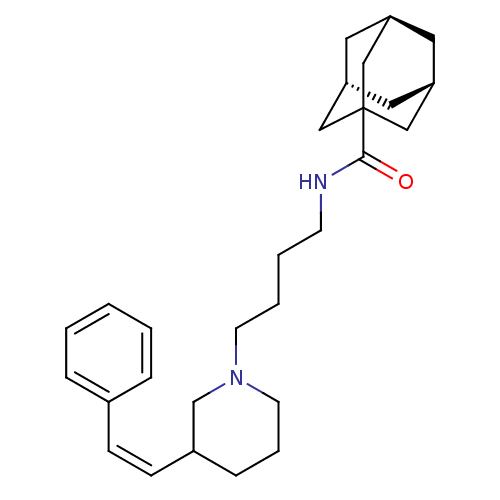
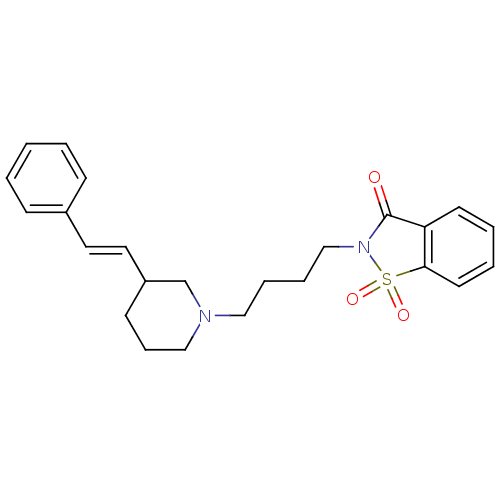
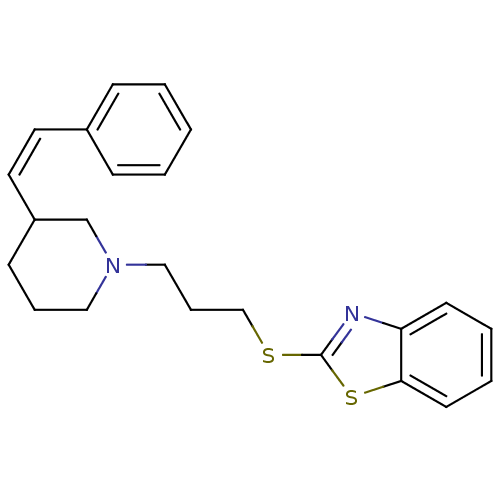
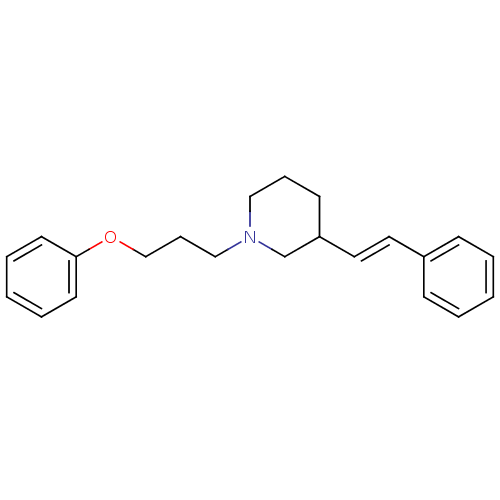
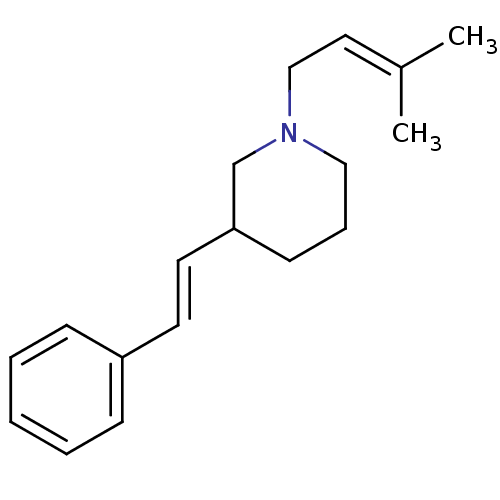
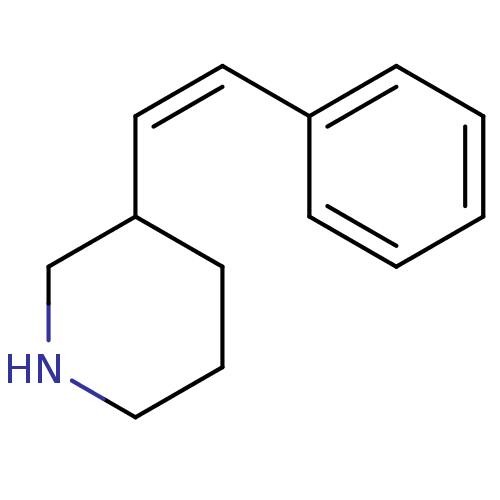
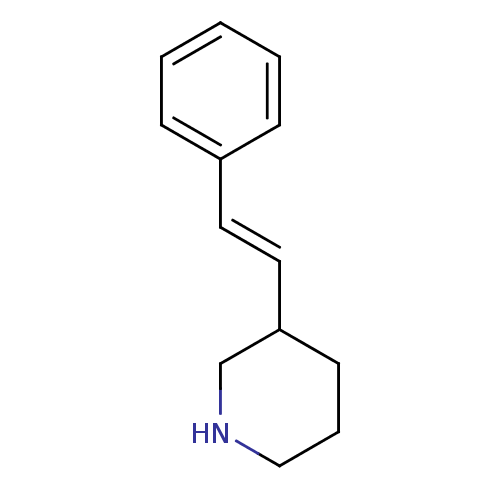
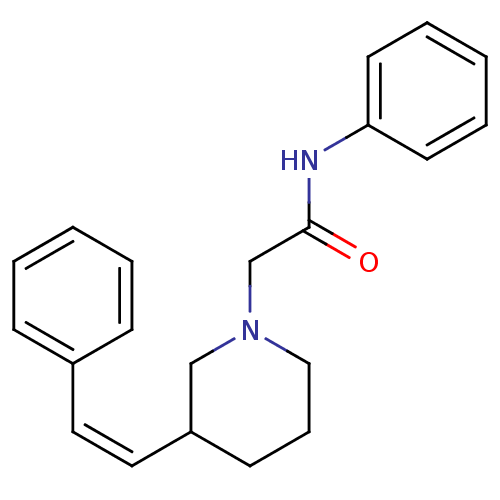
Report error Found 48 Enz. Inhib. hit(s) with all data for entry = 50029942
Affinity DataIC50: 0.600nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 0.800nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 33nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 38nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 320nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 423nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 630nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 640nMAssay Description:Binding affinity at sigma receptor by [3H](+)-SKF-10047 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 870nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.12E+3nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.24E+3nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.33E+3nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.44E+3nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.55E+3nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 5.12E+3nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 5.44E+3nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 5.84E+3nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D2 receptor in rat striatum by [3H]spiperone displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Binding affinity at dopamine D1 receptor in rat striatum by [3H]SCH-22390 displacement.More data for this Ligand-Target Pair