Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
DNA gyrase subunit A/B
Ligand
BDBM50577580
Substrate
n/a
Meas. Tech.
ChEMBL_2130602 (CHEMBL4840031)
IC50
39±n/a nM
Citation
 Lu, Y; Vibhute, S; Li, L; Okumu, A; Ratigan, SC; Nolan, S; Papa, JL; Mann, CA; English, A; Chen, A; Seffernick, JT; Koci, B; Duncan, LR; Roth, B; Cummings, JE; Slayden, RA; Lindert, S; McElroy, CA; Wozniak, DJ; Yalowich, J; Mitton-Fry, MJ Optimization of TopoIV Potency, ADMET Properties, and hERG Inhibition of 5-Amino-1,3-dioxane-Linked Novel Bacterial Topoisomerase Inhibitors: Identification of a Lead with J Med Chem 64:15214-15249 (2021) [PubMed] Article
Lu, Y; Vibhute, S; Li, L; Okumu, A; Ratigan, SC; Nolan, S; Papa, JL; Mann, CA; English, A; Chen, A; Seffernick, JT; Koci, B; Duncan, LR; Roth, B; Cummings, JE; Slayden, RA; Lindert, S; McElroy, CA; Wozniak, DJ; Yalowich, J; Mitton-Fry, MJ Optimization of TopoIV Potency, ADMET Properties, and hERG Inhibition of 5-Amino-1,3-dioxane-Linked Novel Bacterial Topoisomerase Inhibitors: Identification of a Lead with J Med Chem 64:15214-15249 (2021) [PubMed] Article More Info.:
Target
Name:
DNA gyrase subunit A/B
Synonyms:
DNA Gyrase
Type:
A2B2 tetramer
Mol. Mass.:
n/a
Description:
n/a
Components:
This complex has 2 components.
Component 1
Name:
DNA gyrase subunit A
Synonyms:
DNA gyrase subunit A (gyrA) | GYRA_STAAU | gyrA
Type:
Enzyme Subunit
Mol. Mass.:
99588.82
Organism:
Staphylococcus aureus
Description:
n/a
Residue:
889
Sequence:
MAELPQSRINERNITSEMRESFLDYAMSVIVARALPDVRDGLKPVHRRILYGLNEQGMTPDKSYKKSARIVGDVMGKYHPHGDSSIYEAMVRMAQDFSYRYPLVDGQGNFGSMDGDGAAAMRYTEARMTKITLELLRDINKDTIDFIDNYDGNEREPSVLPARFPNLLANGASGIAVGMATNIPPHNLTELINGVLSLSKNPDISIAELMEDIEGPDFPTAGLILGKSGIRRAYETGRGSIQMRSRAVIEERGGGRQRIVVTEIPFQVNKARMIEKIAELVRDKKIDGITDLRDETSLRTGVRVVIDVRKDANASVILNNLYKQTPLQTSFGVNMIALVNGRPKLINLKEALVHYLEHQKTVVRRRTQYNLRKAKDRAHILEGLRIALDHIDEIISTIRESDTDKVAMESLQQRFKLSEKQAQAILDMRLRRLTGLERDKIEAEYNELLNYISELEAILADEEVLLQLVRDELTEIRDRFGDDRRTEIQLGGFEDLEDEDLIPEEQIVITLSHNNYIKRLPVSTYRAQNRGGRGVQGMNTLEEDFVSQLVTLSTHDHVLFFTNKGRVYKLKGYEVPELSRQSKGIPVVNAIELENDEVISTMIAVKDLESEDNFLVFATKRGVVKRSALSNFSRINRNGKIAISFREDDELIAVRLTSGQEDILIGTSHASLIRFPESTLRPLGRTATGVKGITLREGDEVVGLDVAHANSVDEVLVVTENGYGKRTPVNDYRLSNRGGKGIKTATITERNGNVVCITTVTGEEDLMIVTNAGVIIRLDVADISQNGRAAQGVRLIRLGDDQFVSTVAKVKEDAEDETNEDEQSTSTVSEDGTEQQREAVVNDETPGNAIHTEVIDSEENDEDGRIEVRQDFMDRVEEDIQQSLDEDEE
Component 2
Name:
DNA gyrase subunit B
Synonyms:
DNA gyrase | DNA gyrase subunit B (DNA gyraseB) | DNA gyrase subunit B (gyrB) | GYRB_STAAU | gyrB
Type:
Enzyme Subunit
Mol. Mass.:
72530.91
Organism:
Staphylococcus aureus
Description:
n/a
Residue:
644
Sequence:
MVTALSDVNNTDNYGAGQIQVLEGLEAVRKRPGMYIGSTSERGLHHLVWEIVDNSIDEALAGYANQIEVVIEKDNWIKVTDNGRGIPVDIQEKMGRPAVEVILTVLHAGGKFGGGGYKVSGGLHGVGSSVVNALSQDLEVYVHRNETIYHQAYKKGVPQFDLKEVGTTDKTGTVIRFKADGEIFTETTVYNYETLQQRIRELAFLNKGIQITLRDERDEENVREDSYHYEGGIKSYVELLNENKEPIHDEPIYIHQSKDDIEVEIAIQYNSGYATNLLTYANNIHTYEGGTHEDGFKRALTRVLNSYGLSSKIMKEEKDRLSGEDTREGMTAIISIKHGDPQFEGQTKTKLGNSEVRQVVDKLFSEHFERFLYENPQVARTVVEKGIMAARARVAAKKAREVTRRKSALDVASLPGKLADCSSKSPEECEIFLVEGDSAGGSTKSGRDSRTQAILPLRGKILNVEKARLDRILNNNEIRQMITAFGTGIGGDFDLAKARYHKIVIMTDADVDGAHIRTLLLTFFYRFMRPLIEAGYVYIAQPPLYKLTQGKQKYYVYNDRELDKLKSELNPTPKWSIARYKGLGEMNADQLWETTMNPEHRALLQVKLEDAIEADQTFEMLMGDVVENRRQFIEDNAVYANLDF
Inhibitor
Name:
BDBM50577580
Synonyms:
CHEMBL4865826
Type:
Small organic molecule
Emp. Form.:
C23H25FN4O6
Mol. Mass.:
472.4662
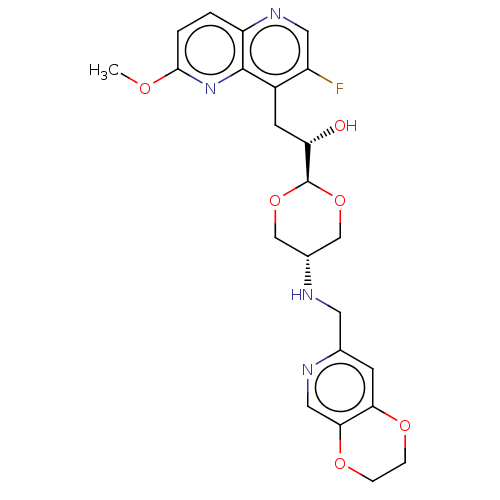
SMILES:
[H][C@@]1(OC[C@@H](CO1)NCc1cc2OCCOc2cn1)[C@@H](O)Cc1c(F)cnc2ccc(OC)nc12 |r,wU:19.22,wD:4.7,1.0,(29.25,-15.08,;27.93,-14.32,;26.58,-13.57,;26.58,-12.02,;27.92,-11.25,;29.26,-12.02,;29.26,-13.55,;27.92,-9.71,;29.25,-8.93,;30.58,-9.7,;31.91,-8.93,;33.24,-9.69,;34.57,-8.91,;35.91,-9.69,;35.91,-11.23,;34.57,-12,;33.24,-11.23,;31.91,-12.01,;30.58,-11.24,;27.93,-15.86,;29.27,-16.62,;26.61,-16.63,;26.61,-18.17,;27.95,-18.93,;29.28,-18.16,;27.95,-20.48,;26.62,-21.26,;25.29,-20.49,;23.95,-21.27,;22.62,-20.5,;22.62,-18.95,;21.29,-18.18,;21.29,-16.64,;23.95,-18.18,;25.29,-18.95,)|