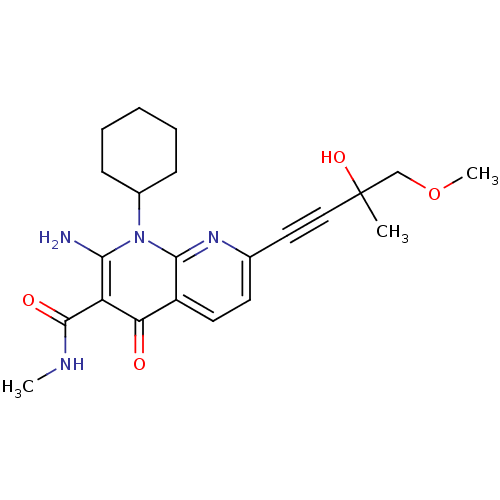
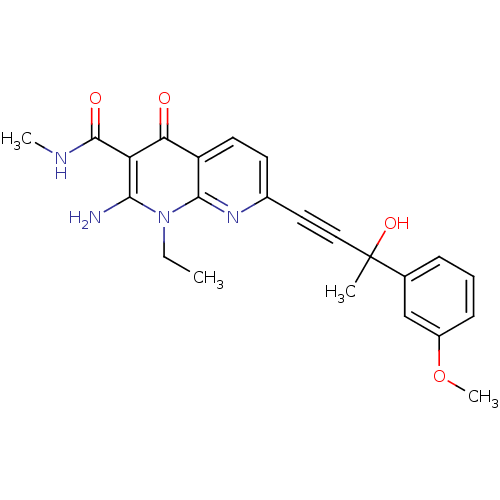
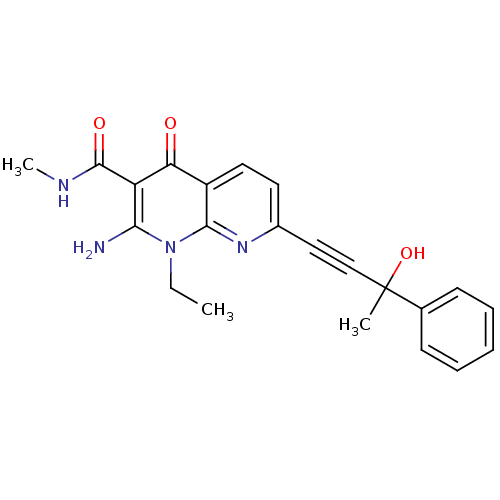
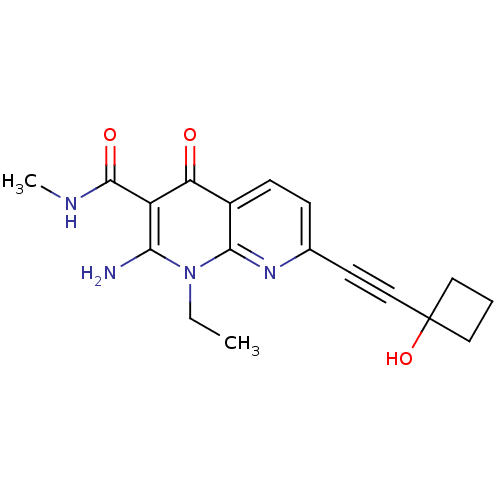
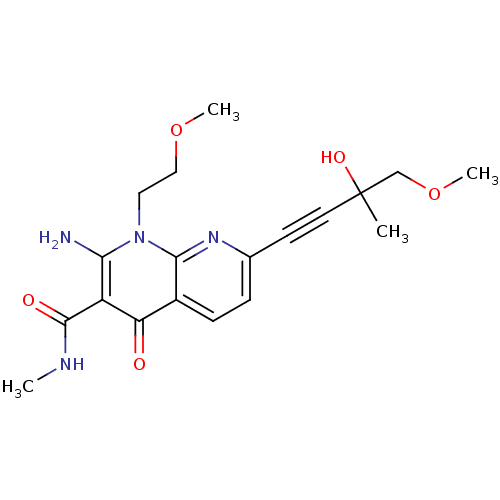
Report error Found 15 Enz. Inhib. hit(s) with all data for entry = 5888
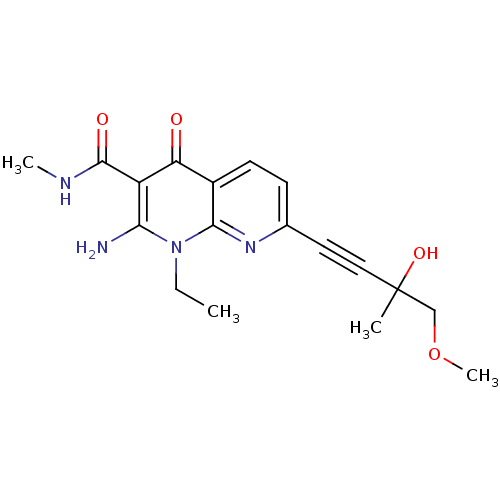
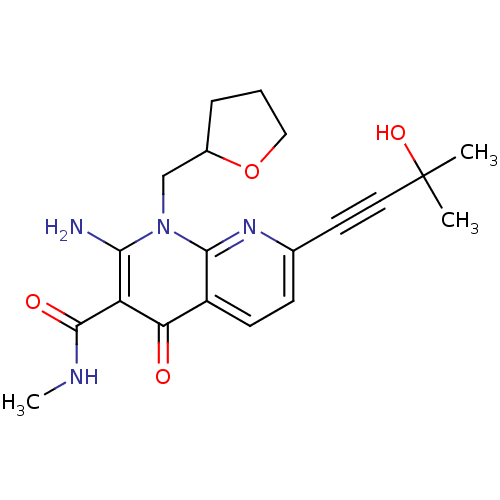
Affinity DataIC50: 2nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
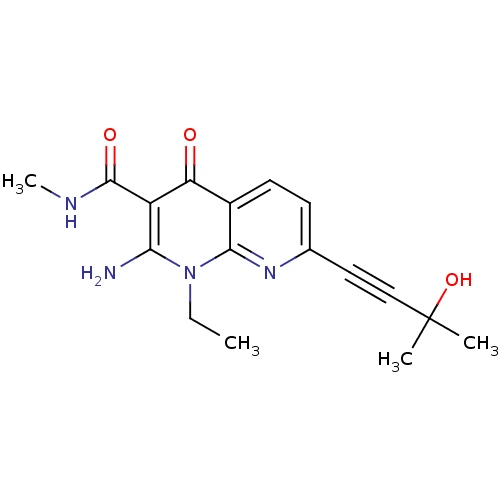
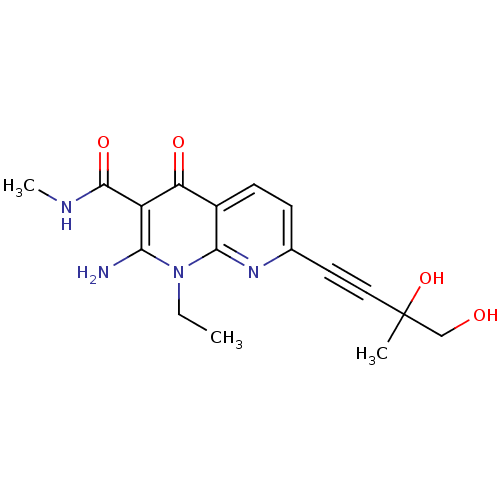
Affinity DataIC50: 5nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
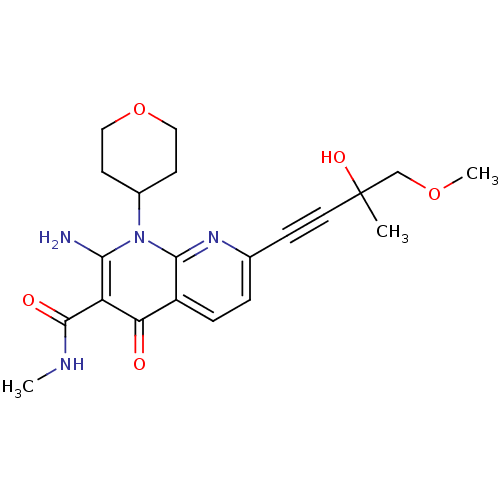
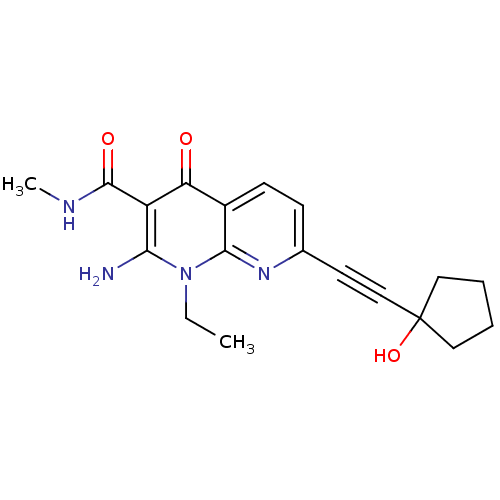
Affinity DataIC50: 6nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
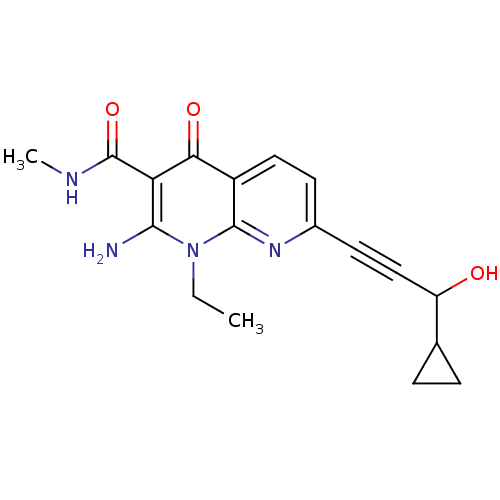
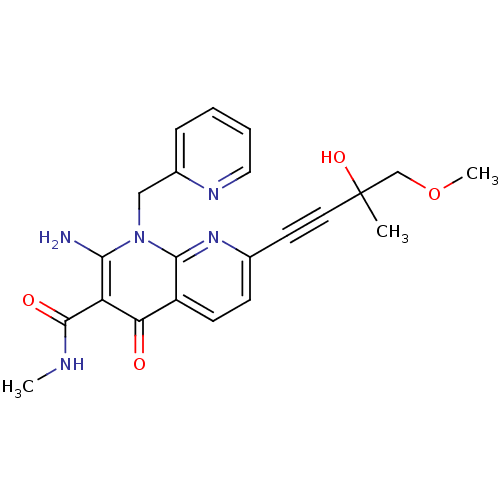
Affinity DataIC50: 14nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 21nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 23nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 23nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 34nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 37nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 38nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 40nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 43nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 46nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 65nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair
Affinity DataIC50: 257nMT: 2°CAssay Description:Activity of VEGFR-3 is measured by ELISA assay by measuring the intensity of phosphorylation of the substrate poly Glu-Tyr.More data for this Ligand-Target Pair