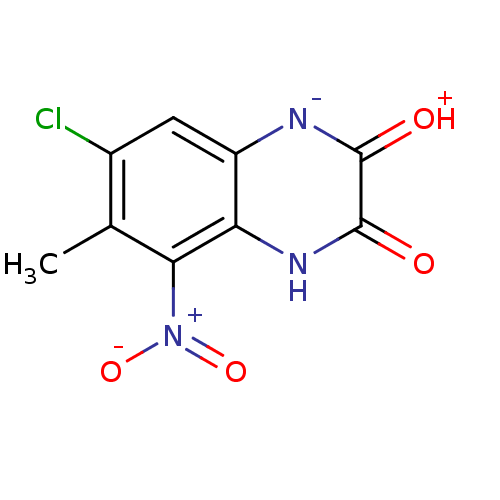
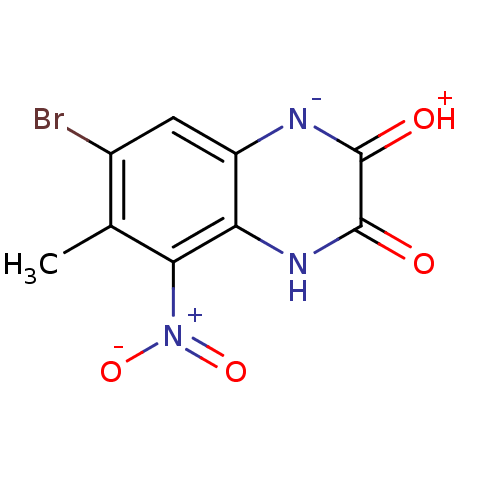
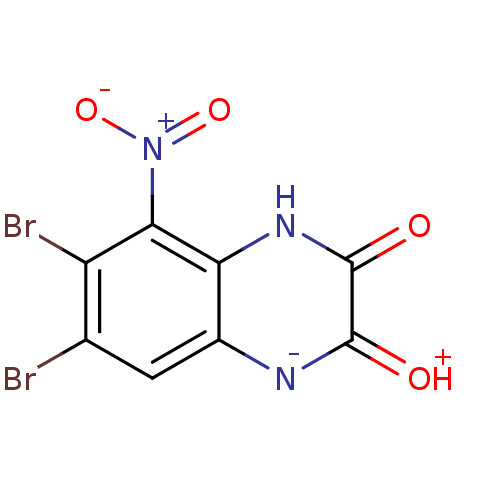
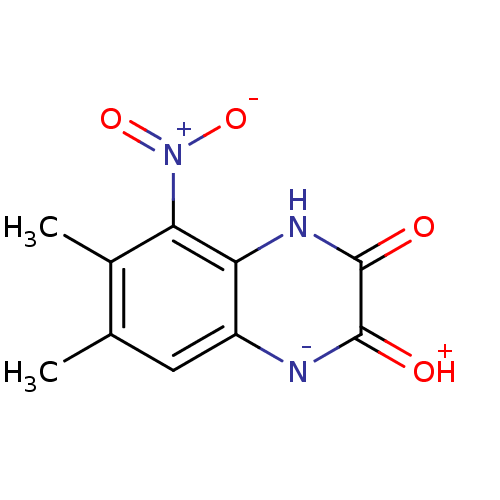
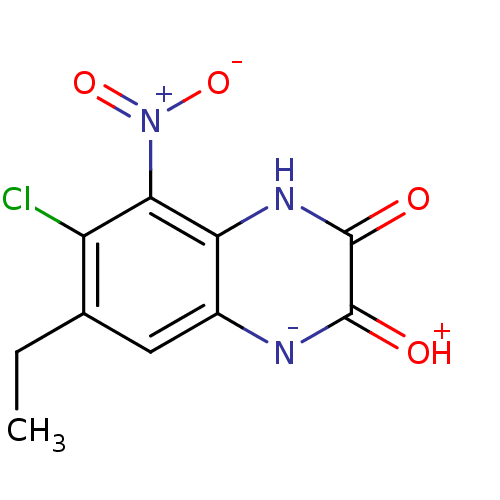
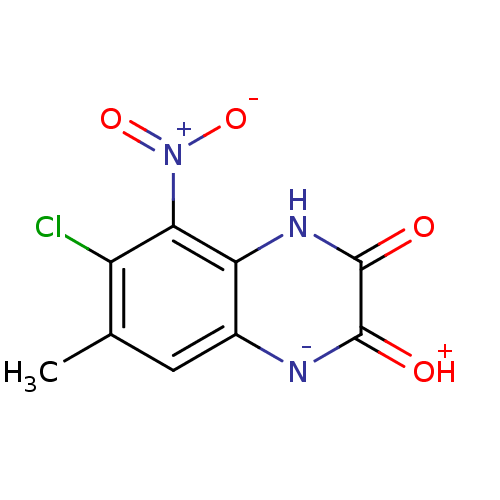
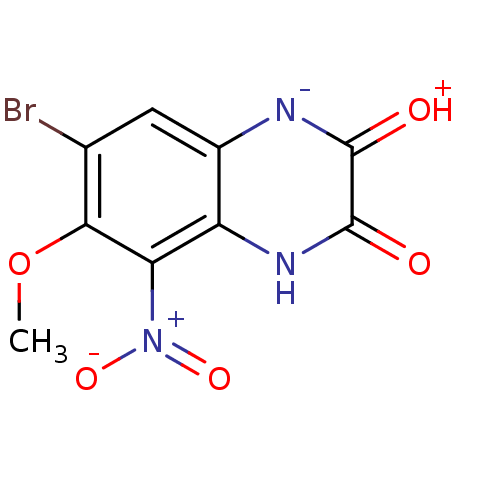
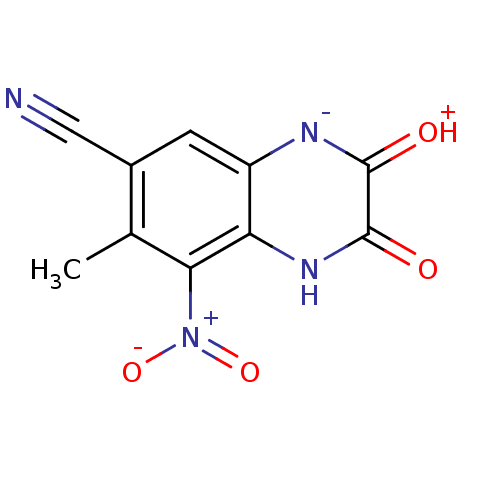
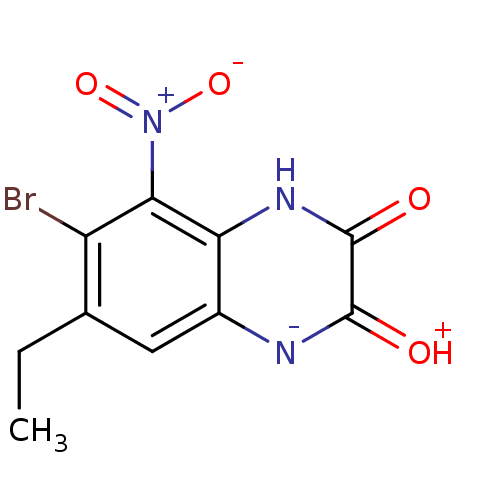
