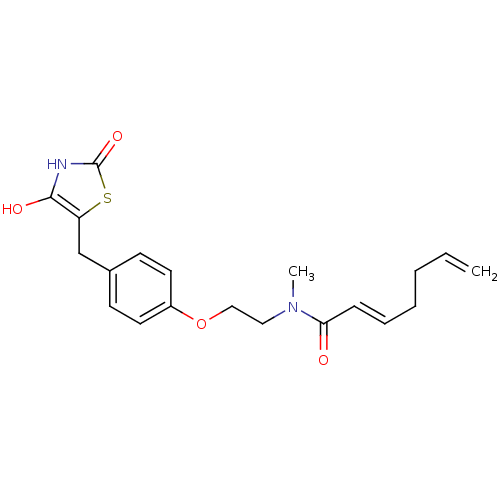
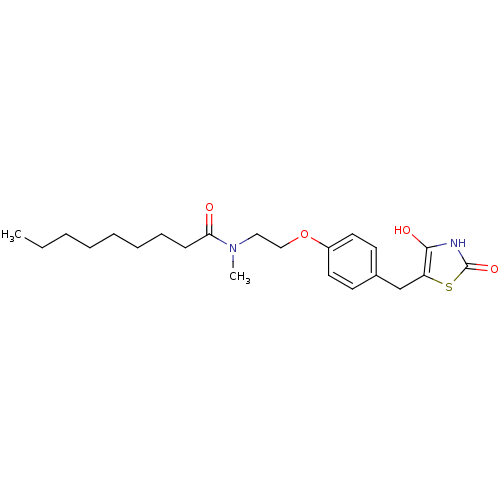
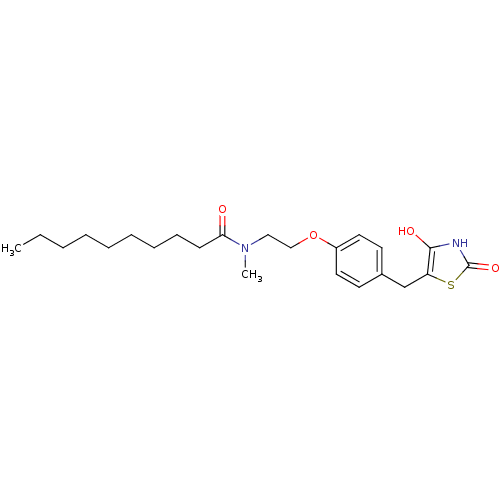
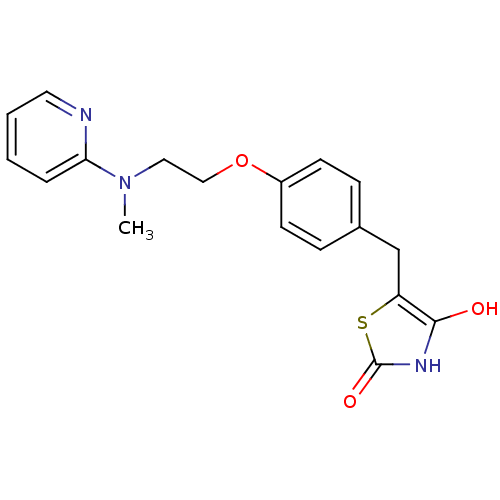
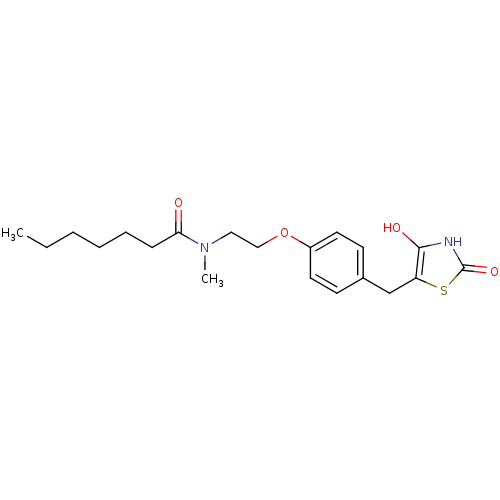
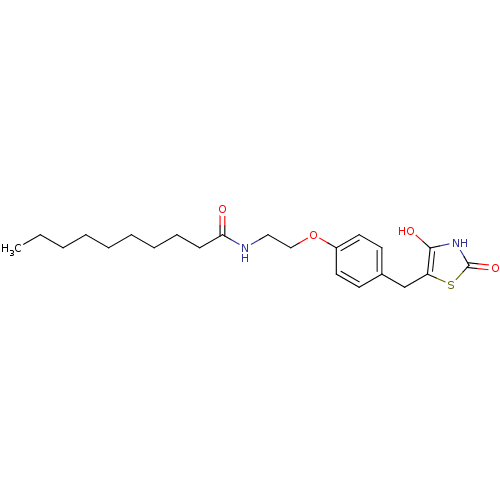
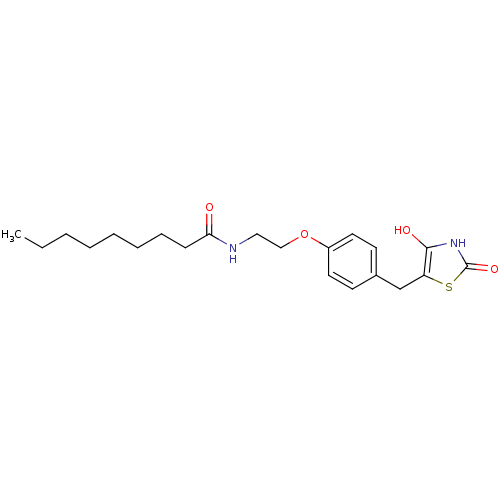
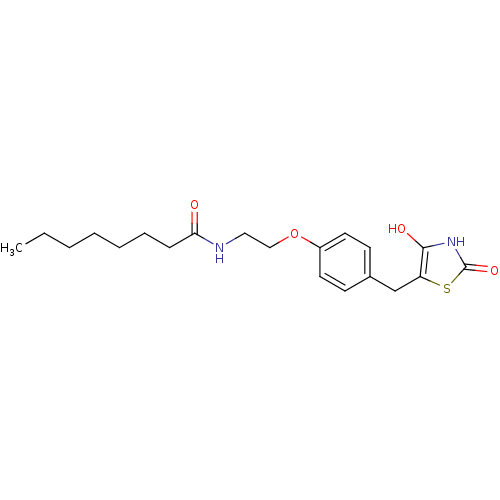
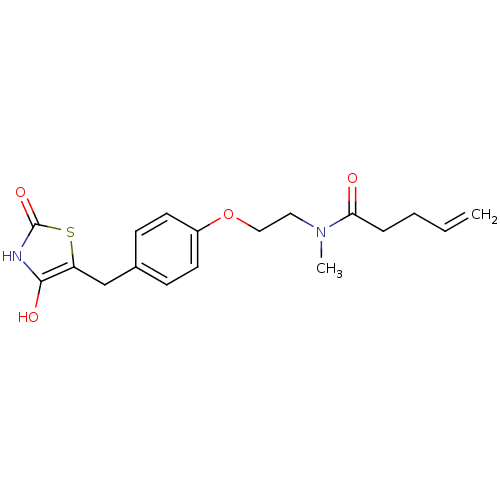
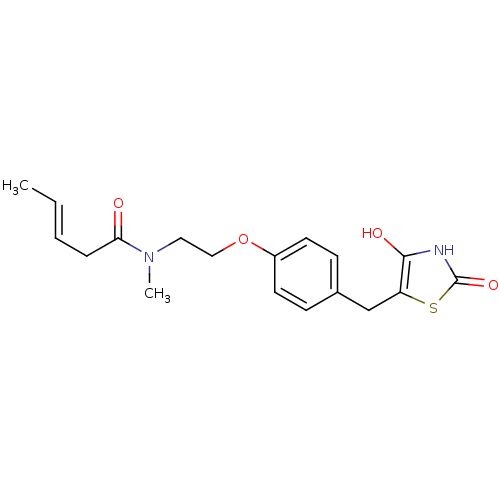
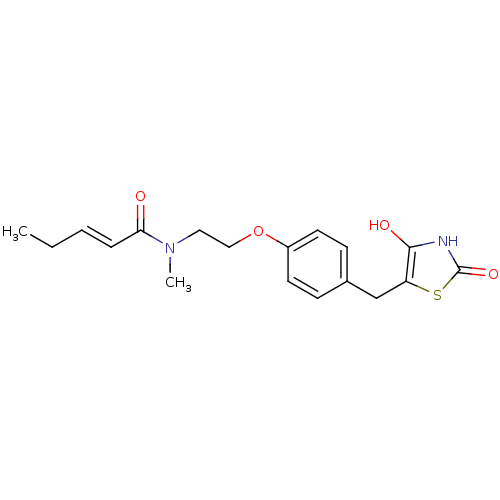
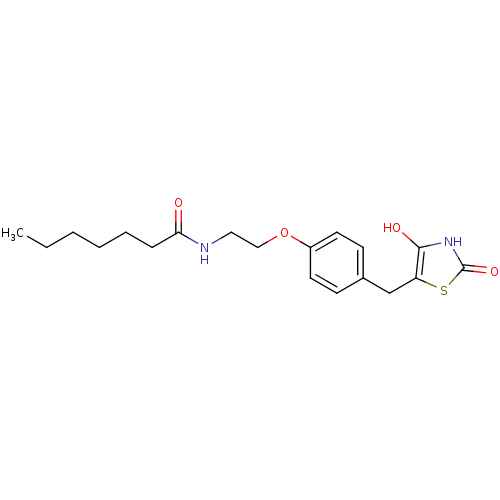
