Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Cyclin-A1/Cyclin-dependent kinase 2
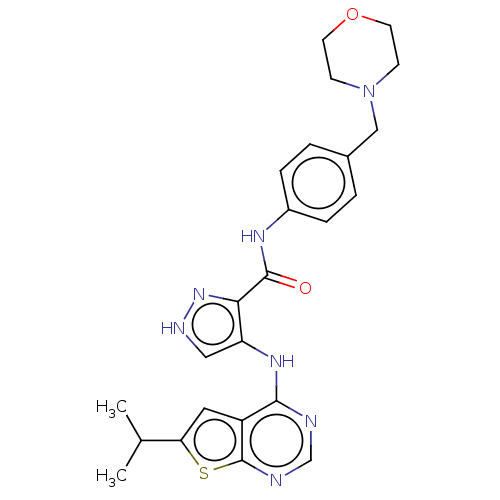
Ligand
BDBM50270334
Substrate
n/a
Meas. Tech.
ChEMBL_1707952 (CHEMBL4059185)
IC50
9.9±n/a nM
Citation
 Wang, Y; Zhi, Y; Jin, Q; Lu, S; Lin, G; Yuan, H; Yang, T; Wang, Z; Yao, C; Ling, J; Guo, H; Li, T; Jin, J; Li, B; Zhang, L; Chen, Y; Lu, T Discovery of 4-((7H-Pyrrolo[2,3-d]pyrimidin-4-yl)amino)-N-(4-((4-methylpiperazin-1-yl)methyl)phenyl)-1H-pyrazole-3-carboxamide (FN-1501), an FLT3- and CDK-Kinase Inhibitor with Potentially High Efficiency against Acute Myelocytic Leukemia. J Med Chem 61:1499-1518 (2018) [PubMed] Article
Wang, Y; Zhi, Y; Jin, Q; Lu, S; Lin, G; Yuan, H; Yang, T; Wang, Z; Yao, C; Ling, J; Guo, H; Li, T; Jin, J; Li, B; Zhang, L; Chen, Y; Lu, T Discovery of 4-((7H-Pyrrolo[2,3-d]pyrimidin-4-yl)amino)-N-(4-((4-methylpiperazin-1-yl)methyl)phenyl)-1H-pyrazole-3-carboxamide (FN-1501), an FLT3- and CDK-Kinase Inhibitor with Potentially High Efficiency against Acute Myelocytic Leukemia. J Med Chem 61:1499-1518 (2018) [PubMed] Article More Info.:
Target
Name:
Cyclin-A1/Cyclin-dependent kinase 2
Synonyms:
CDK2/CycA | CDK2/Cyclin A1
Type:
n/a
Mol. Mass.:
n/a
Description:
ASSAY_ID of ChEMBL is 1979627
Components:
This complex has 2 components.
Component 1
Name:
Cyclin-dependent kinase 2
Synonyms:
CDK2 | CDK2-Kinase | CDK2_HUMAN | CDKN2 | Cell division protein kinase 2 | Cyclin-dependent kinase 2 (CDK2) | Protein cereblon/Cyclin-dependent kinase 2 | p33 protein kinase
Type:
Enzyme Subunit
Mol. Mass.:
33938.17
Organism:
Human
Description:
P24941
Residue:
298
Sequence:
MENFQKVEKIGEGTYGVVYKARNKLTGEVVALKKIRLDTETEGVPSTAIREISLLKELNHPNIVKLLDVIHTENKLYLVFEFLHQDLKKFMDASALTGIPLPLIKSYLFQLLQGLAFCHSHRVLHRDLKPQNLLINTEGAIKLADFGLARAFGVPVRTYTHEVVTLWYRAPEILLGCKYYSTAVDIWSLGCIFAEMVTRRALFPGDSEIDQLFRIFRTLGTPDEVVWPGVTSMPDYKPSFPKWARQDFSKVVPPLDEDGRSLLSQMLHYDPNKRISAKAALAHPFFQDVTKPVPHLRL
Component 2
Name:
Cyclin-A1
Synonyms:
CCNA1 | CCNA1_HUMAN | CDK2/Cyclin A1 | Cyclin-A1 | Cyclin-A1/Cyclin-dependent kinase 4
Type:
Enzyme
Mol. Mass.:
52343.37
Organism:
Human
Description:
P78396
Residue:
465
Sequence:
METGFPAIMYPGSFIGGWGEEYLSWEGPGLPDFVFQQQPVESEAMHCSNPKSGVVLATVARGPDACQILTRAPLGQDPPQRTVLGLLTANGQYRRTCGQGITRIRCYSGSENAFPPAGKKALPDCGVQEPPKQGFDIYMDELEQGDRDSCSVREGMAFEDVYEVDTGTLKSDLHFLLDFNTVSPMLVDSSLLSQSEDISSLGTDVINVTEYAEEIYQYLREAEIRHRPKAHYMKKQPDITEGMRTILVDWLVEVGEEYKLRAETLYLAVNFLDRFLSCMSVLRGKLQLVGTAAMLLASKYEEIYPPEVDEFVYITDDTYTKRQLLKMEHLLLKVLAFDLTVPTTNQFLLQYLRRQGVCVRTENLAKYVAELSLLEADPFLKYLPSLIAAAAFCLANYTVNKHFWPETLAAFTGYSLSEIVPCLSELHKAYLDIPHRPQQAIREKYKASKYLCVSLMEPPAVLLLQ