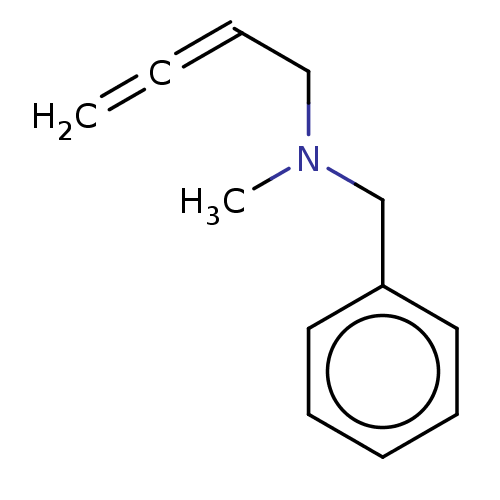
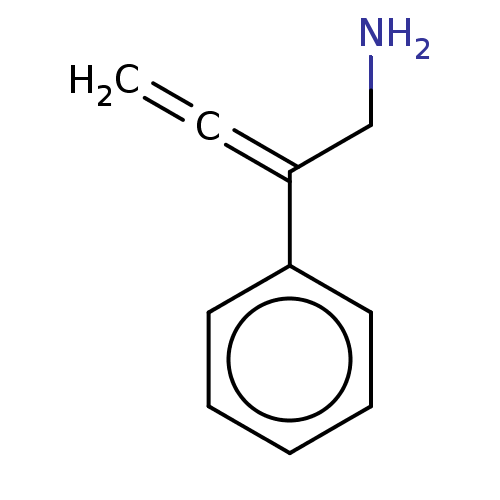
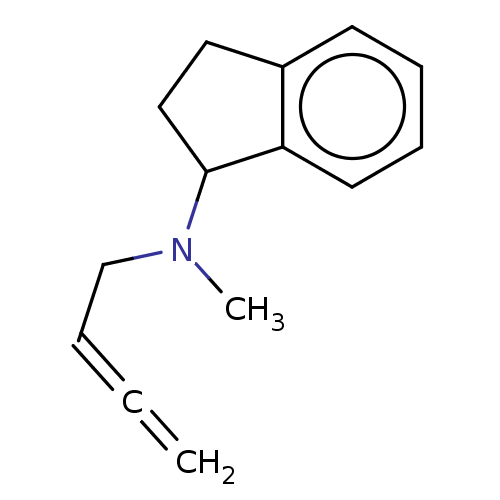
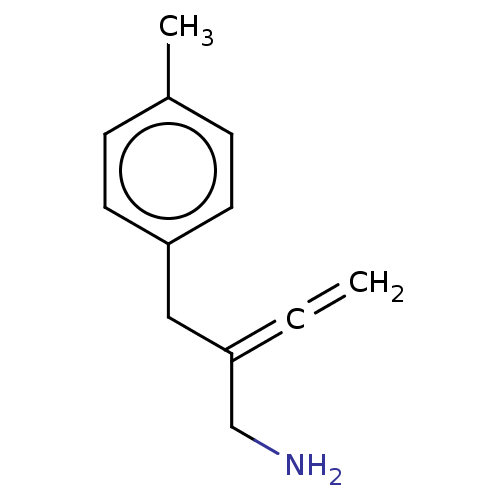
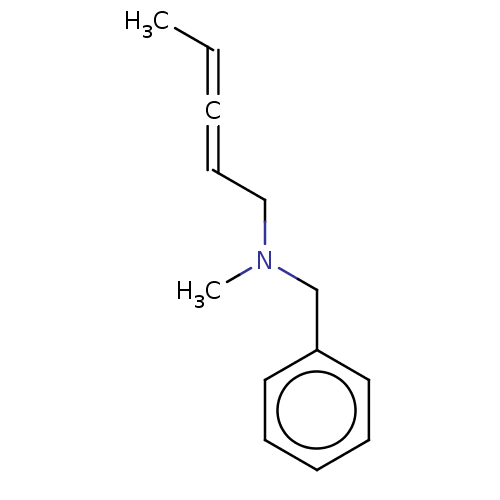
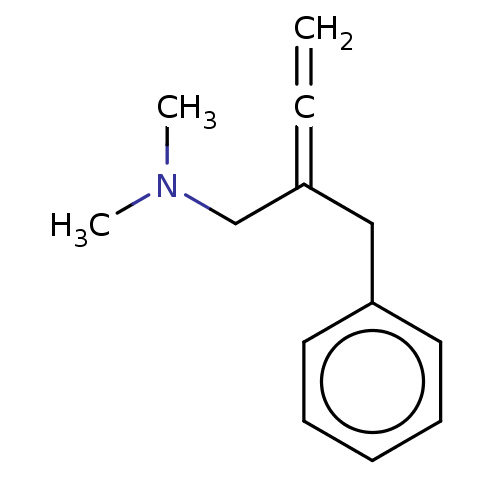
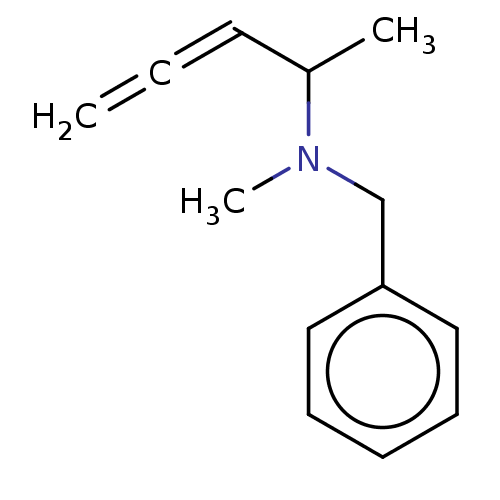
Report error Found 38 Enz. Inhib. hit(s) with all data for entry = 50048660
Affinity DataIC50: 0.700nMAssay Description:In vitro Inhibitory activity of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT (serotonin)More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 260nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 360nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 360nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 440nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 650nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 680nMAssay Description:In vitro Inhibitory activity of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA (phenethylamine)More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 2.30E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 4.70E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 6.20E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 7.20E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+3nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+4nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial 5-HT by displacing 2.5 uM of [14C]5-HT.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:In vitro inhibition of Monoamine oxidase at rat hyphalamic mitochondrial PEA by displacing 2.5 uM of [14C]PEA.More data for this Ligand-Target Pair
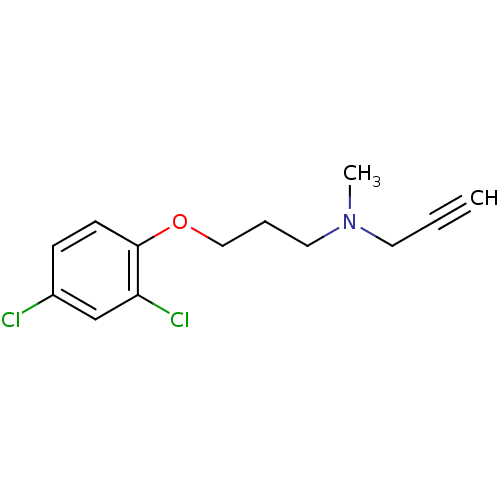
 3D Structure (crystal)
3D Structure (crystal)