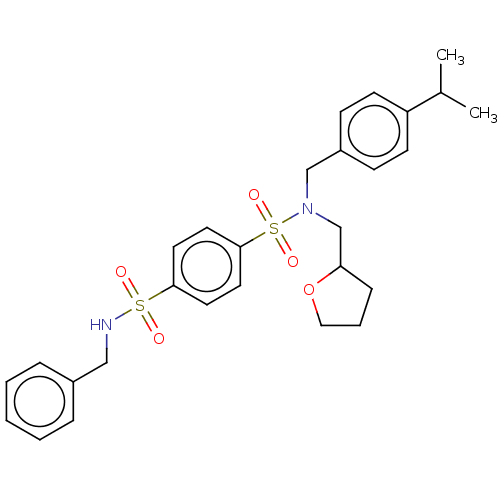
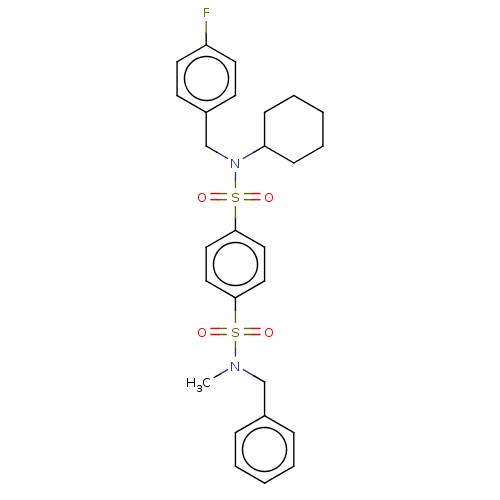
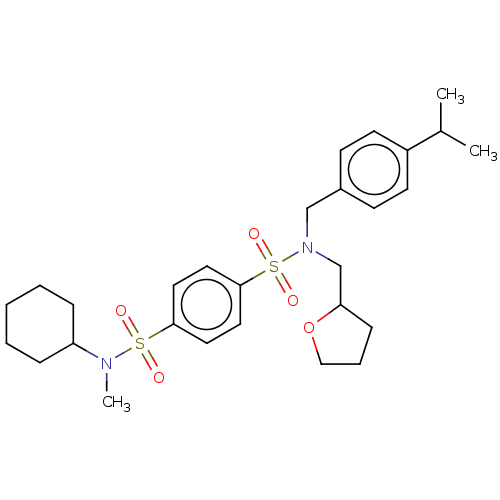
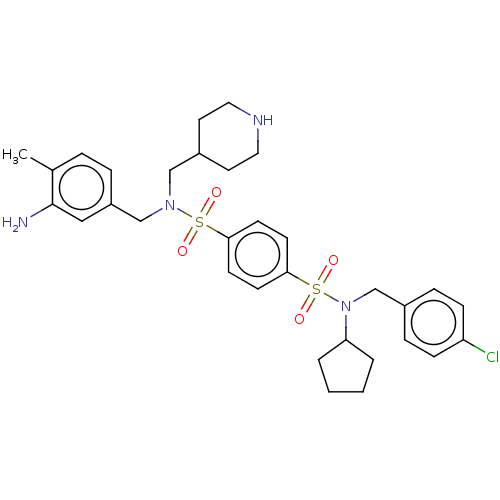
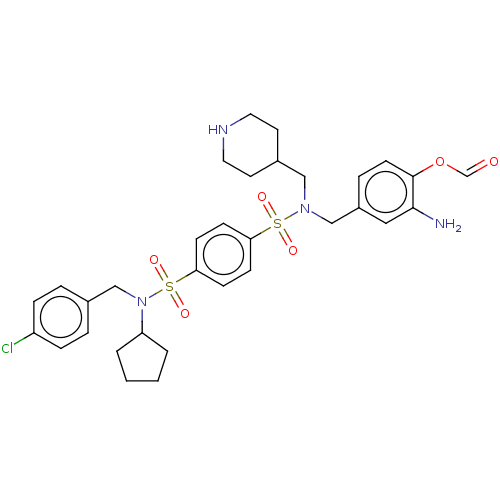
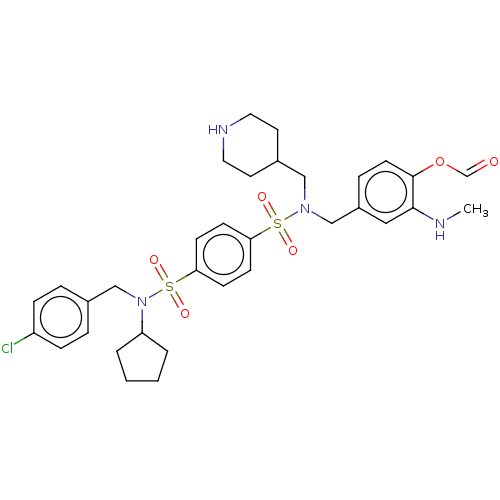
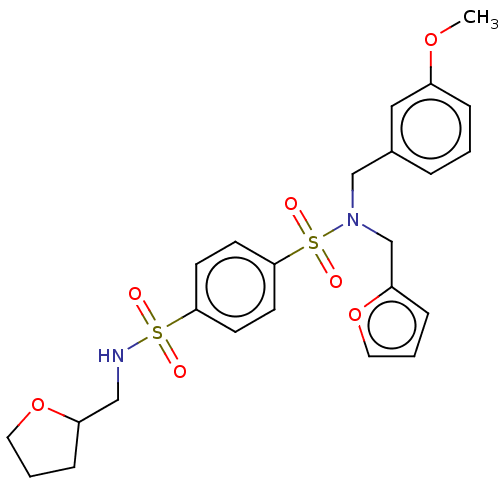
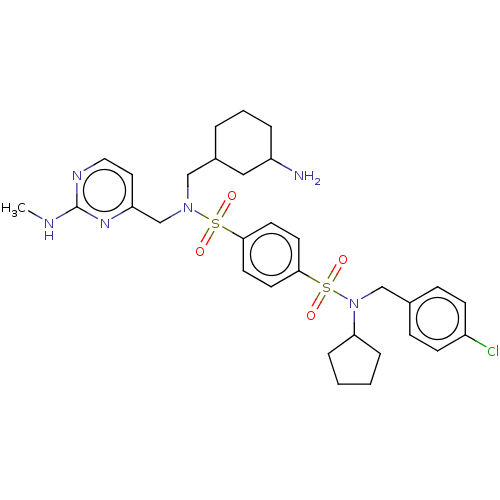
