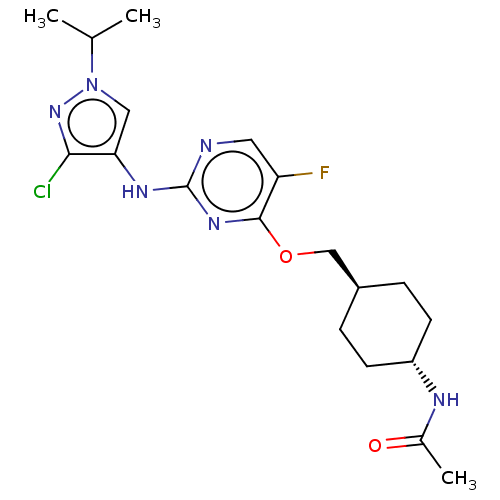
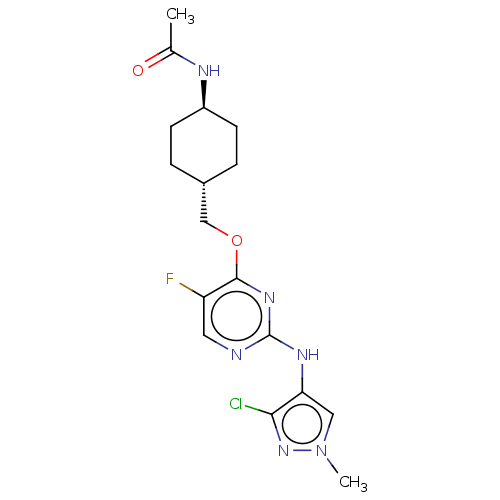
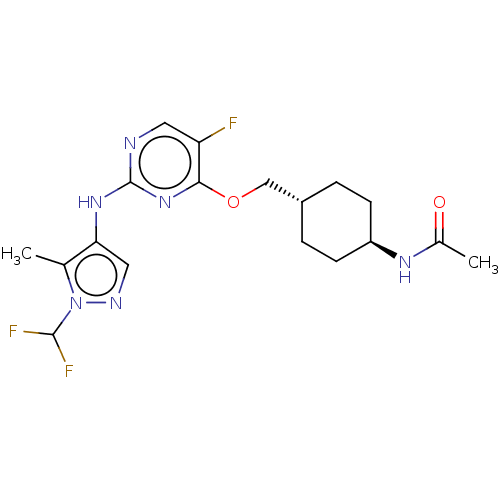
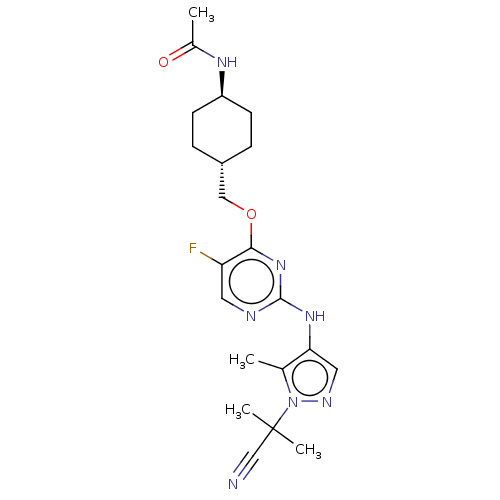
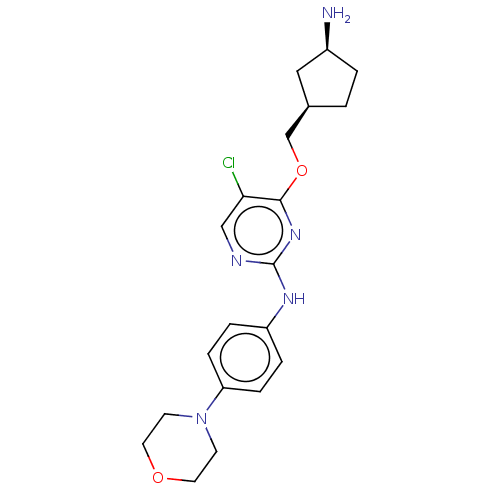
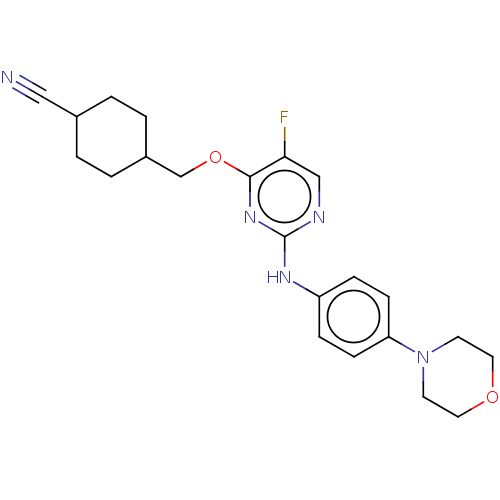
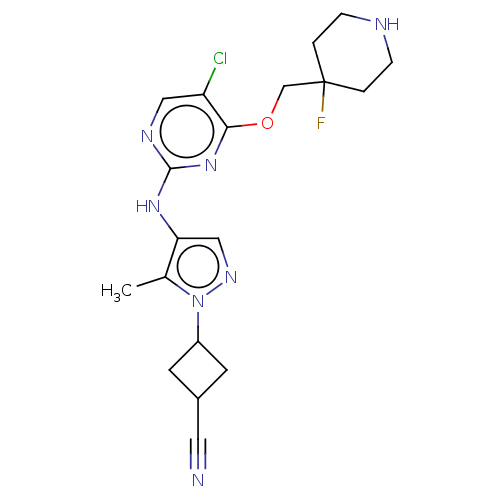
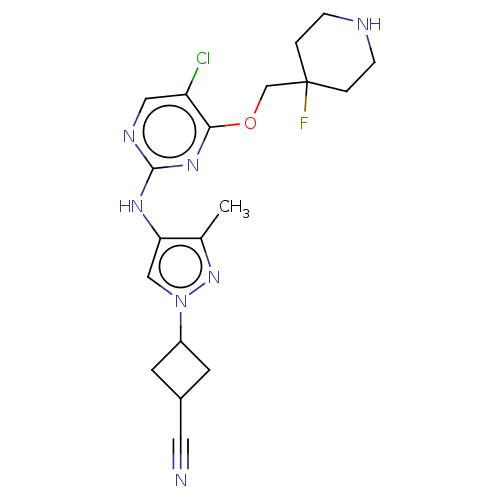
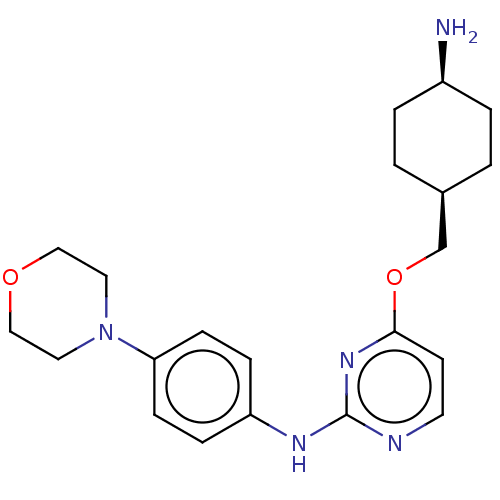
