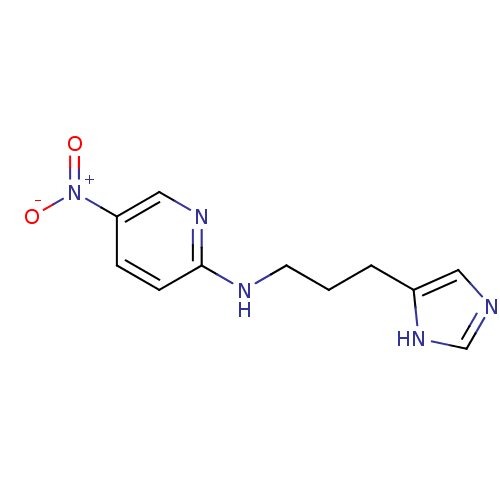
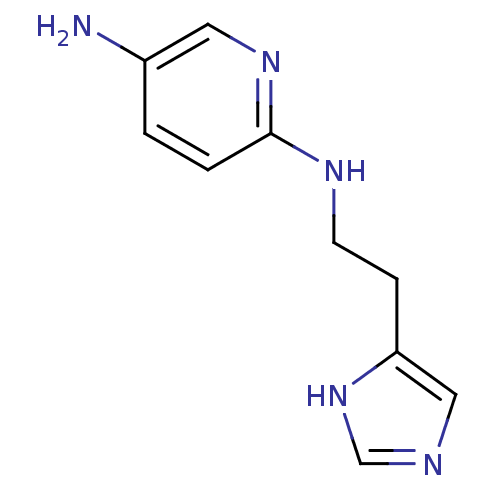
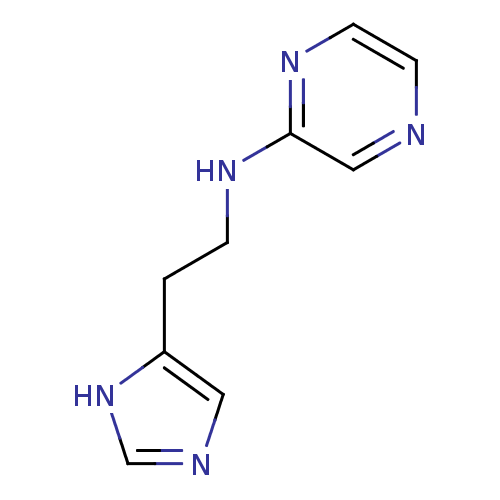
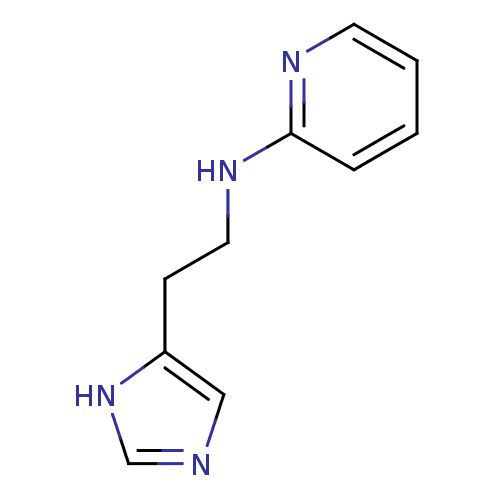
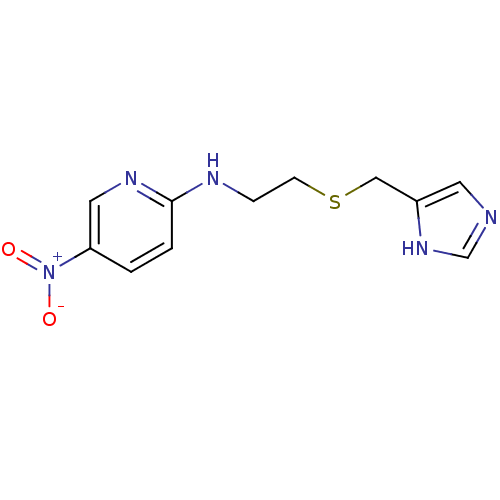
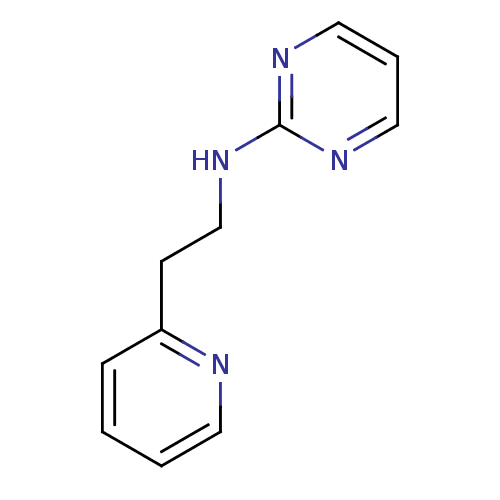
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50005115
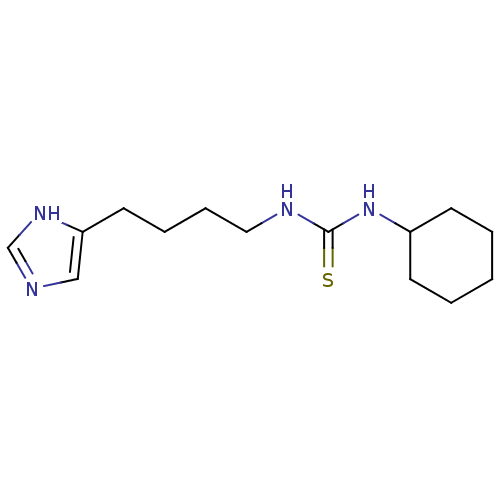
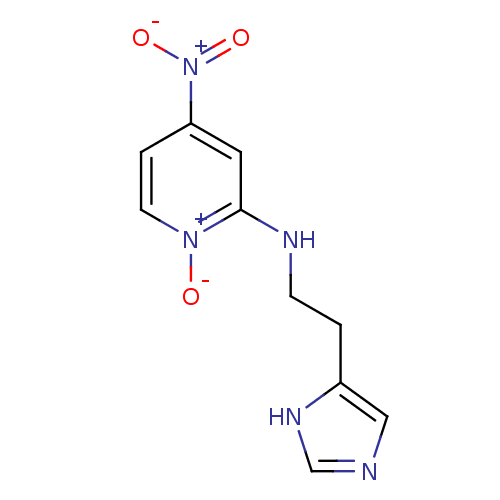
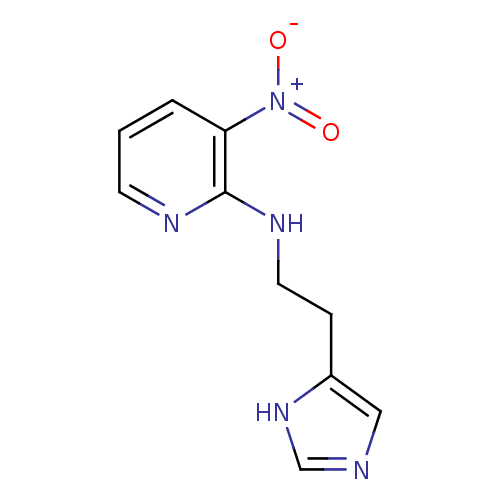
Affinity DataKi: 4.30nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
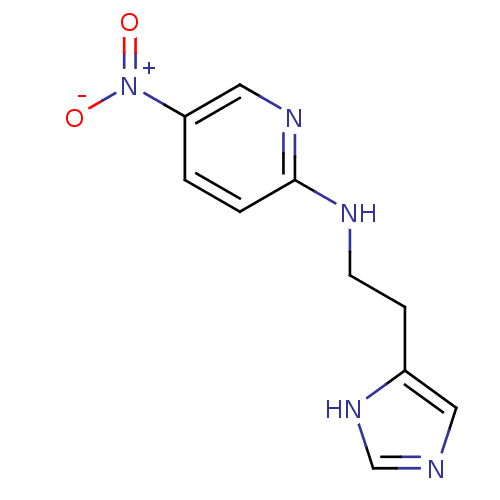
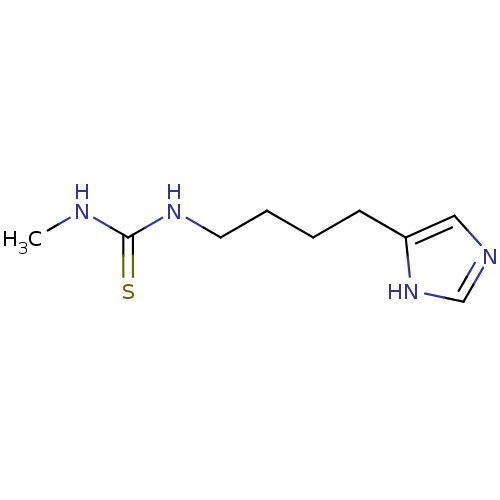