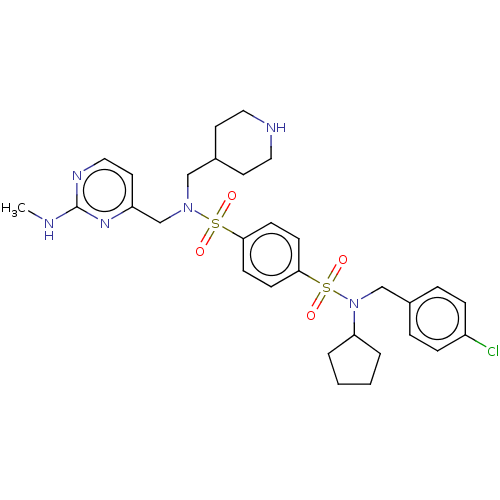
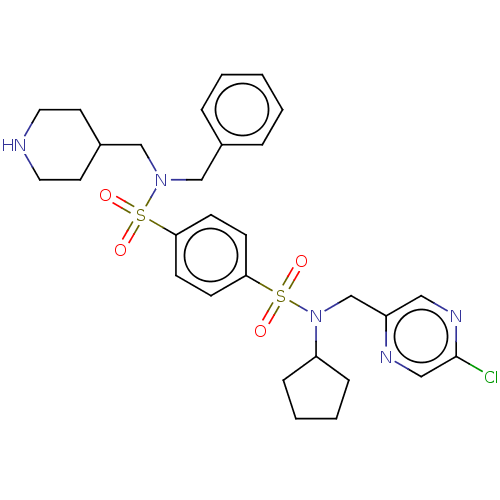
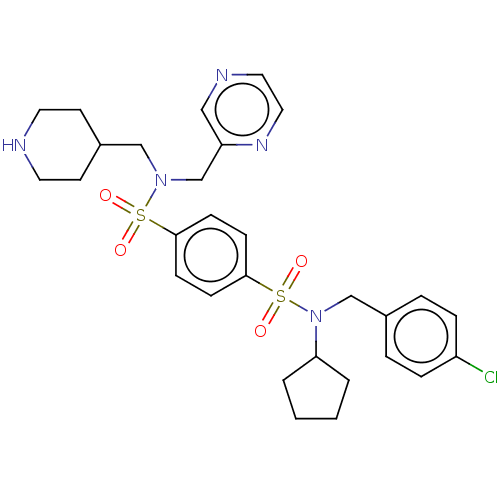
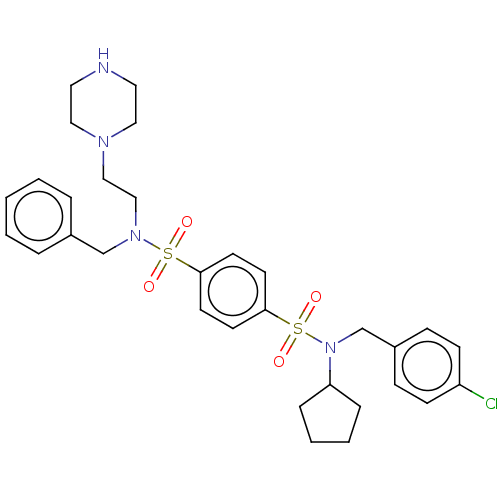
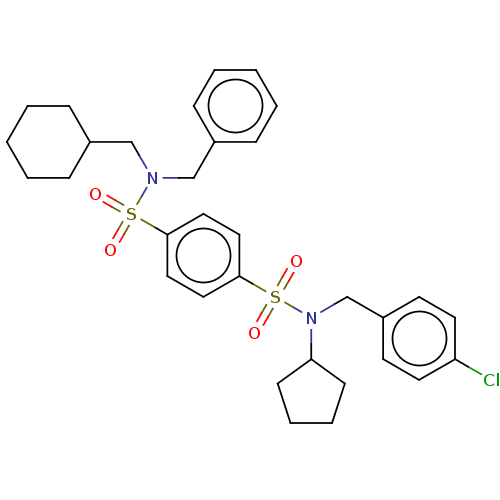
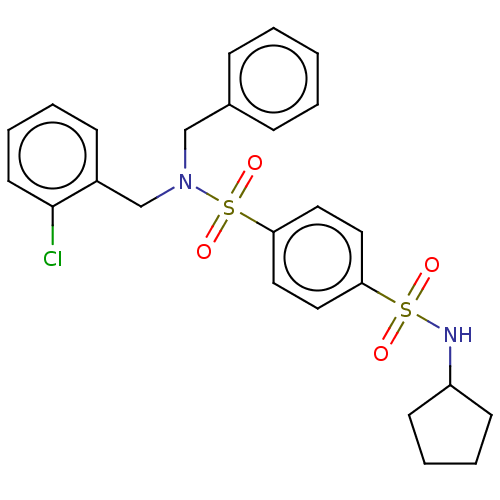
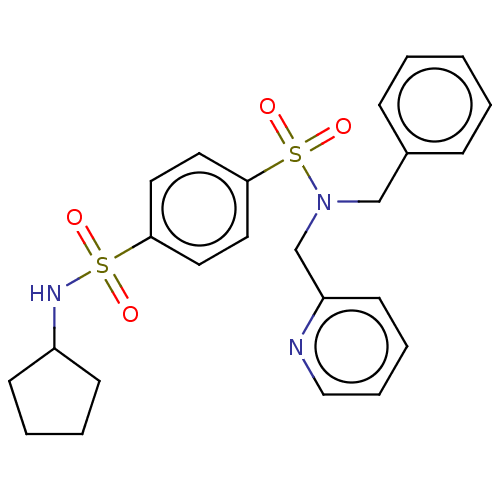
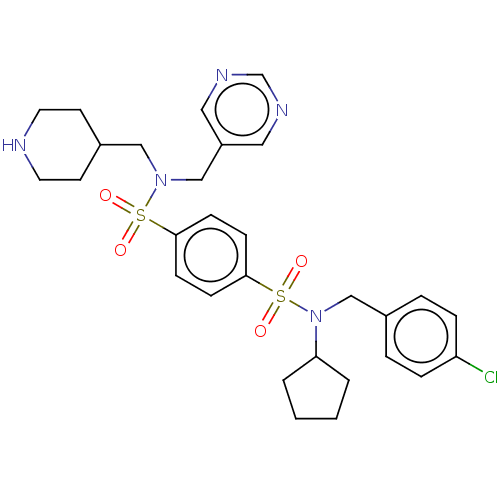
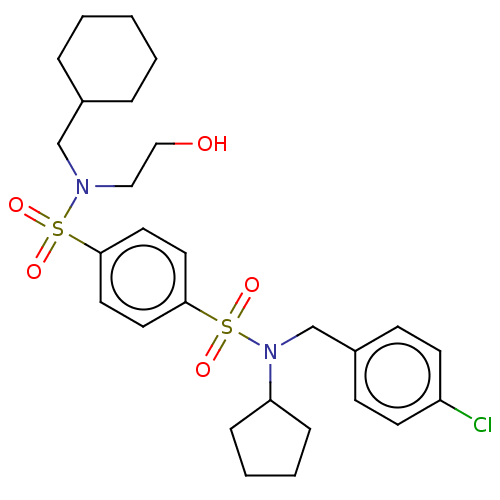
