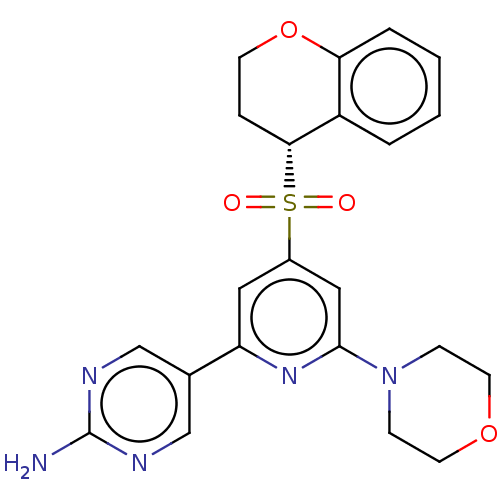
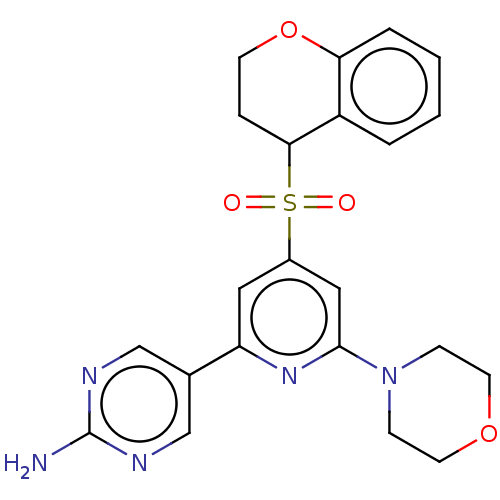
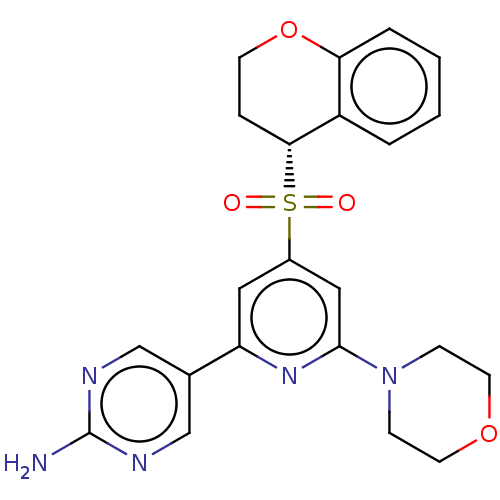
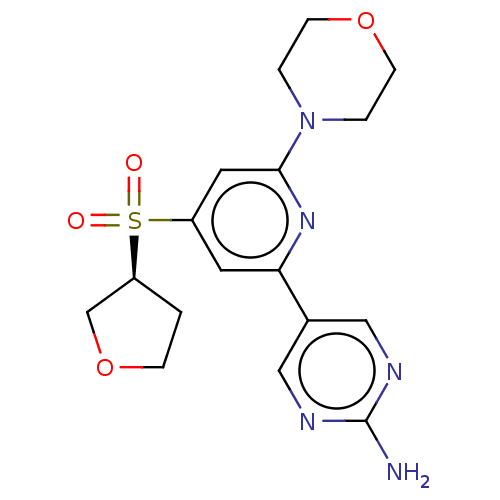
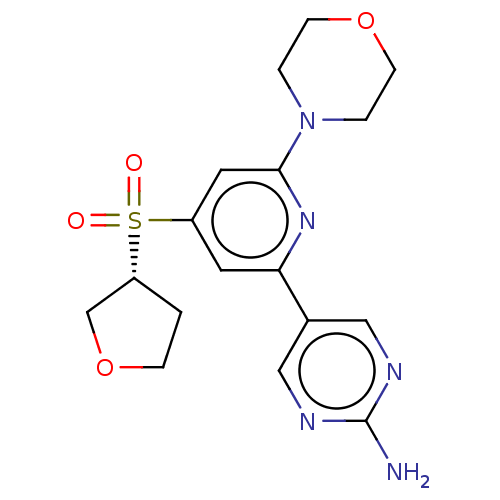
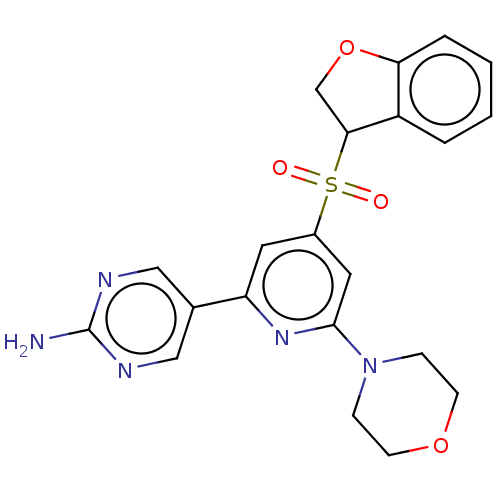
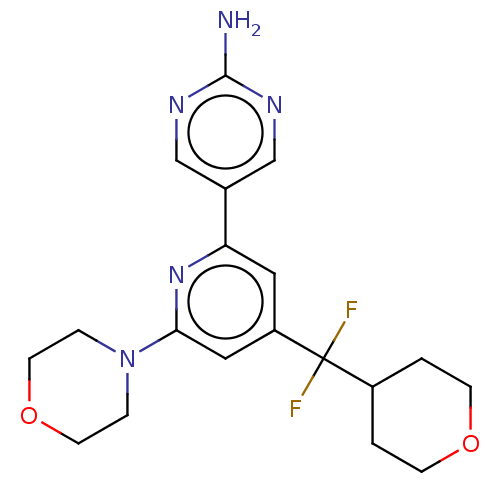
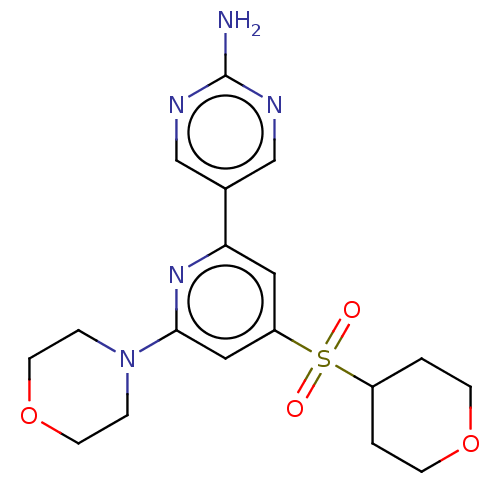
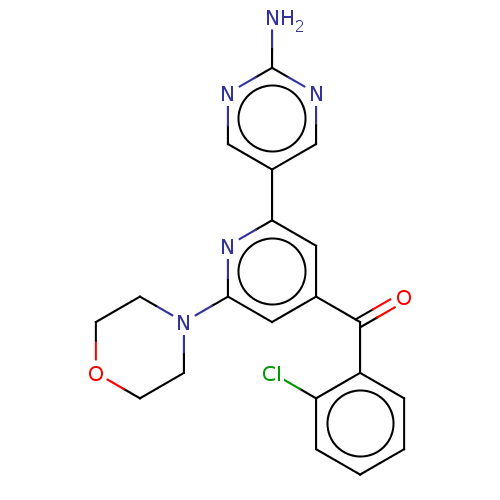
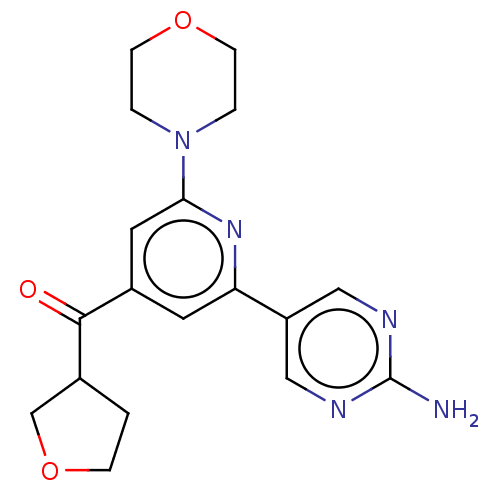
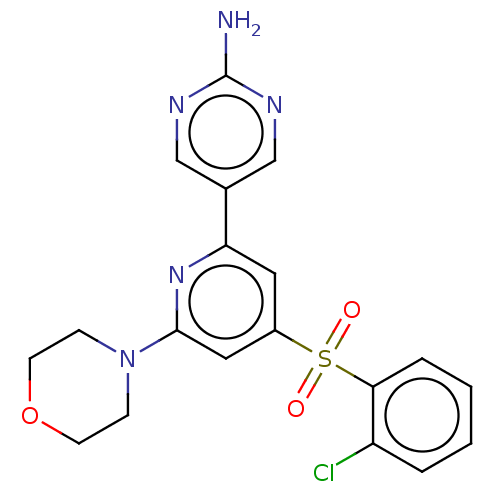
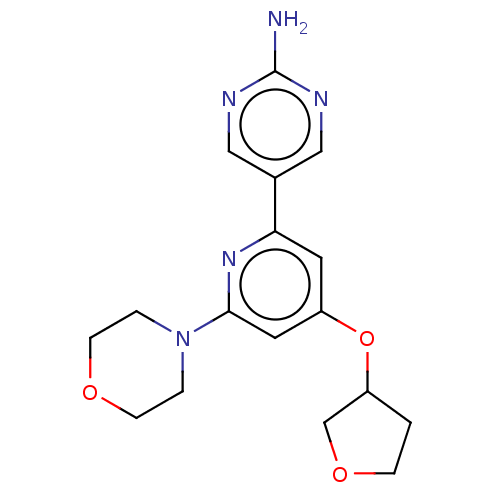
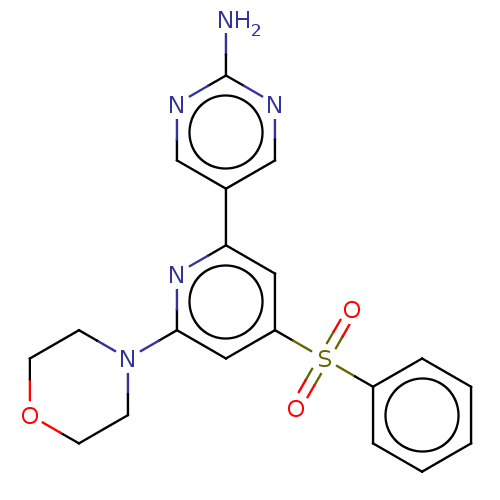
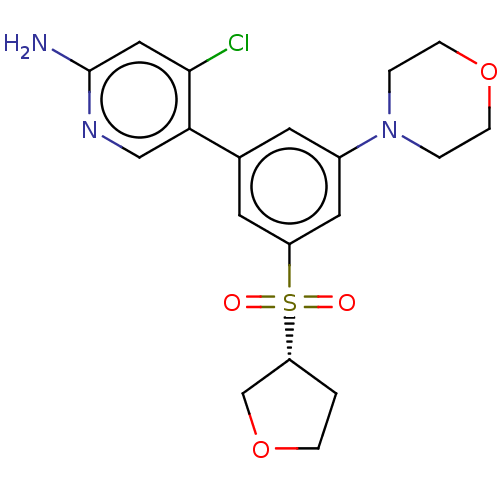
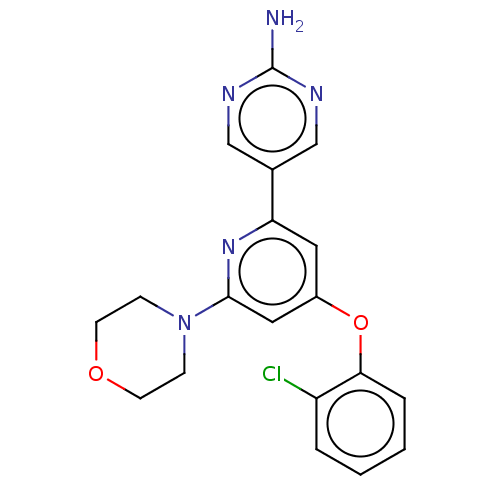
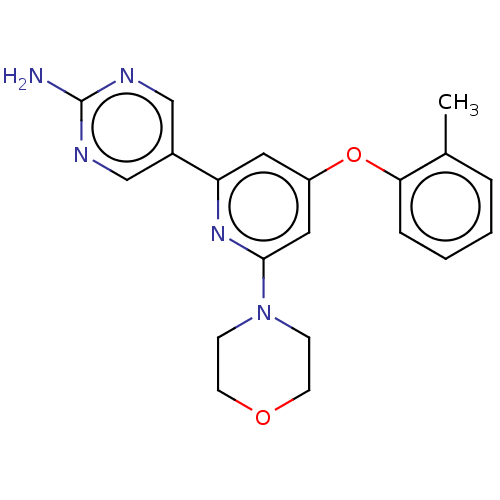
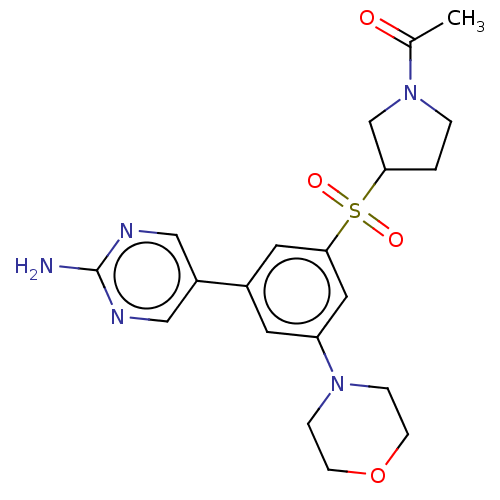
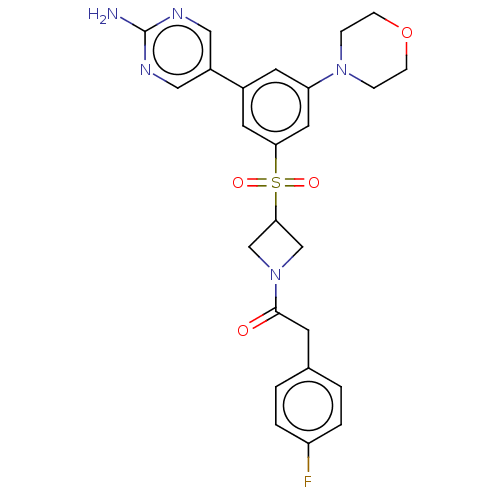