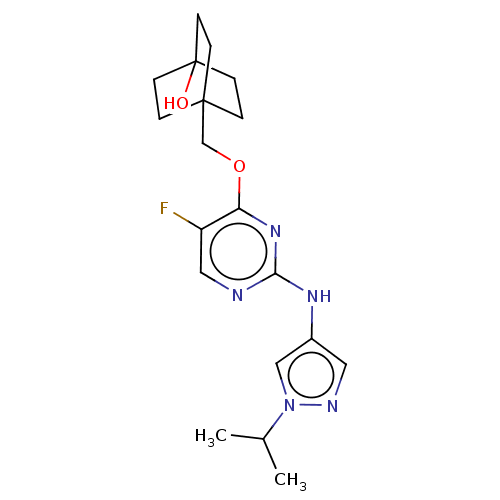
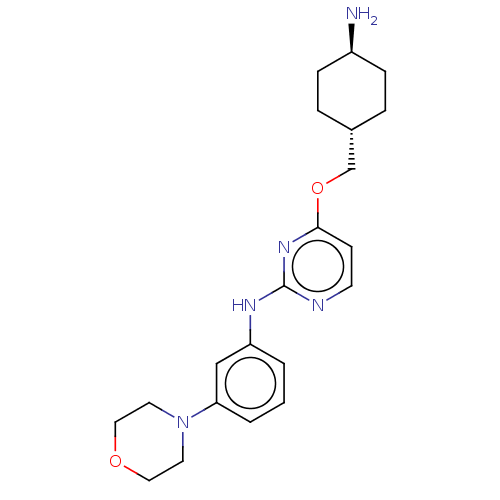
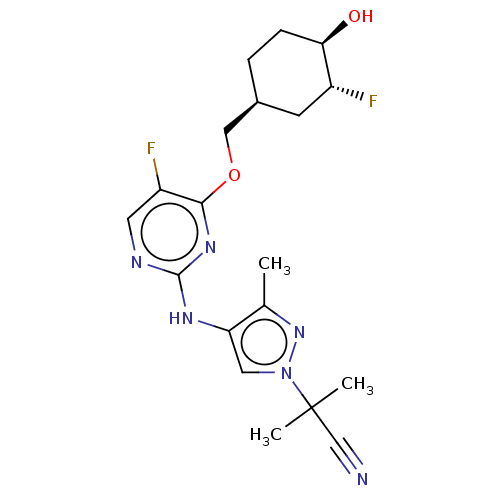
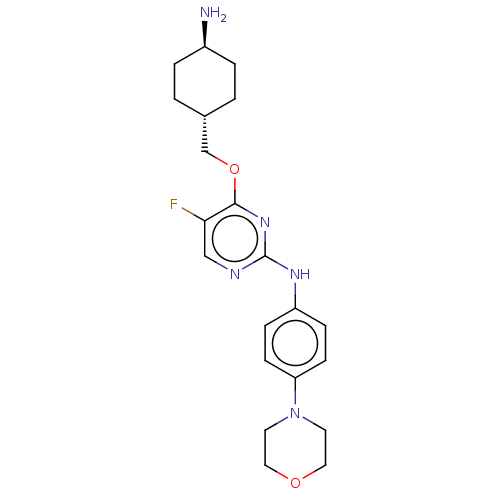
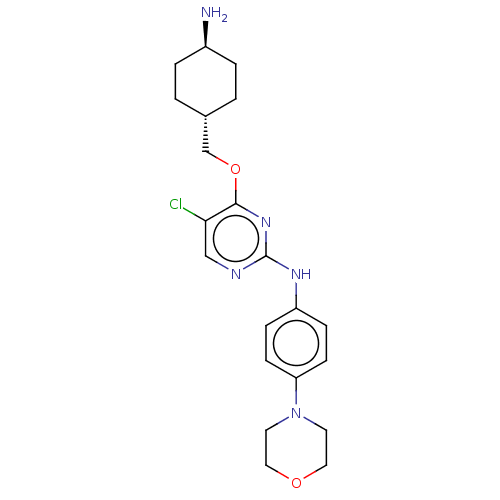
Report error Found 290 Enz. Inhib. hit(s) with all data for entry = 11128
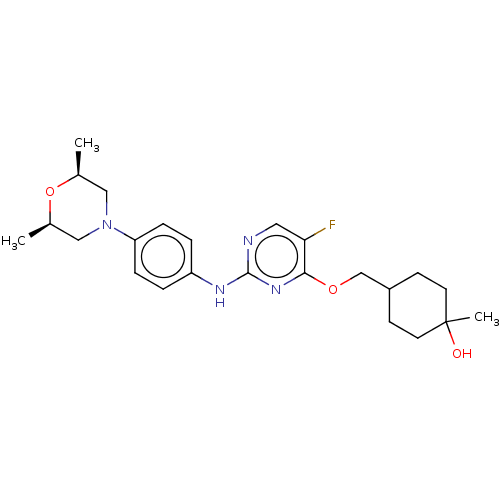
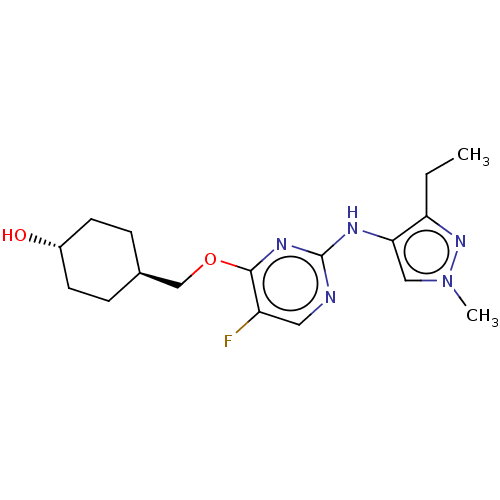