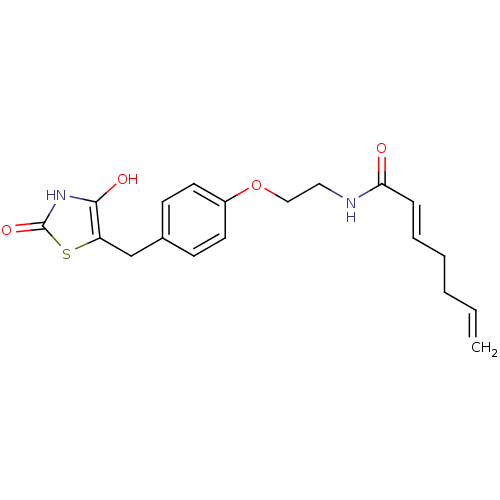
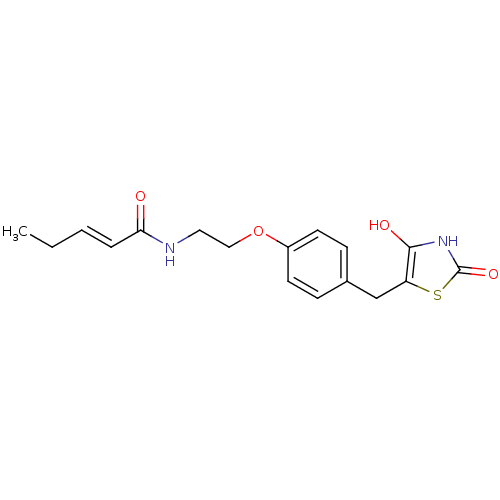
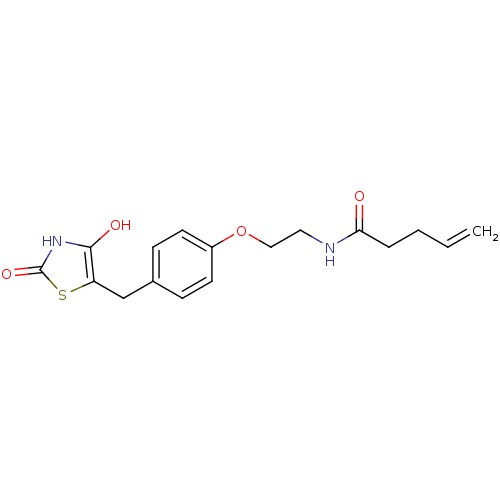
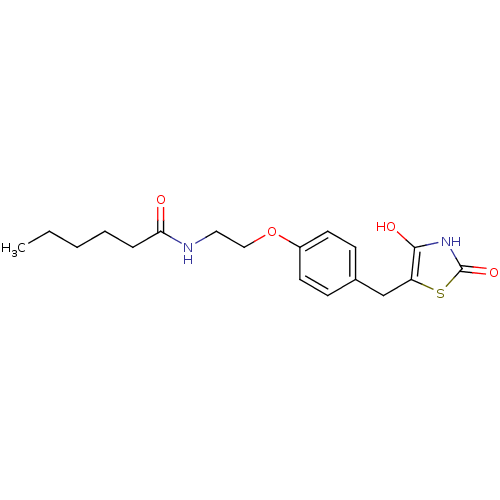
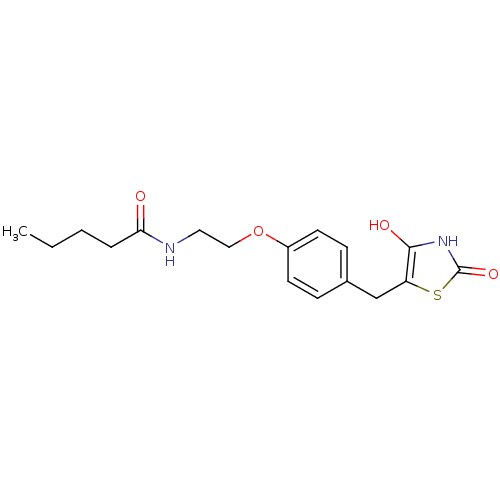
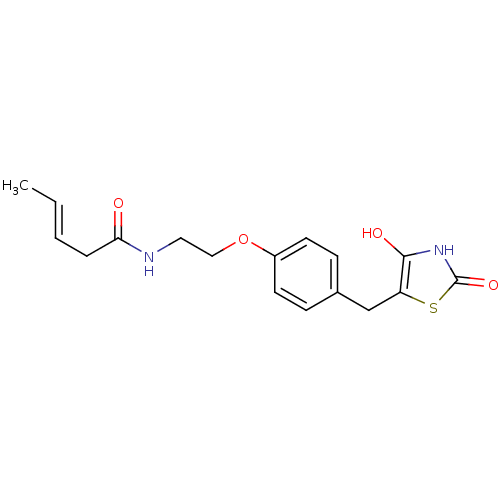
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50030018
Affinity DataKi: 18nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 48nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 49nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 270nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 290nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 600nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 1.20E+3nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 1.50E+3nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 6.80E+3nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity was determined by displacement of 20 nM [3H]- thiazolidinedione from 4 nM biotinylated human Peroxisome proliferator activated recep...More data for this Ligand-Target Pair