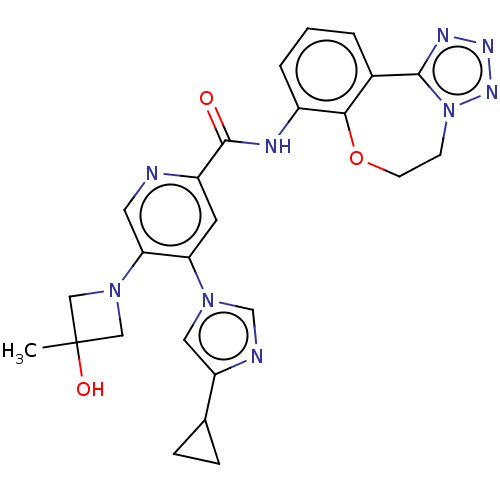
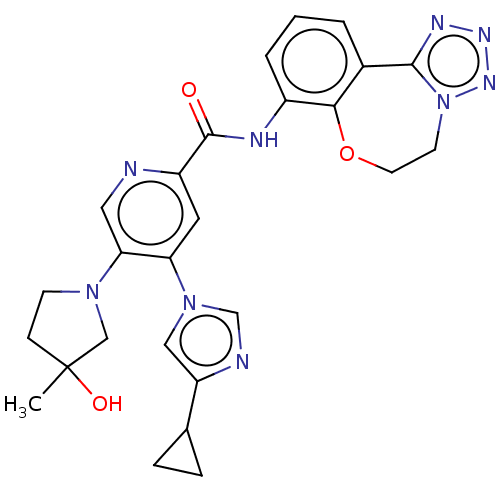
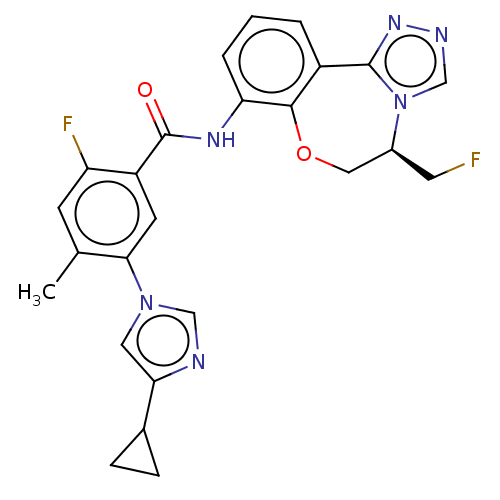
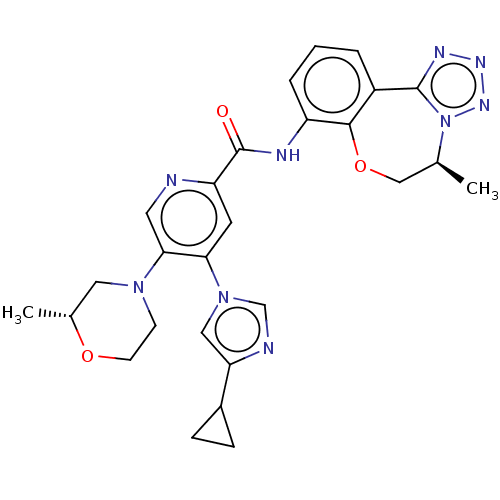
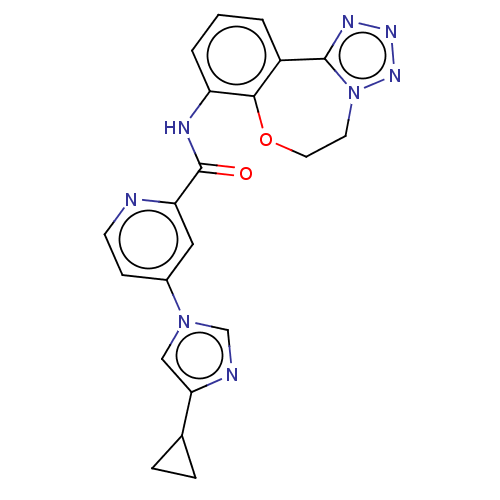
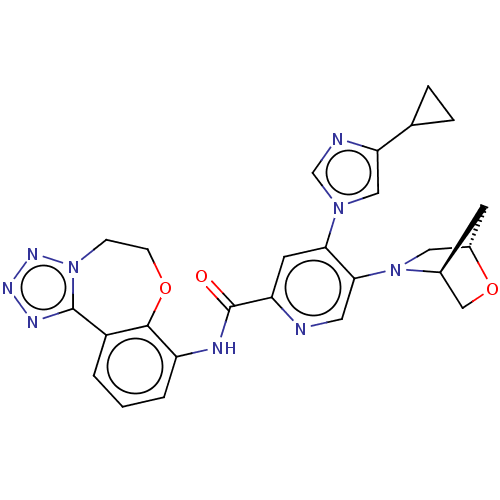
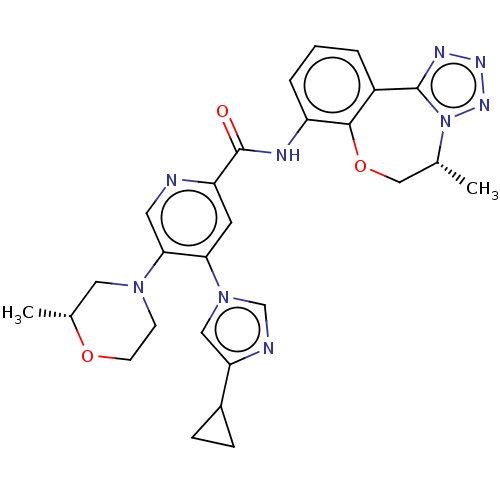
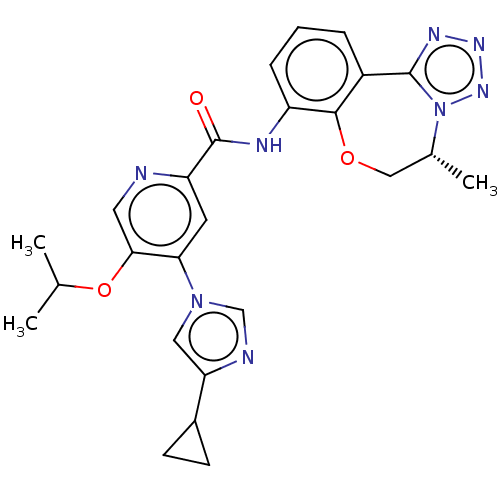
Report error Found 81 Enz. Inhib. hit(s) with all data for entry = 9711
Affinity DataIC50: 0.5nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.900nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.5nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.5nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.5nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.60nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:Base Reaction buffer: 20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM DTT, and 1% DMSO were used for ...More data for this Ligand-Target Pair