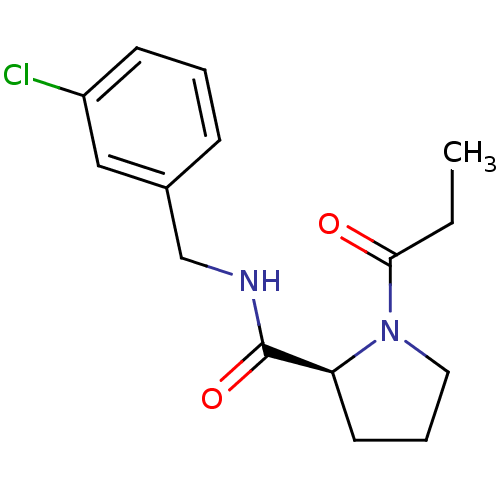
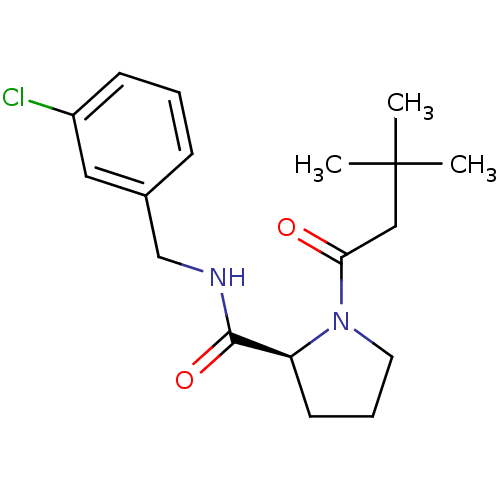
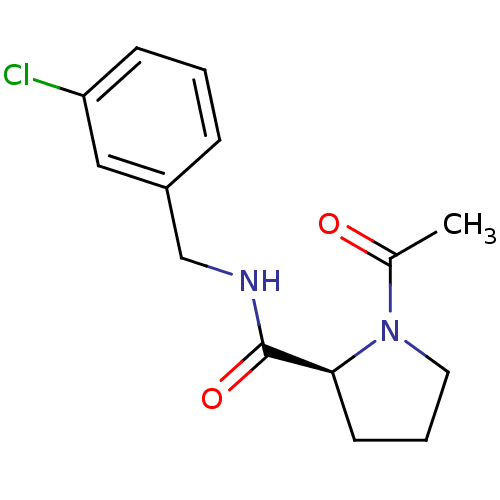
Report error Found 41 Enz. Inhib. hit(s) with all data for entry = 50031131
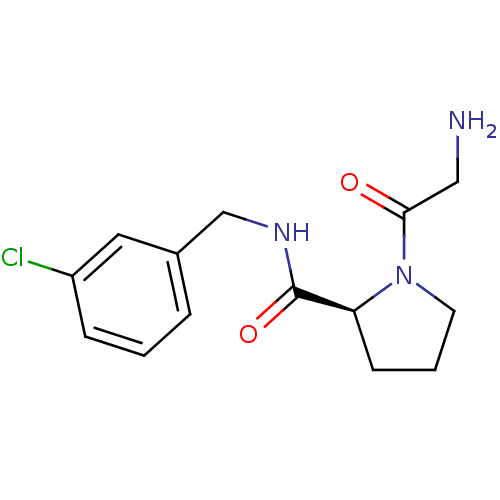
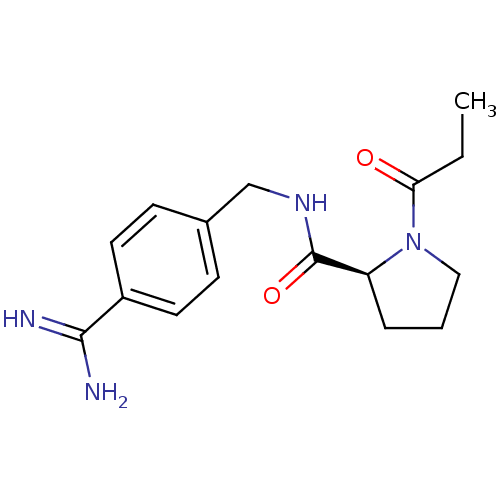
Affinity DataKi: 4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
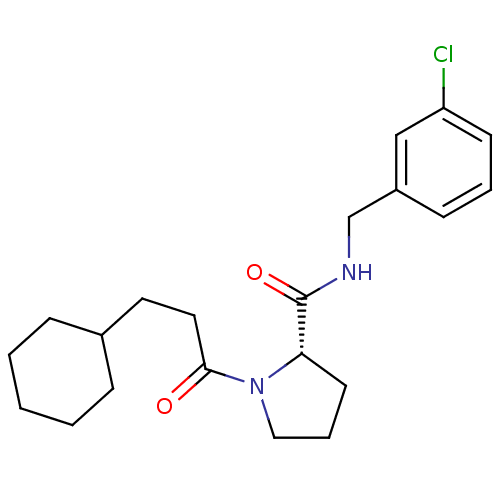
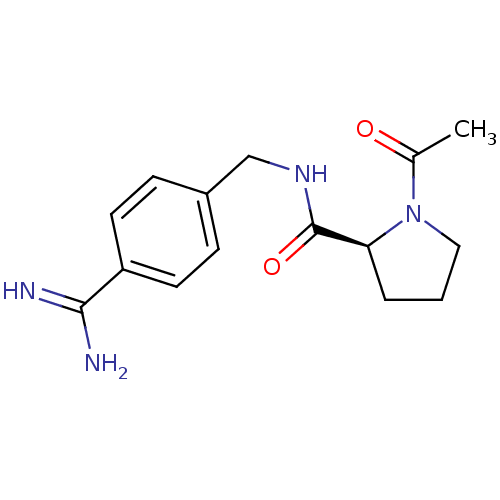
Affinity DataKi: 11nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
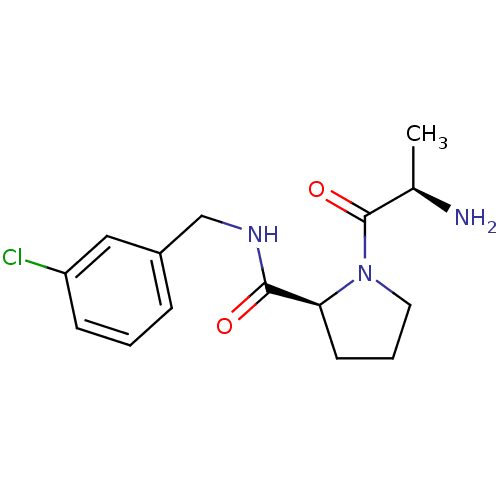
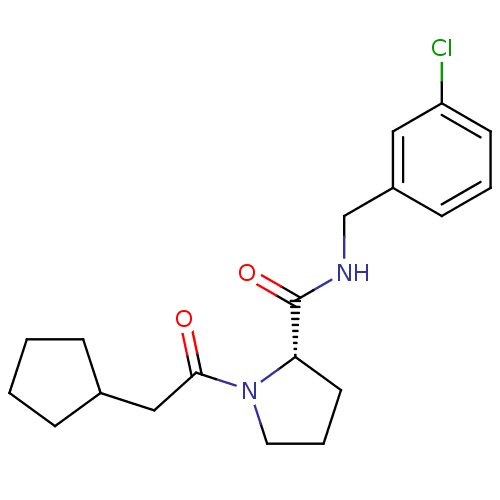
Affinity DataKi: 20nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
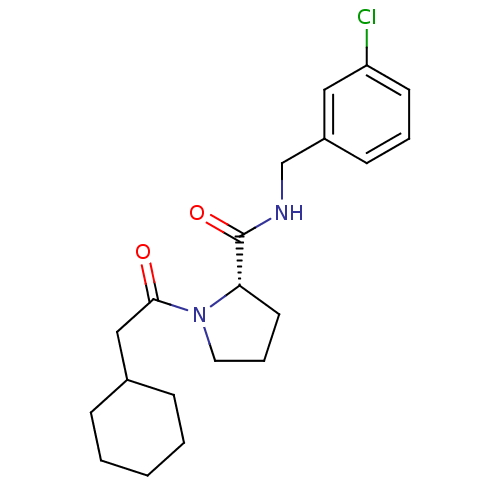
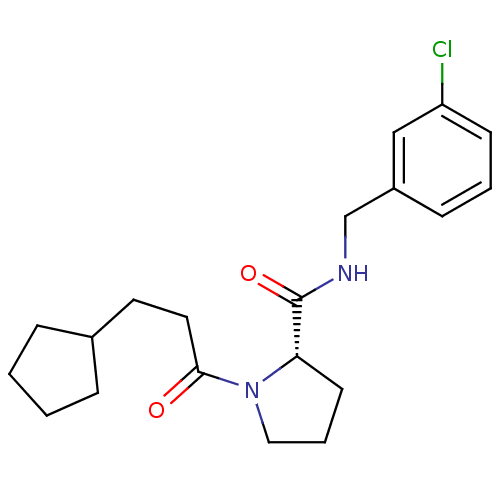
