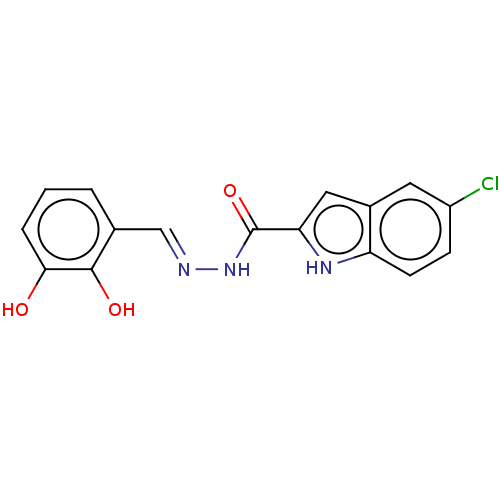
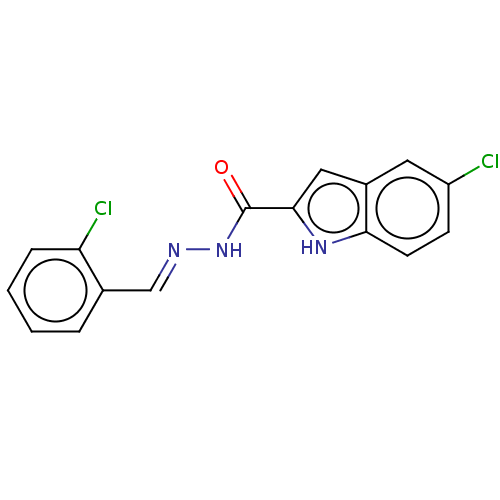
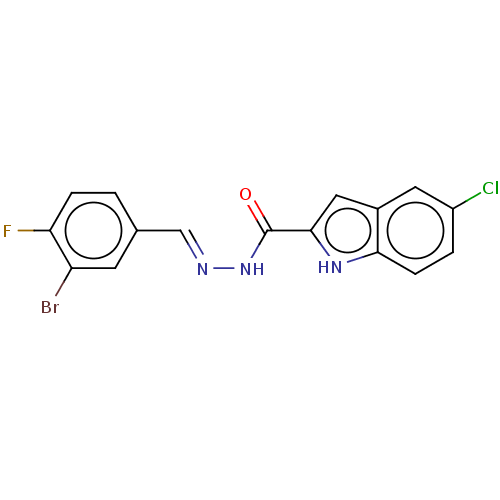
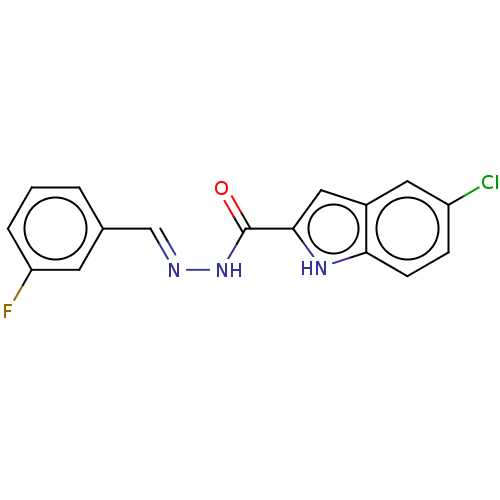
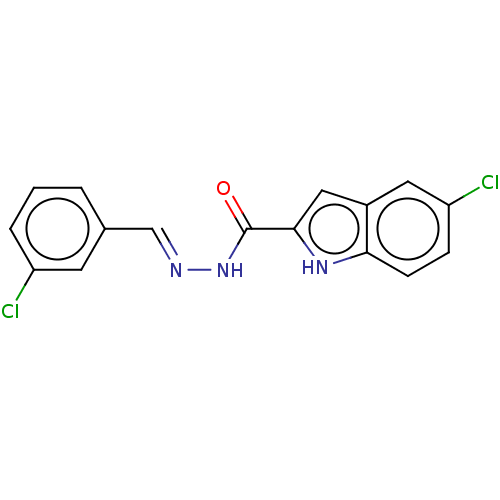
Report error Found 32 Enz. Inhib. hit(s) with all data for entry = 7991
Affinity DataIC50: 500nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+3nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 7.50E+3nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 8.70E+3nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 1.34E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 1.36E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 1.44E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 1.44E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 1.46E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.15E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.25E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.26E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.28E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.43E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.46E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 2.52E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 3.25E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 3.43E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 3.72E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 3.94E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 4.14E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 4.24E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 4.35E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 4.45E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 4.84E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 5.34E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 5.36E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair
Affinity DataIC50: 5.65E+4nMAssay Description:β-glucuronidase activity was determined in accordance to method used by Taha et al. [Taha et al., Bioorg. Med. Chem., 23:7394-7404] by measuring...More data for this Ligand-Target Pair