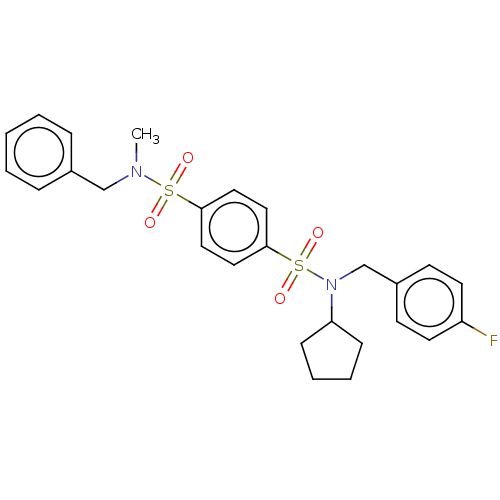
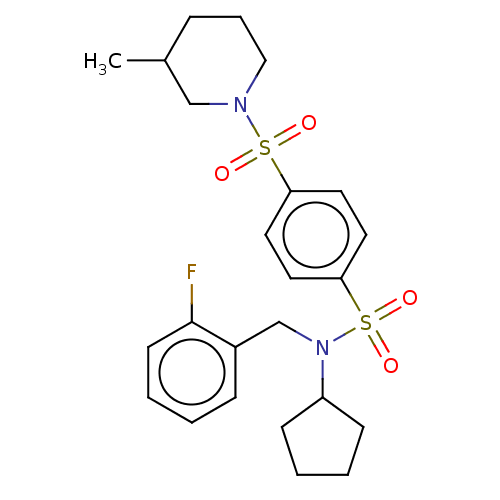
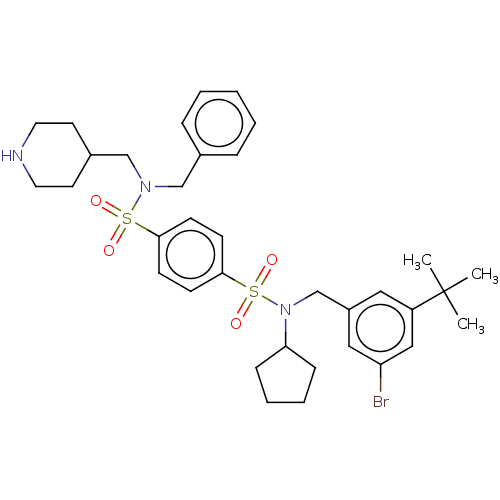
Report error Found 100 Enz. Inhib. hit(s) with all data for entry = 9684
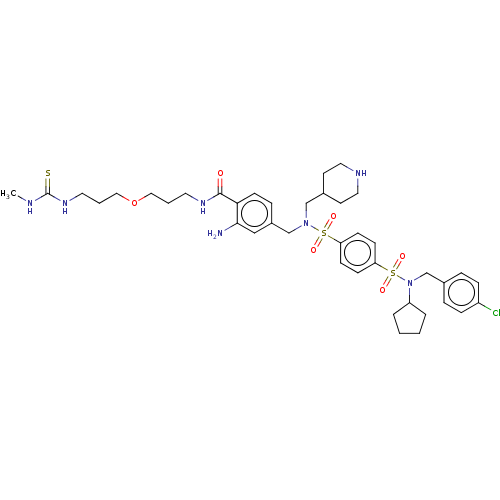
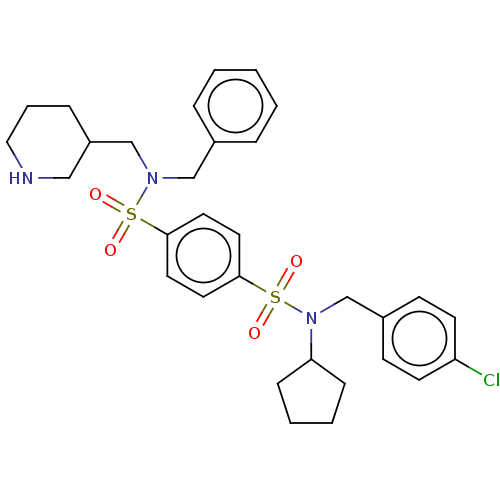
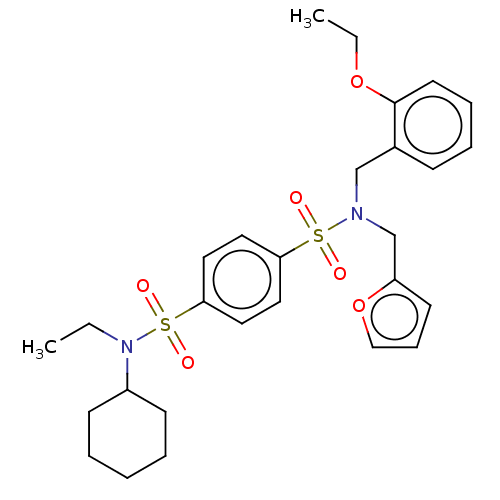
