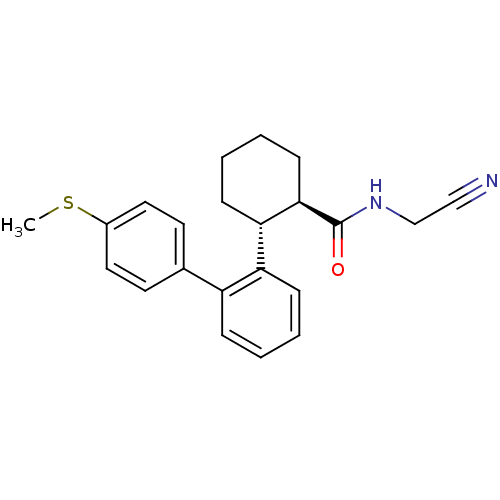
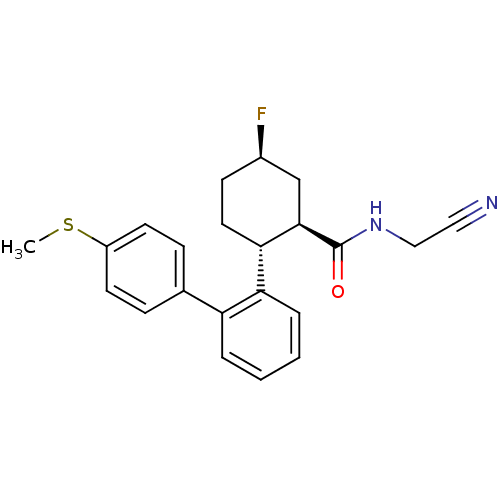
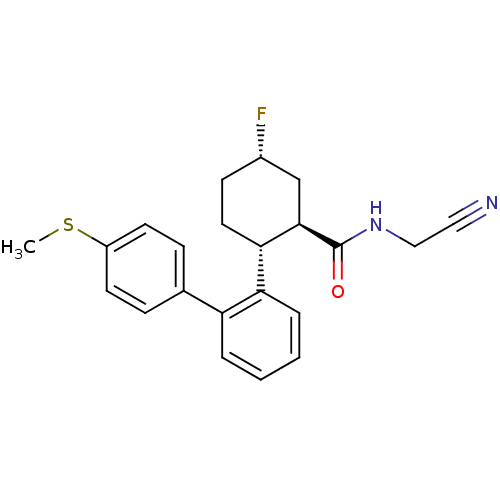
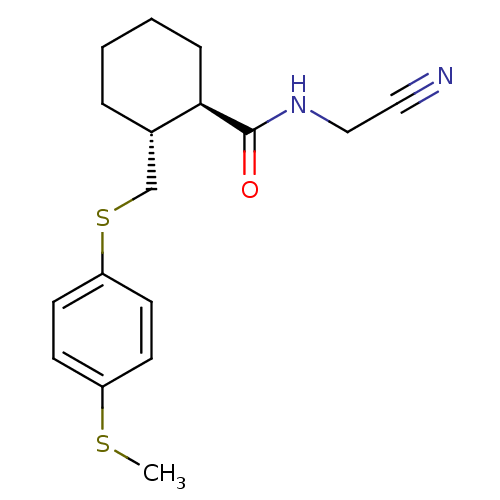
Report error Found 44 Enz. Inhib. hit(s) with all data for entry = 2373
Affinity DataIC50: 0.280nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.320nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.460nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.580nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.590nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.10nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.20nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 17nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 33nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 100nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 172nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 280nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 330nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 350nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 370nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 410nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 460nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 480nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 490nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 700nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 950nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair
Affinity DataIC50: 990nMpH: 5.5 T: 2°CAssay Description:Prior to the addition of substrate, different concentrations of the inhibitor ranging from 100 uM to 0.2 nM were preincubated for 15 min with enzyme ...More data for this Ligand-Target Pair