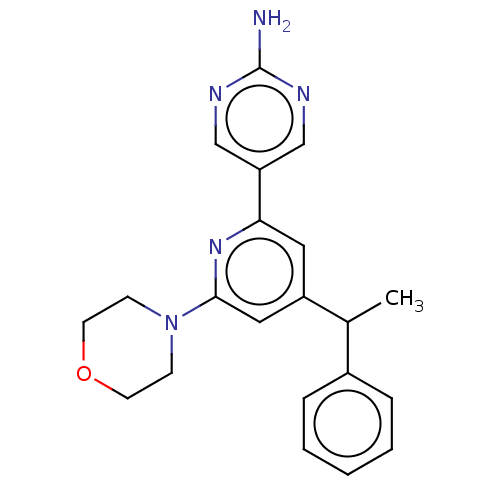
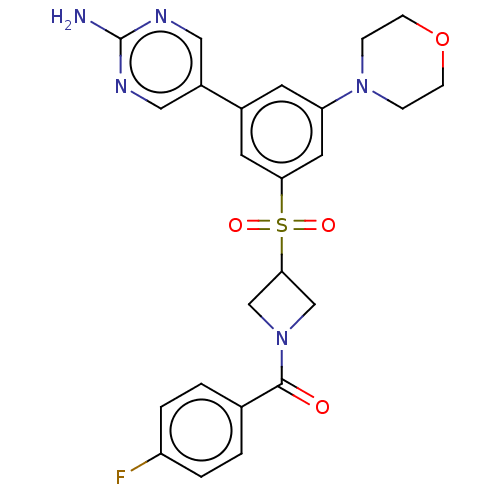
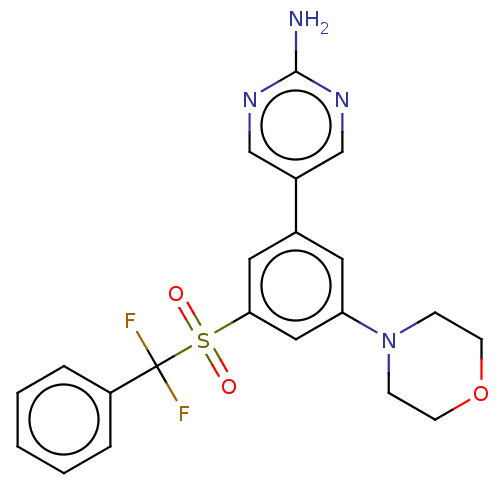
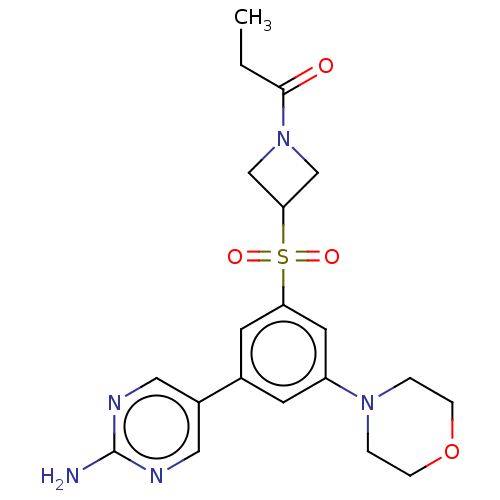
Report error Found 311 Enz. Inhib. hit(s) with all data for entry = 10952
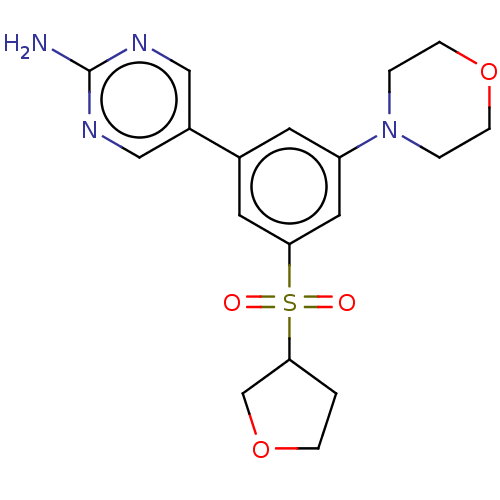
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
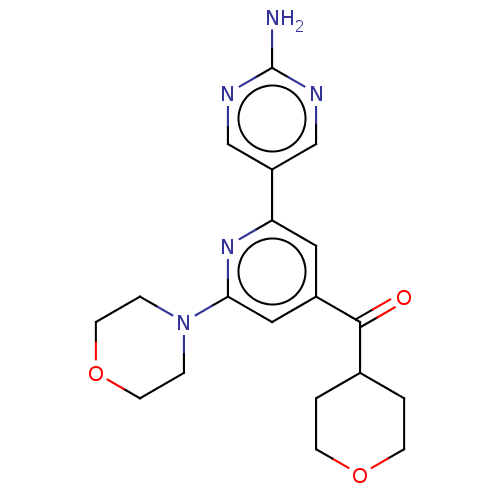
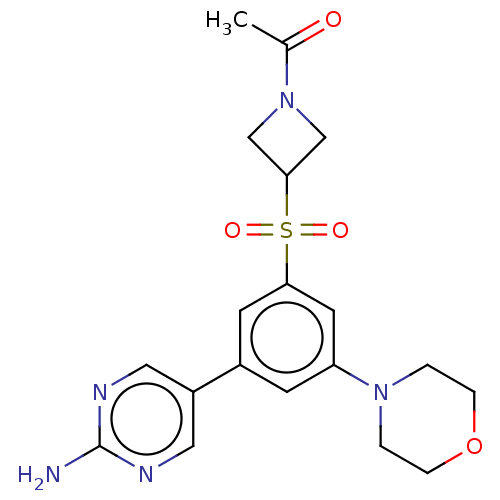
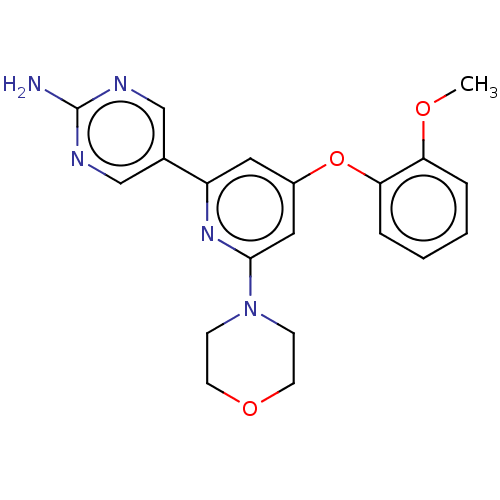
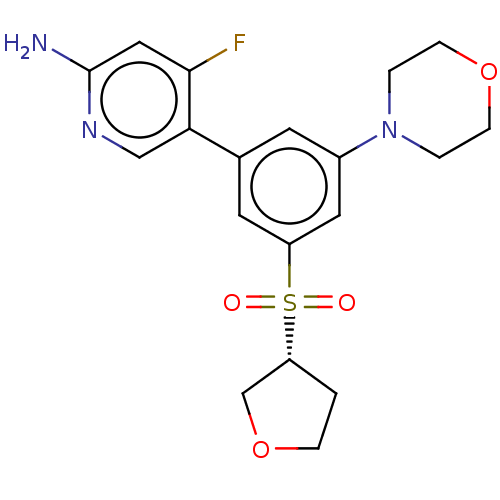
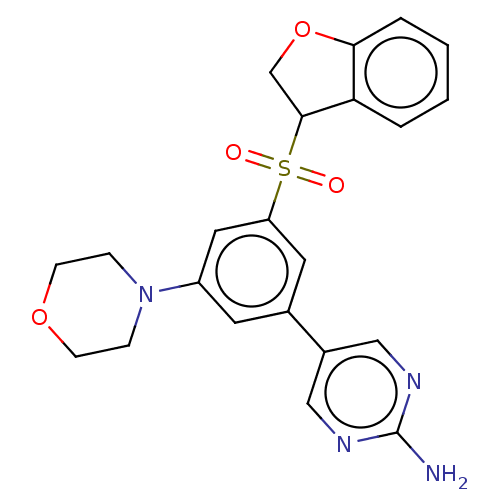
Affinity DataIC50: 1nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
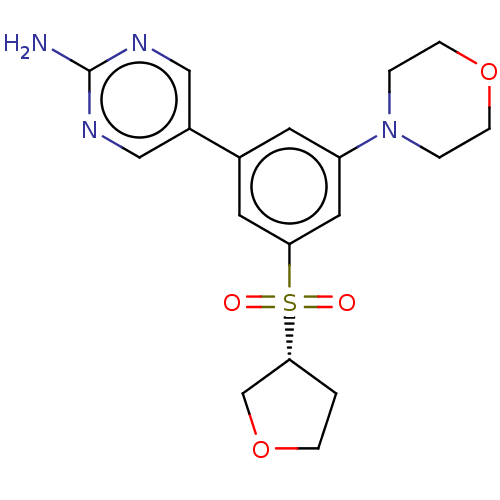
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
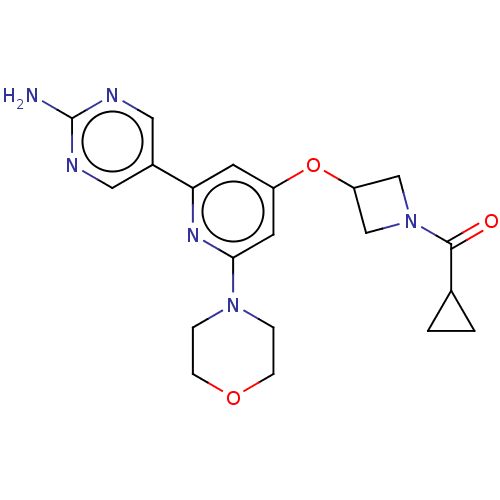