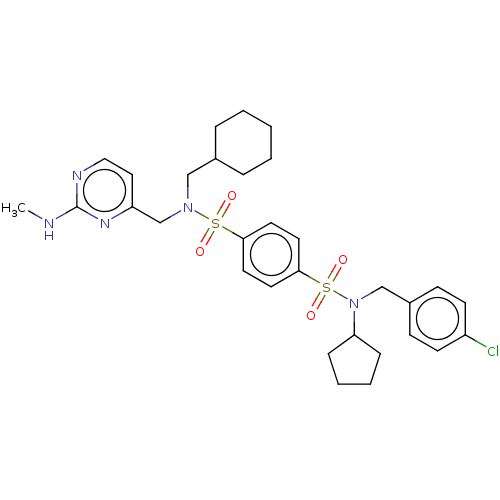
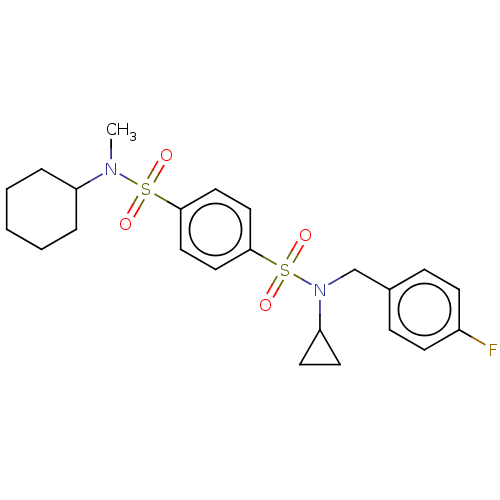
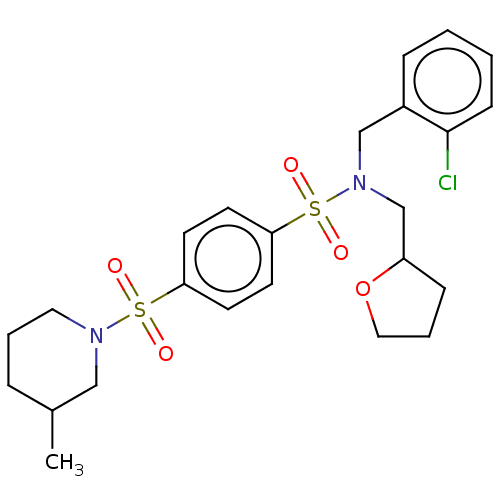
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
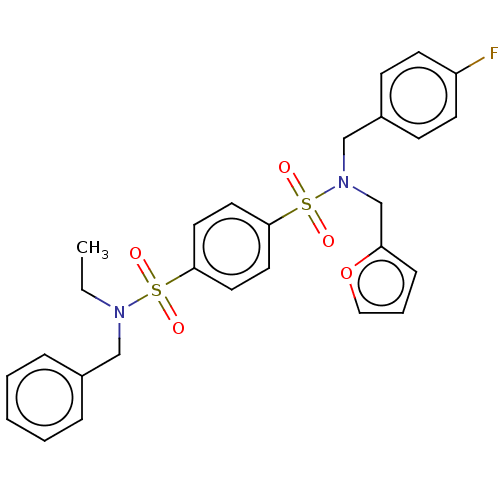
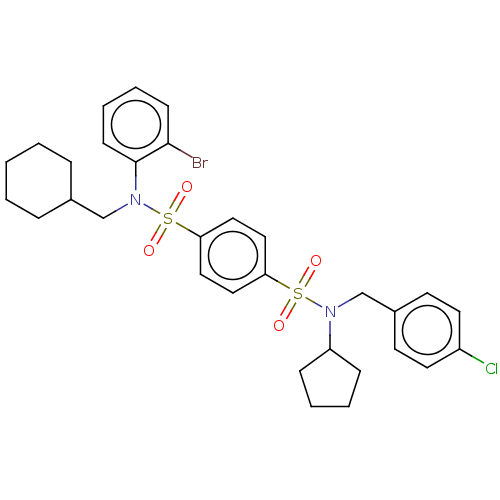
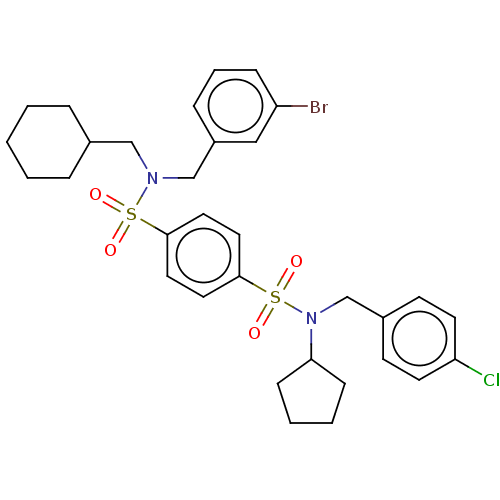
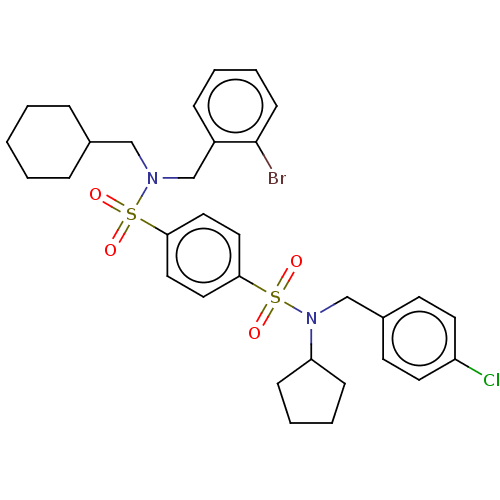