Affinity DataKi: 23nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 70nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 150nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 150nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 180nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 410nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 430nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 540nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 750nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 920nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 940nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.17E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.18E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.28E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.42E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.64E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.54E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.61E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.81E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.70E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.81E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.08E+3nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.97E+4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.30E+4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.45E+4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.90E+4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.61E+4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.24E+4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 9.29E+4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 9.43E+4nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.09E+5nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.55E+5nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.22E+5nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.44E+5nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.84E+5nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.46E+6nMAssay Description:Binding affinity to human thrombin by standard kinetic photometric assayMore data for this Ligand-Target Pair
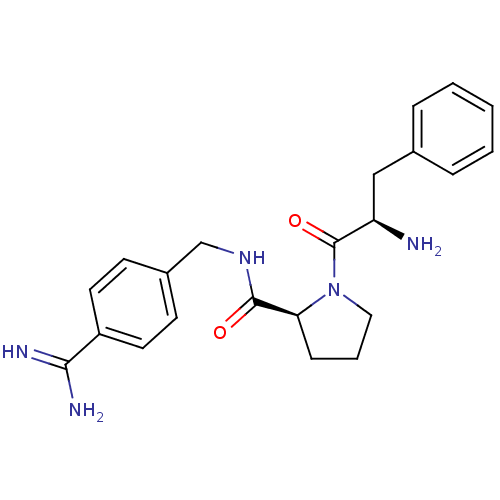
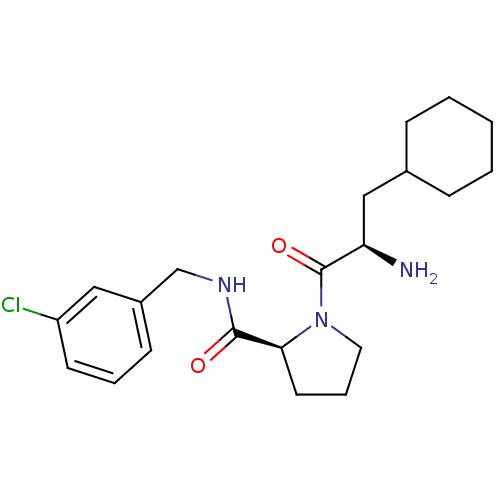
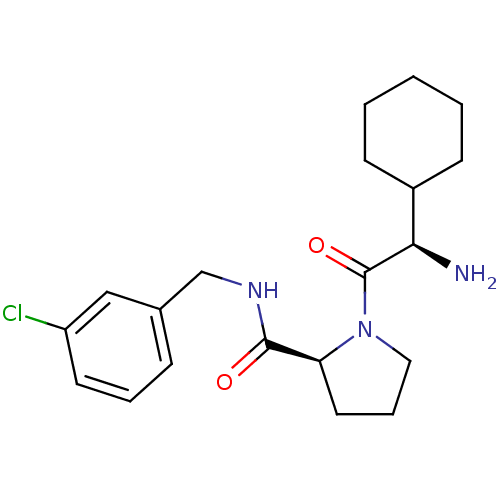
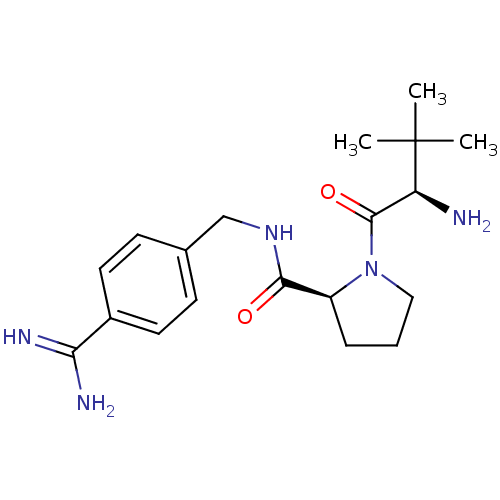

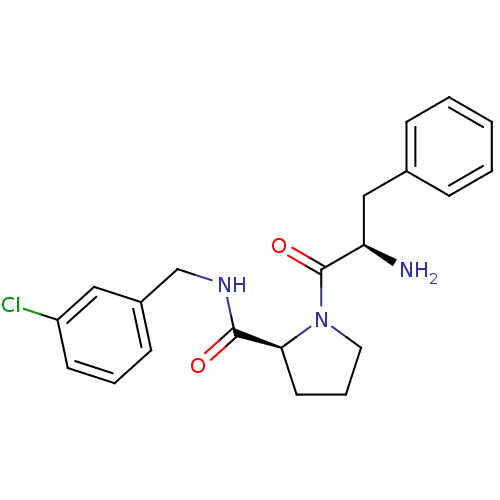
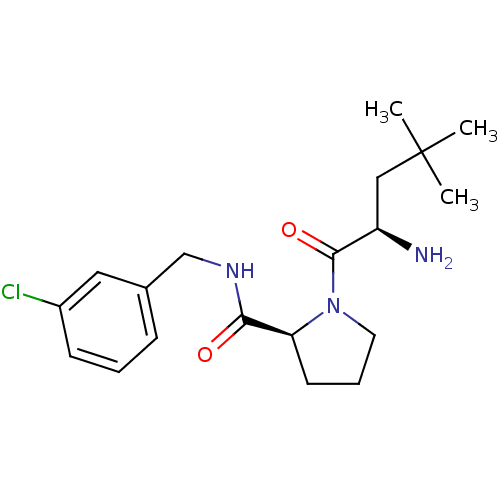
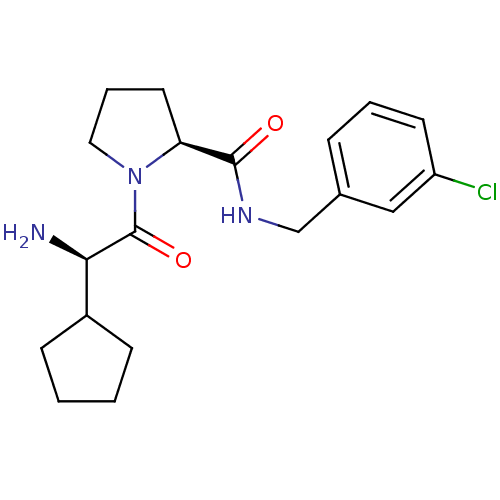
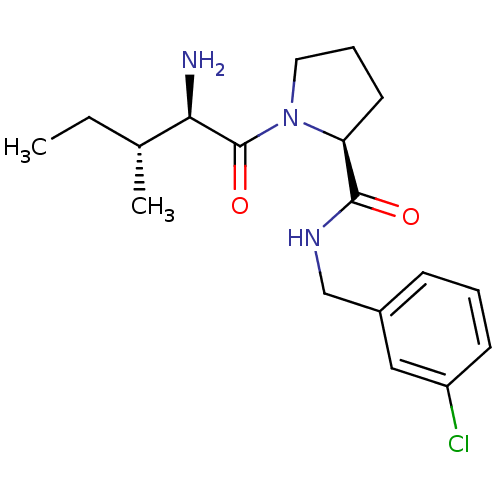
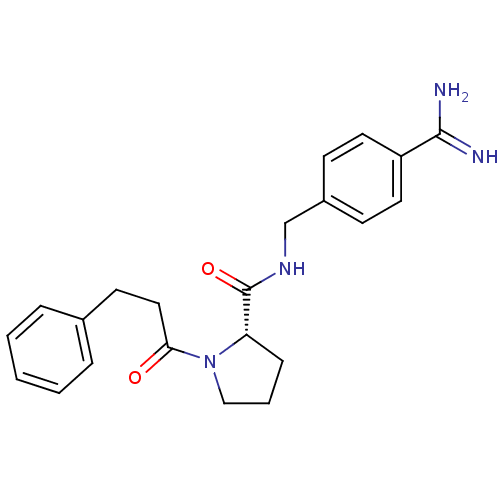
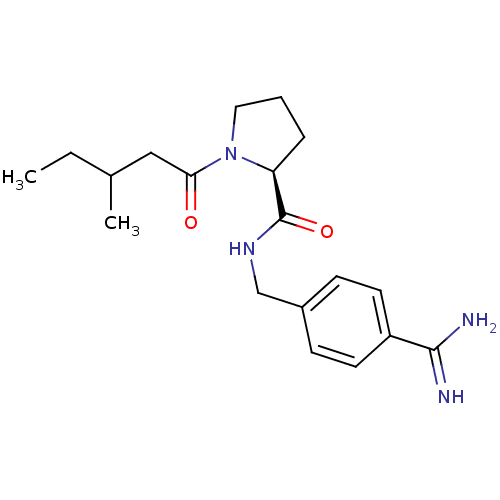
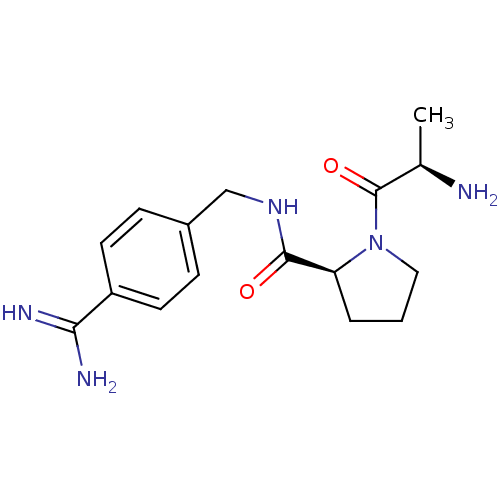
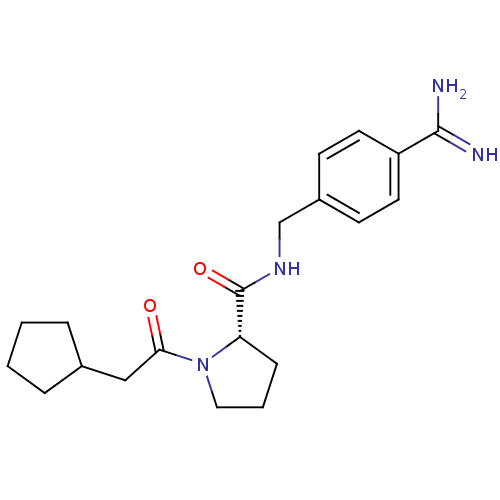
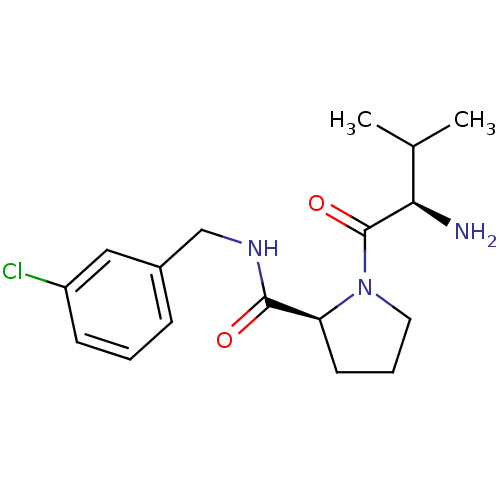
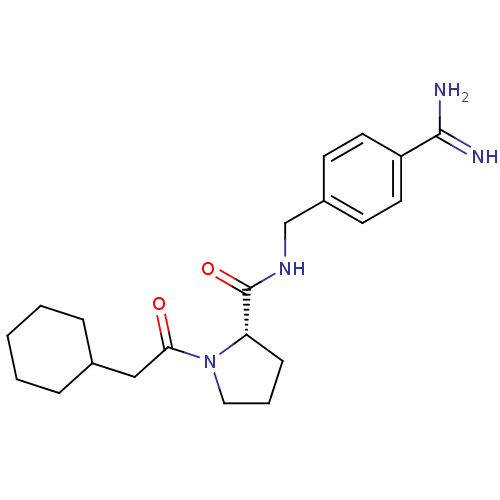
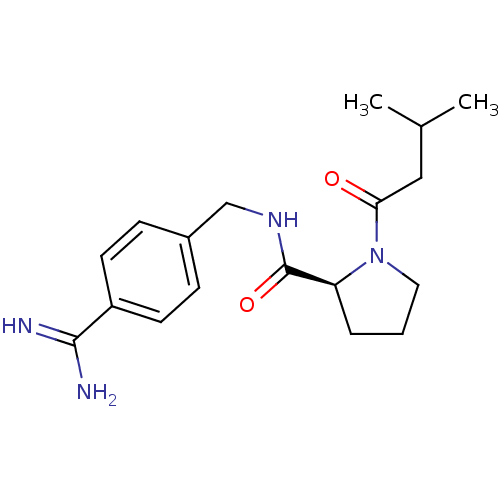

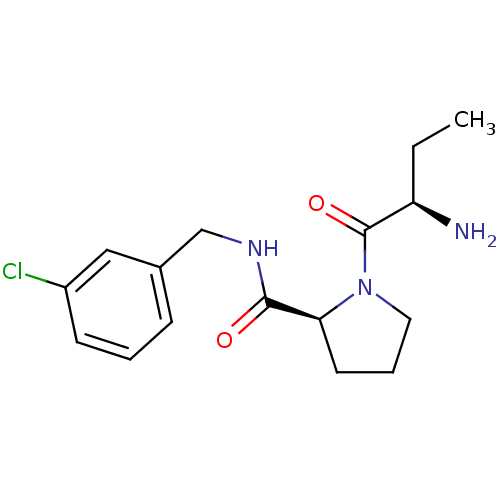
 3D Structure (crystal)
3D Structure (crystal)