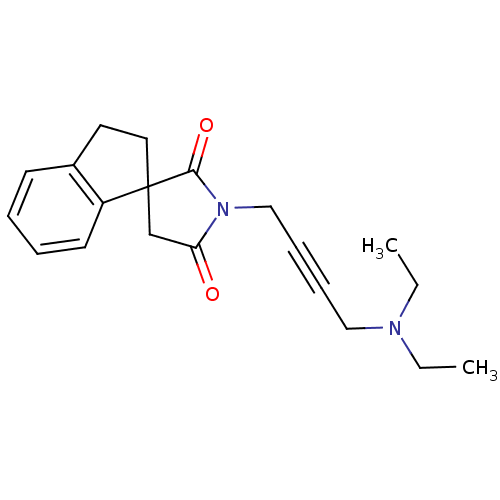
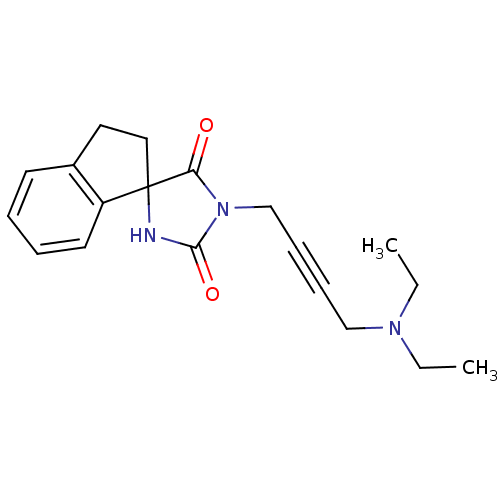
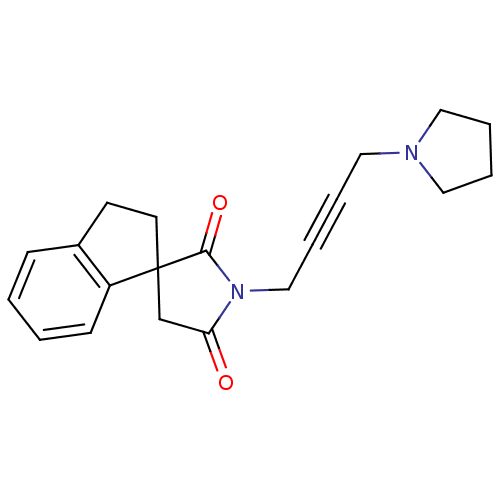
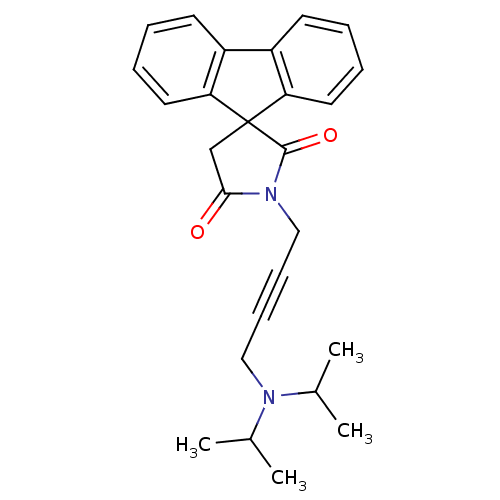
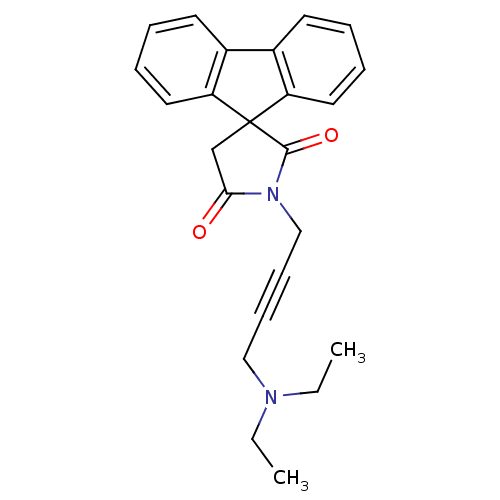
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 50034925
Affinity DataKi: 0.0300nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 5.20nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 100nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 120nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 267nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 380nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 380nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 440nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 460nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 530nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 640nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 680nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 950nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 1.05E+3nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 1.59E+3nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 1.62E+3nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 1.63E+3nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 2.09E+3nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: 2.30E+3nMAssay Description:Displacement of [3H]pirenzepine from muscarinic M1 receptor of rat cortex homogenates.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]QNB (quinuclidinyl benzylate) from muscarinic M2 receptor of rat heart homogenates.More data for this Ligand-Target Pair
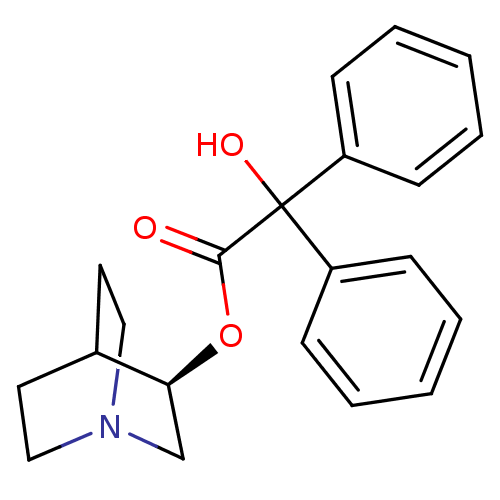
 3D Structure (crystal)
3D Structure (crystal)