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 16nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
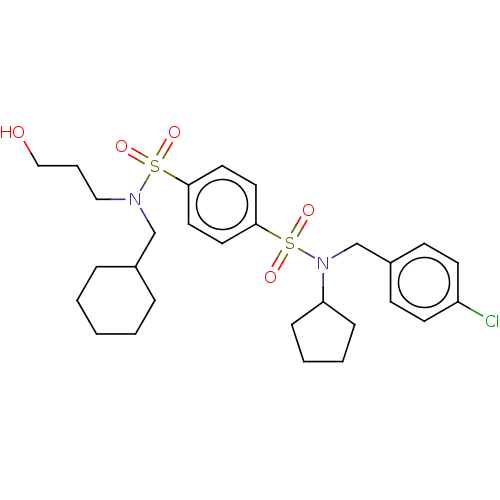
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
TargetRetinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta(Human)
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Max-Planck-Gesellschaft Zur Forderung Der Wissenschaften
US Patent
Affinity DataKd: 65nMAssay Description:Binding to PDEδ was validated and quantified by means of a displacement assay employing a fluorescent-tagged analog of the HMG-CoA reductase inh...More data for this Ligand-Target Pair
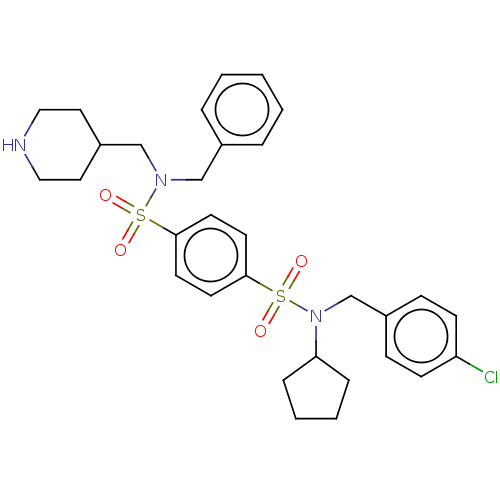
 3D Structure (crystal)
3D Structure (crystal)