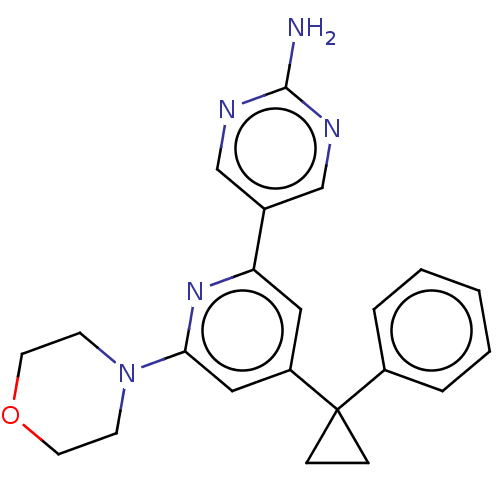
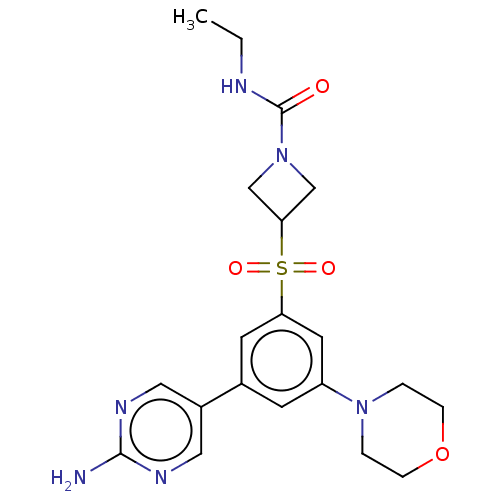
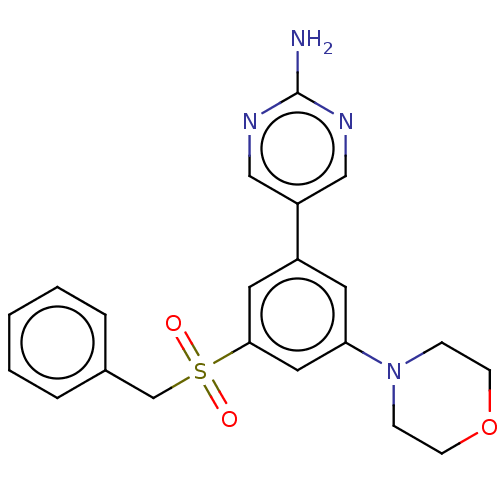
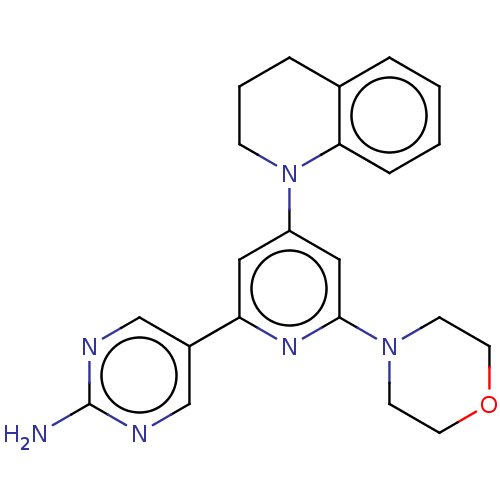
Affinity DataIC50: 1.70nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
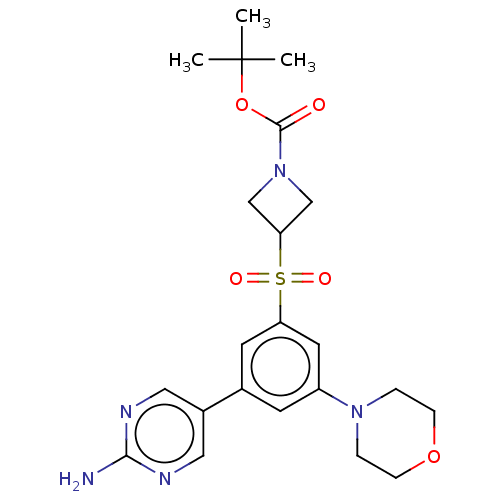
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
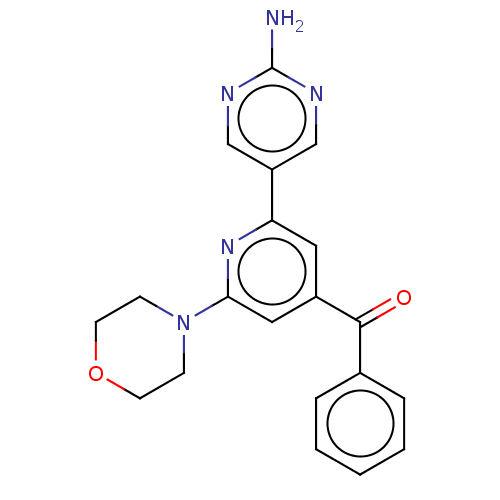
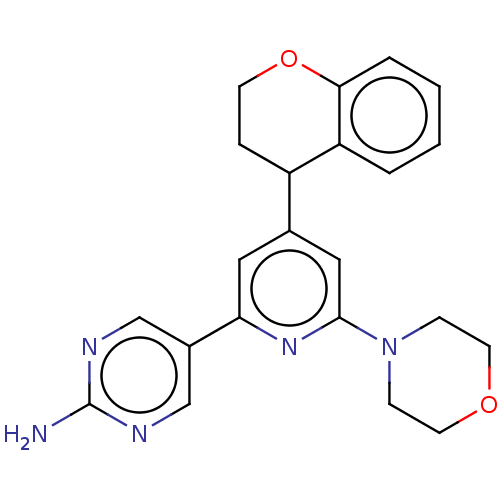
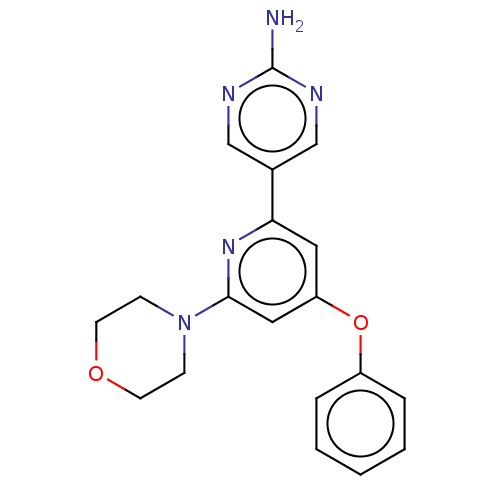
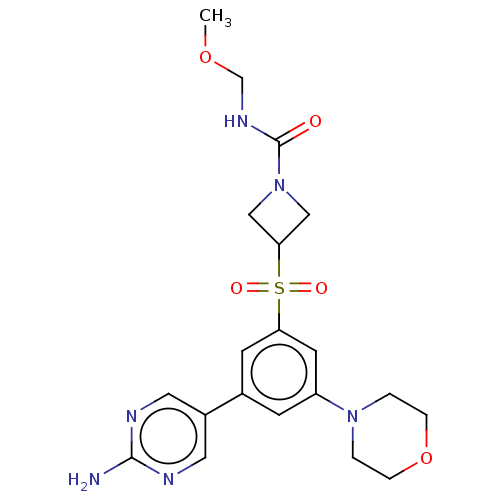
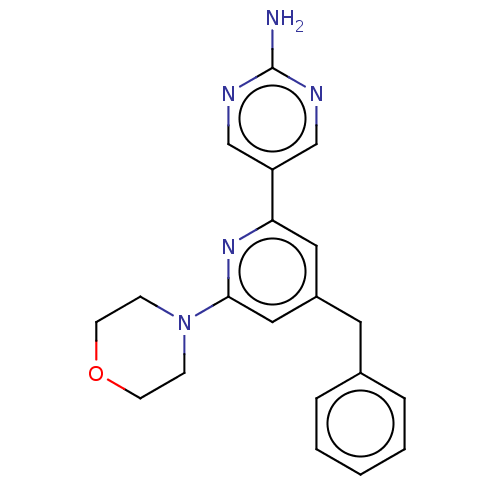
Affinity DataIC50: 4.5nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
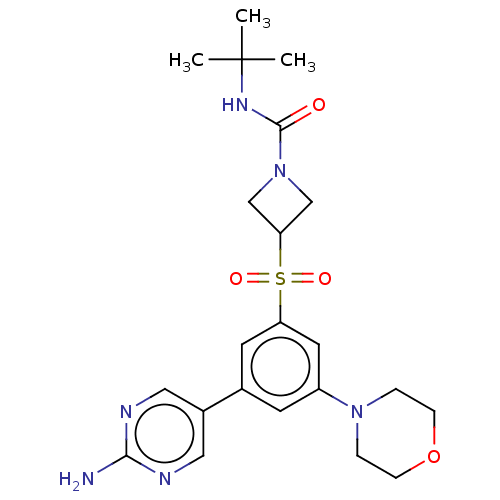
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