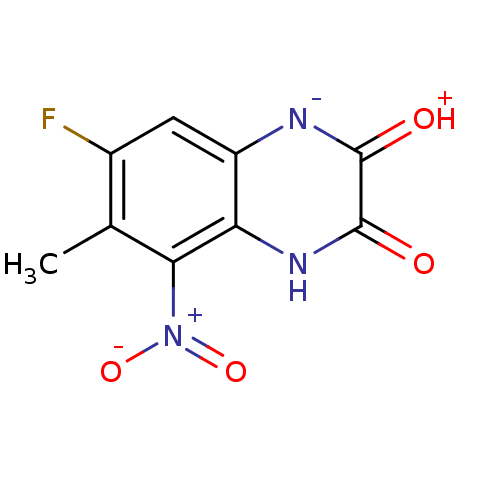
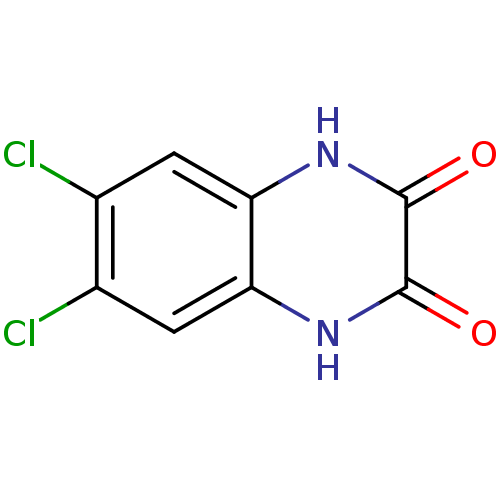
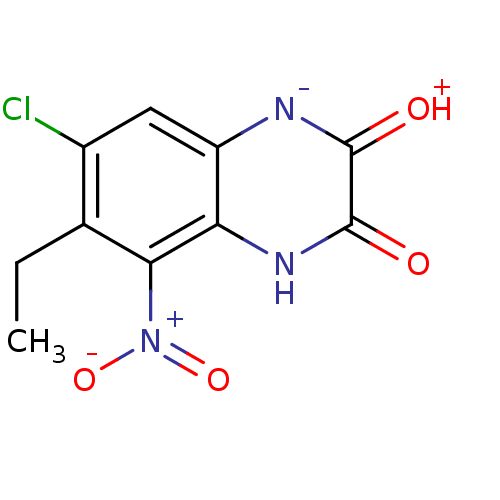
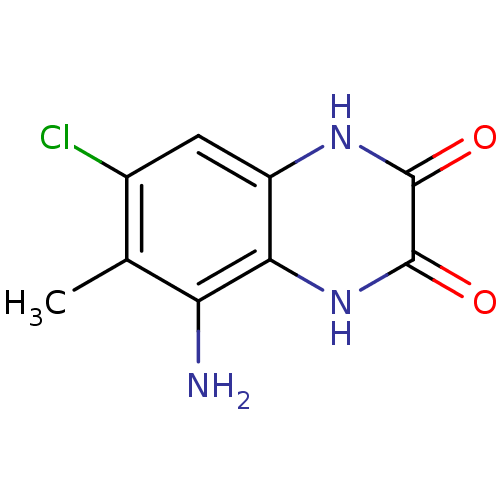
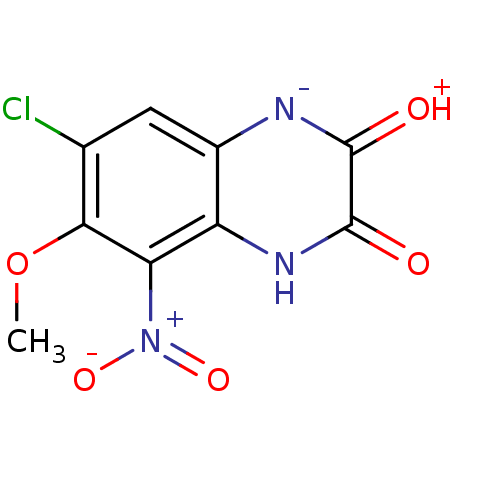
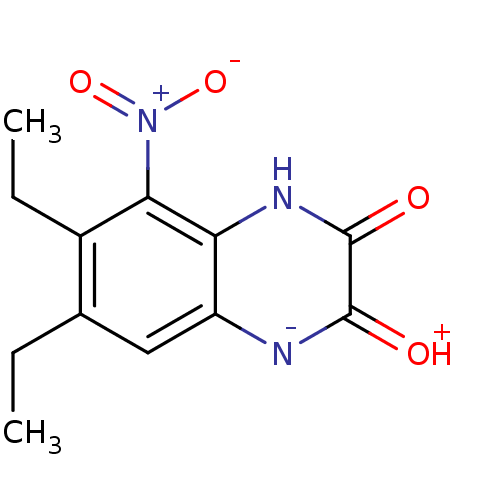
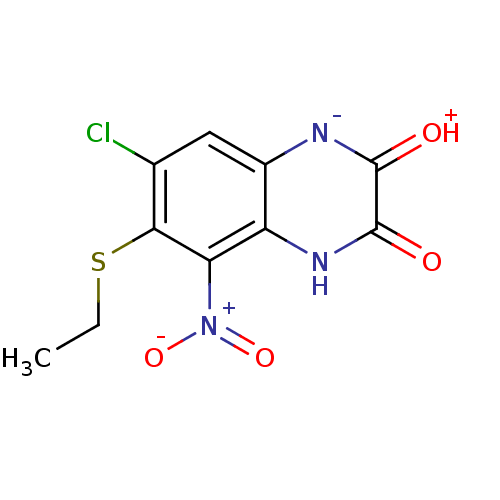
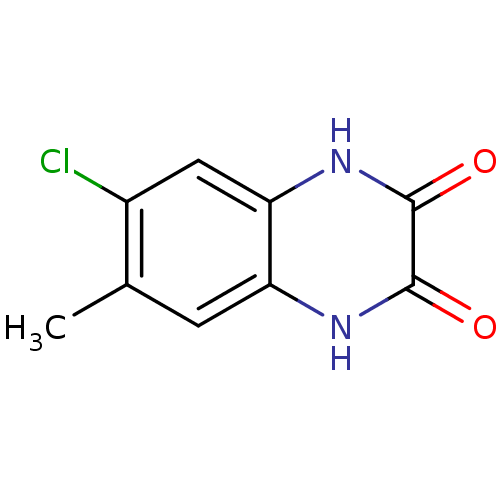