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
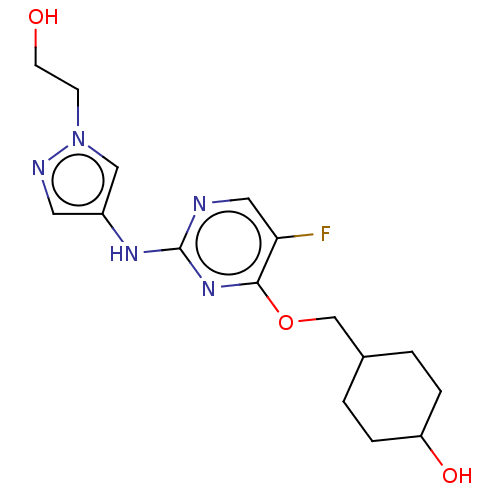
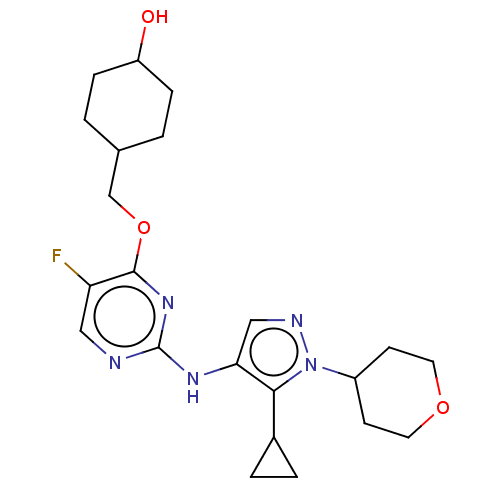
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
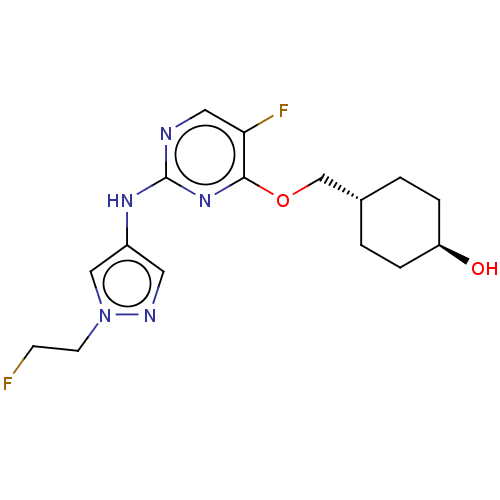
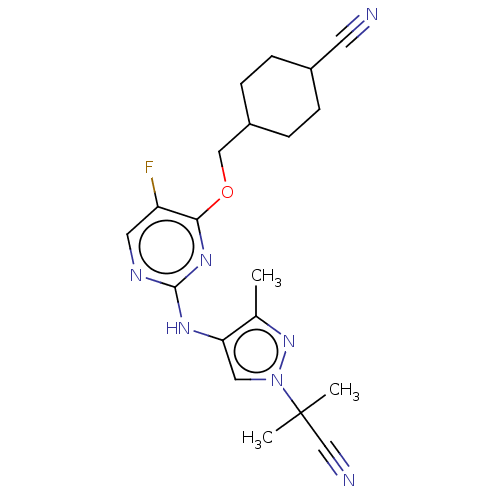
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
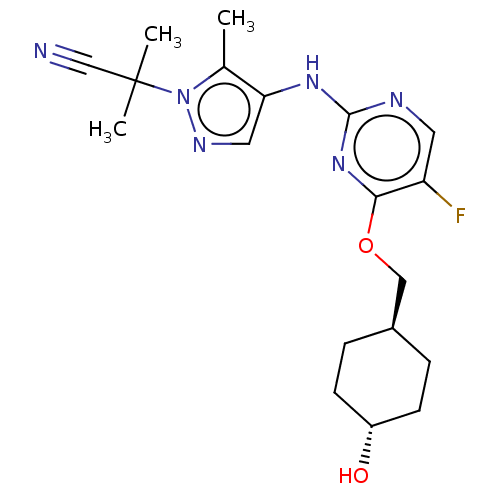
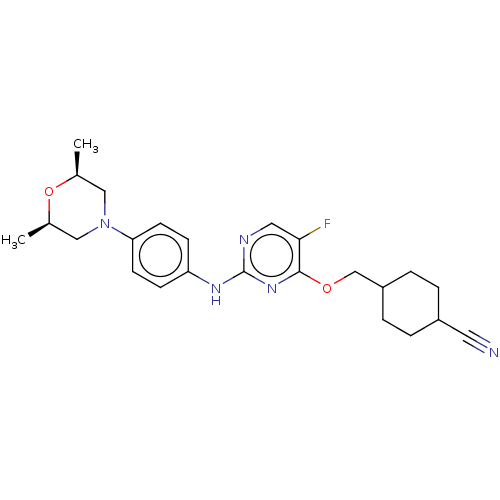
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2 [970-2527](Human)
Halia Therapeutics
US Patent
Halia Therapeutics
US Patent
Affinity DataIC50: 50nMAssay Description:LRRKtide substrate (peptide sequence RLGRDKYKTLRQIRQ, derived from human ezrin [amino acids 561-573], moesin [amino acids 539-553] and radixin [amino...More data for this Ligand-Target Pair