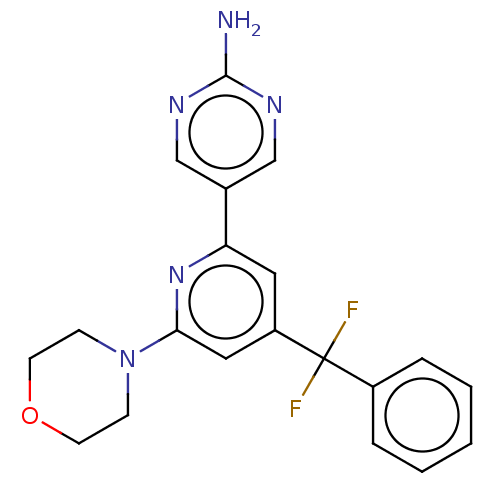
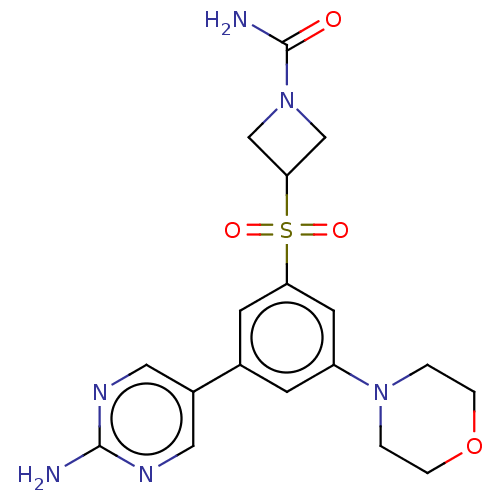
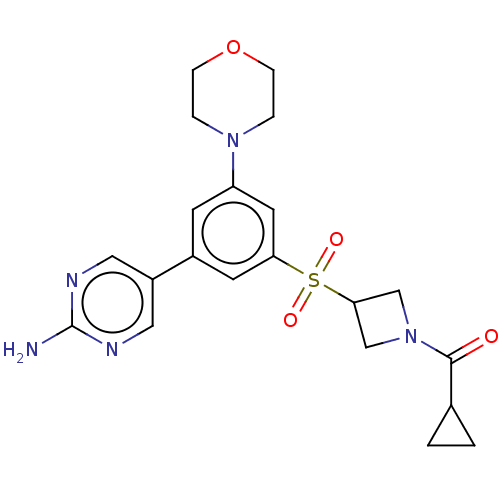
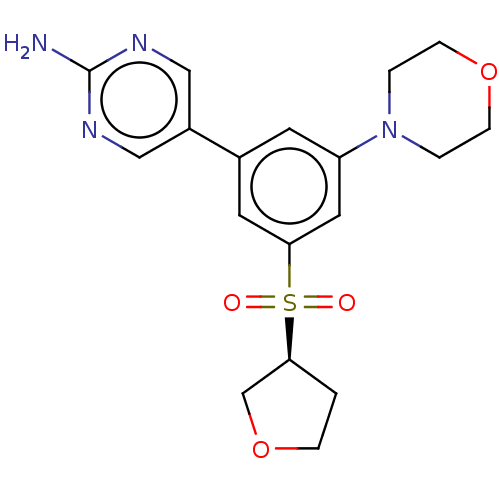
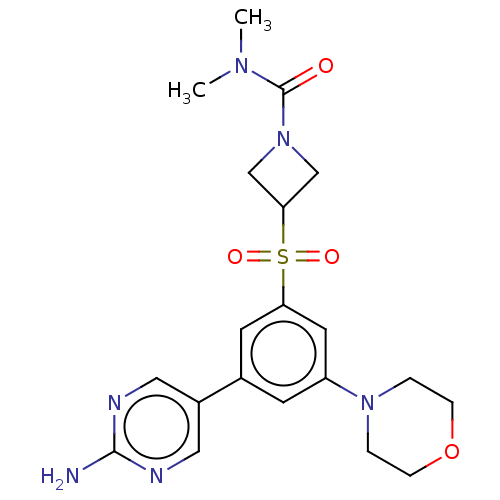
Affinity DataIC50: 5.70nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 6.80nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 7nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 12nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 13nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 14nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 14nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of the PI3K-AKT-mTOR pathway was measured by quantifying the loss of (Ser-473) pAKT using AlphaScreen (Perkin Elmer). B103 (Rat Neuroblast...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 17nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 23nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 23nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 34nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 37nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 46nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 47nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 56nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 62nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 62nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:The substrate was prepared in base reaction buffer (20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM D...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 66nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 67nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 70nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 75nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 79nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 88nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 91nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 100nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 100nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 100nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 110nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 110nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 110nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 120nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 120nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 120nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 120nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 130nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:The substrate was prepared in base reaction buffer (20 mM Hepes (pH 7.5), 10 mM MgCl2, 1 mM EGTA, 0.02% Brij35, 0.02 mg/ml BSA, 0.1 mM Na3VO4, 2 mM D...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 130nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 150nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 180nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 180nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:Inhibition of the PI3K-AKT-mTOR pathway was measured by quantifying the loss of (Ser-473) pAKT using AlphaScreen (Perkin Elmer). B103 (Rat Neuroblast...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 210nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 210nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 210nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Neuropore Therapies
US Patent
Neuropore Therapies
US Patent
Affinity DataIC50: 250nMAssay Description:Quantification of ATP to ADP conversion as a measure of PI3Kα activity. Active PI3Kα (Life Technologies), in the presence or absence of PI3...More data for this Ligand-Target Pair