Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform
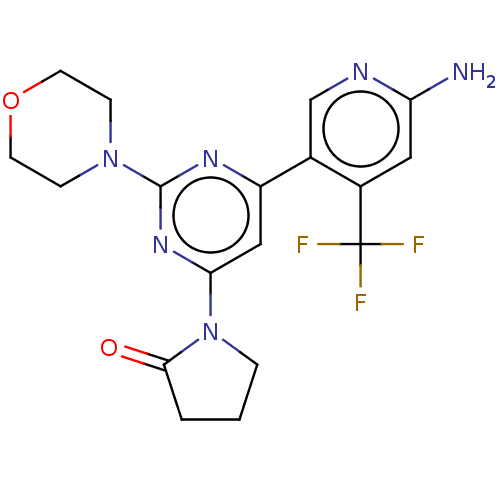
Ligand
BDBM50598027
Substrate
n/a
Meas. Tech.
ChEMBL_2225467 (CHEMBL5138980)
IC50
157±n/a nM
Citation
 Fairhurst, RA; Furet, P; Imbach-Weese, P; Stauffer, F; Rueeger, H; McCarthy, C; Ripoche, S; Oswald, S; Arnaud, B; Jary, A; Maira, M; Schnell, C; Guthy, DA; Wartmann, M; Kiffe, M; Desrayaud, S; Blasco, F; Widmer, T; Seiler, F; Gutmann, S; Knapp, M; Caravatti, G Identification of NVP-CLR457 as an Orally Bioavailable Non-CNS-Penetrant pan-Class IA Phosphoinositol-3-Kinase Inhibitor. J Med Chem 65:8345-8379 (2022) [PubMed]
Fairhurst, RA; Furet, P; Imbach-Weese, P; Stauffer, F; Rueeger, H; McCarthy, C; Ripoche, S; Oswald, S; Arnaud, B; Jary, A; Maira, M; Schnell, C; Guthy, DA; Wartmann, M; Kiffe, M; Desrayaud, S; Blasco, F; Widmer, T; Seiler, F; Gutmann, S; Knapp, M; Caravatti, G Identification of NVP-CLR457 as an Orally Bioavailable Non-CNS-Penetrant pan-Class IA Phosphoinositol-3-Kinase Inhibitor. J Med Chem 65:8345-8379 (2022) [PubMed] More Info.:
Target
Name:
Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform
Synonyms:
PI3-kinase subunit beta | PI3K-beta | PI3Kbeta | PK3CB_RAT | Phosphatidylinositol 4,5-bisphosphate 3-kinase 110 kDa catalytic subunit beta | Pik3cb | PtdIns-3-kinase subunit beta | PtdIns-3-kinase subunit p110-beta | p110beta
Type:
Enzyme Catalytic Domain
Mol. Mass.:
122616.58
Organism:
Rat
Description:
Q9Z1L0
Residue:
1070
Sequence:
MCFRSIMPPAMADTLDIWAVDSQIASDGSISVDFLLPTGIYIQLEVPREATISYIKQMLWKQVHNYPMFNLLMDIDSYMFACVNQTAVYEELEDETRRLCDVRPFLPVLKLVTRSCDPAEKLDSKIGVLIGKGLHEFDALKDPEVNEFRRKMRKFSEDKIQSLVGLSWIDWLKHTYPPEHEPSVLENLEDKLYGGKLVVAVHFENSQDVFSFQVSPNLNPIKINELAIQKRLTIRGKEEEASPCDYVLQVSGRVEYVFGDHPLIQFQYIRNCVMNRTLPHFILVECCKIKKMYEQEMIAIEAAINRNSSSLPLPLPPKKTRVISHVWGNNNPFQIVLVKGNKLNTEETVKVHVRAGLFHGTELLCKTVVSSEISGKNDHIWNEQLEFDINICDLPRMARLCFAVYAVLDKVKTKKSTKTINPSKYQTIRKAGKVHYPVAWVNTMVFDFKGQLRSGDVILHSWSSFPDELEEMLNPMGTVQTNPYAENATALHIKFPENKKQPYYYPPFDKIIEKAAEIASGDSANVSSRGGKKFLAVLKEILDRDPLSQLCENEMDLIWTLRQDCRENFPQSLPKLLLSIKWNKLEDVAQLQALLQIWPKLPPREALELLDFNYPDQYVREYAVGCLRQMSDEELSQYLLQLVQVLKYEPFLDCALSRFLLERALDNRRIGQFLFWHLRSEVHTPAVSIQFGVILEAYCRGSVGHMKVLSKQVEALNKLKTLNSLIKLNAMKLNRAKGKEAMHTCLKQSAYREALSDLQSPLNPCVILSELYVEKCRYMDSKMKPLWLVYSNRAFGEDAVGVIFKNGDDLRQDMLTLQMLRLMDLLWKEAGLDLRMLPYGCLATGDRSGLIEVVSTSETIADIQLNSSNVAATAAFNKDALLNWLKEYNSGDDLDRAIEEFTLSCAGYCVASYVLGIGDRHSDNIMVKKTGQLFHIDFGHILGNFKSKFGIKRERVPFILTYDFIHVIQQGKTGNTEKFGRFRQCCEDAYLILRRHGNLFITLFALMLTAGLPELTSVKDIQYLKDSLALGKSEEEALKQFKQKFDEALRESWTTKVNWMAHTVRKDYRS