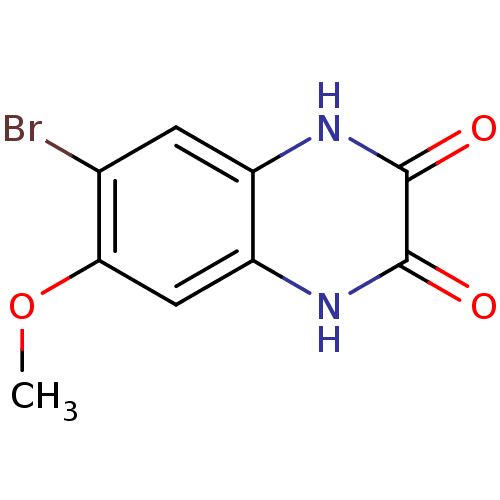
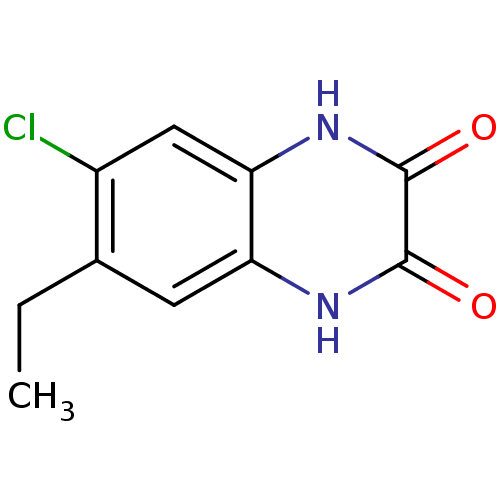
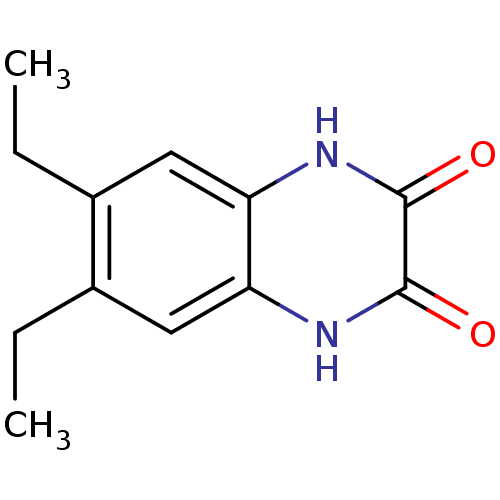
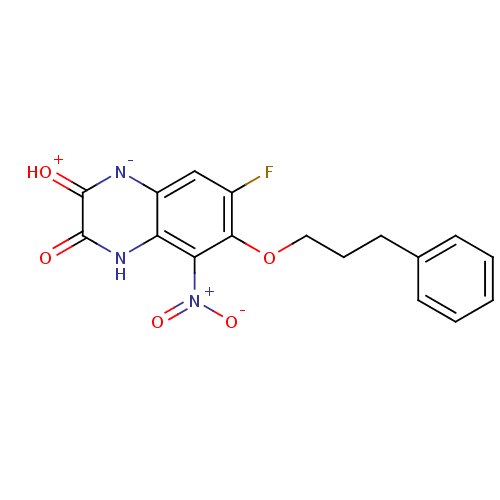
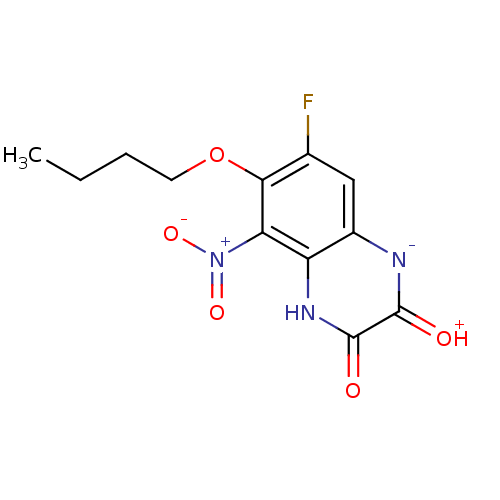
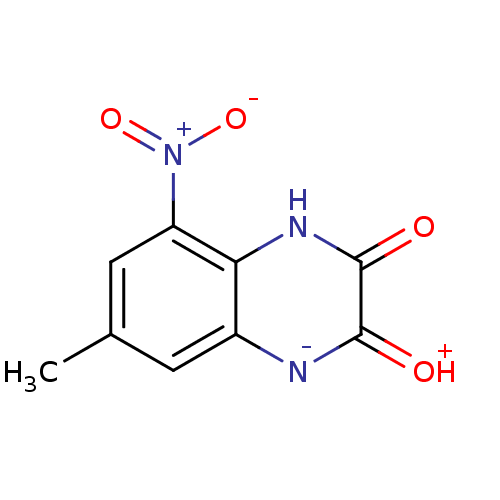
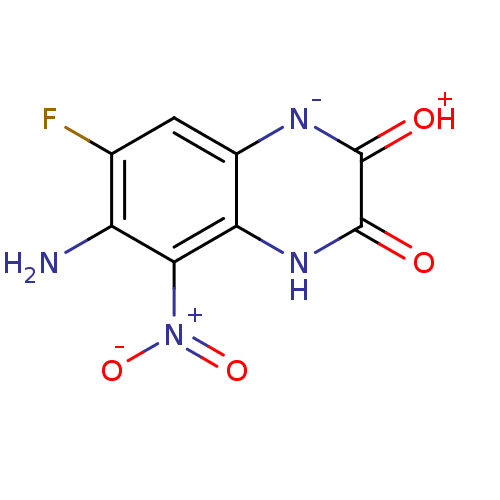
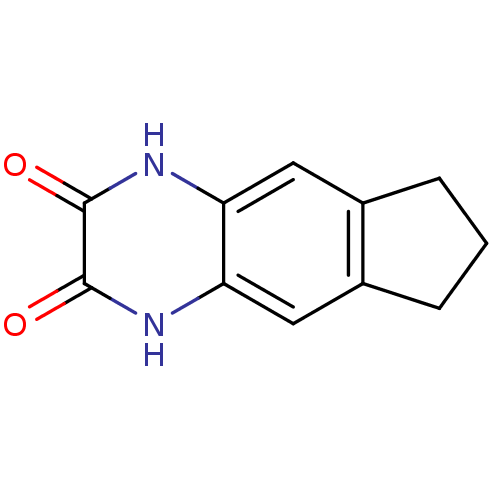
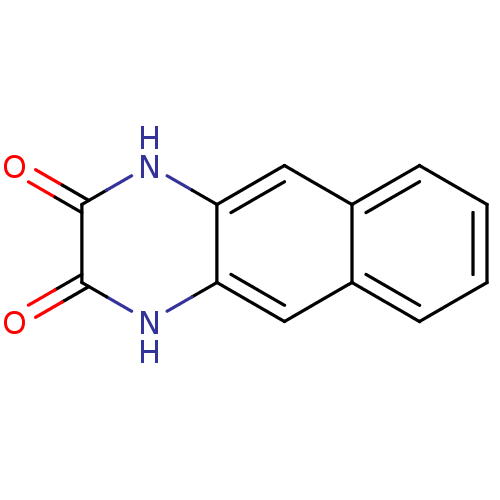