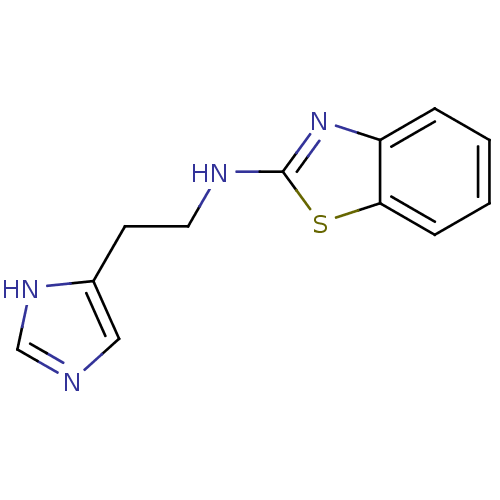
Affinity DataKi: 4.80nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
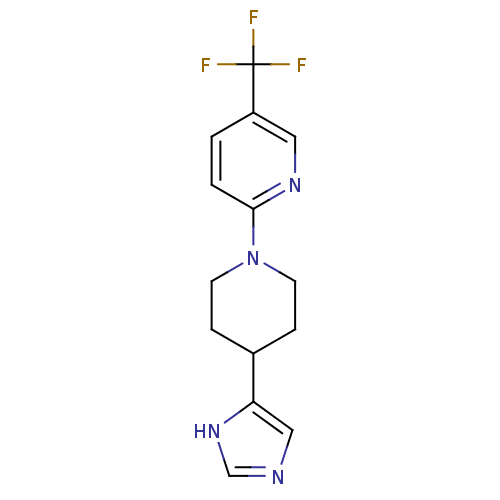
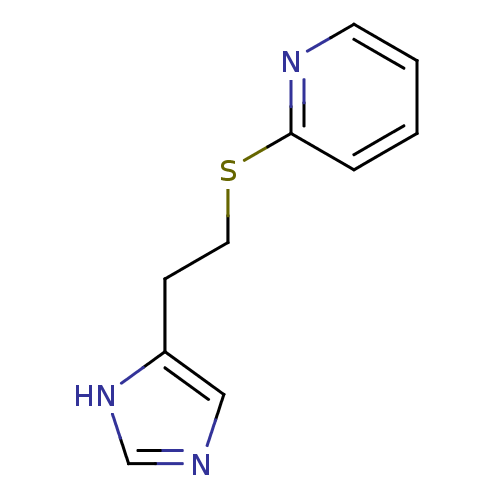
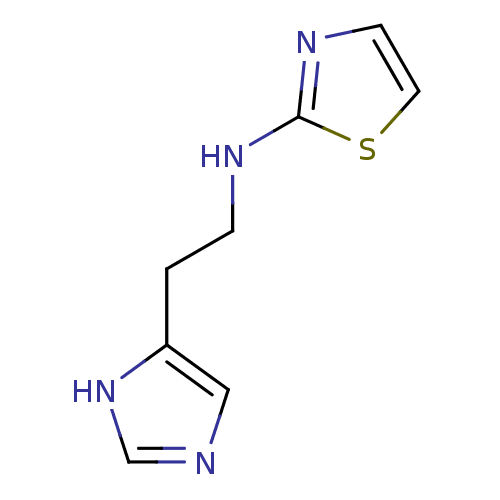
Affinity DataKi: 13nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
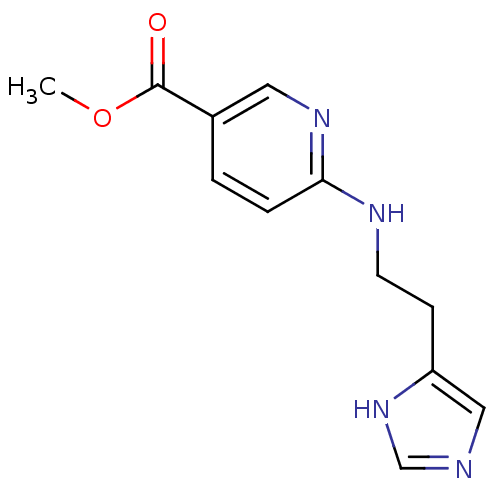
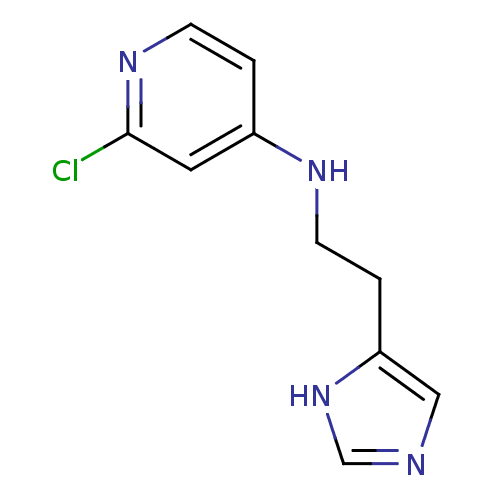
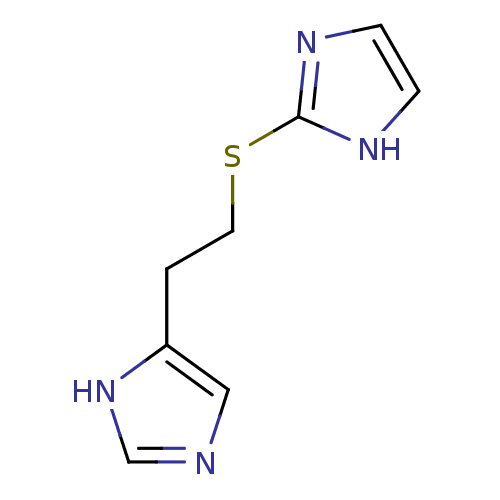
Affinity DataKi: 17nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 29nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 53nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 70nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 92nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 186nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 330nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 340nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 400nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+3nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:In vitro antagonism of the histamine H3-receptor in the rat cerebral cortex.More data for this Ligand-Target Pair