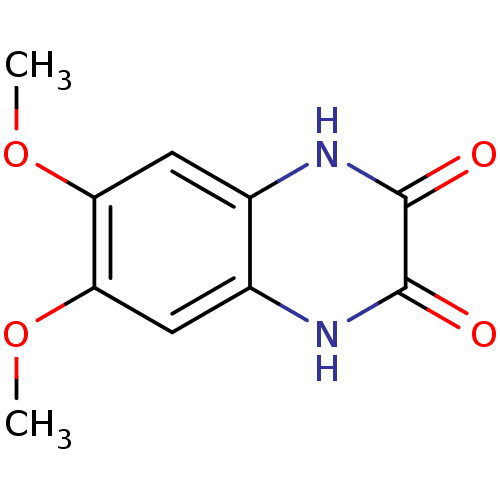
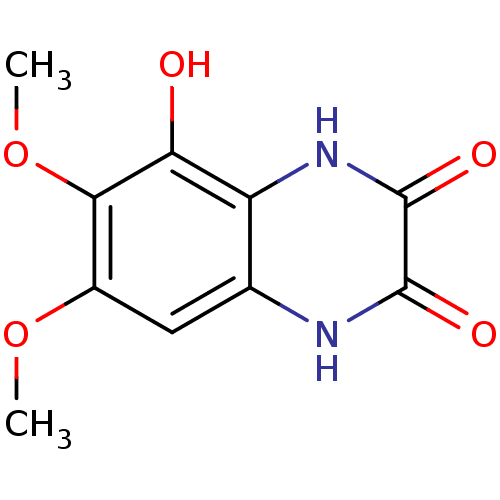
Report error Found 37 Enz. Inhib. hit(s) with all data for entry = 50006995
Affinity DataIC50: 4.70nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.90nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.70nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 45nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 73nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 82nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 95nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 640nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 780nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+3nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+4nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+4nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Potency for the N-methyl-D-aspartate glutamate receptor 1 glycine site by displacement of [3H]5,7-dichlorokynurenic acid (DCKA) binding in rat brain ...More data for this Ligand-Target Pair