Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
cGMP-specific 3',5'-cyclic phosphodiesterase
Ligand
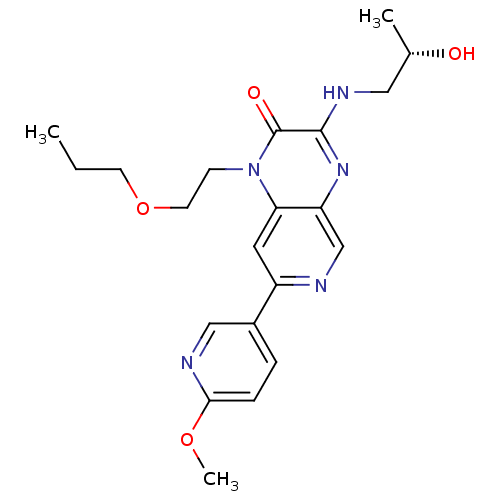
BDBM50300989
Substrate
n/a
Meas. Tech.
ChEMBL_597959 (CHEMBL1042744)
IC50
0.090000±n/a nM
Citation
 Hughes, RO; Walker, JK; Rogier, DJ; Heasley, SE; Blevis-Bal, RM; Benson, AG; Jacobsen, EJ; Cubbage, JW; Fobian, YM; Owen, DR; Freskos, JN; Molyneaux, JM; Brown, DL; Acker, BA; Maddux, TM; Tollefson, MB; Moon, JB; Mischke, BV; Rumsey, JM; Zheng, Y; MacInnes, A; Bond, BR; Yu, Y Optimization of the aminopyridopyrazinones class of PDE5 inhibitors: discovery of 3-[(trans-4-hydroxycyclohexyl)amino]-7-(6-methoxypyridin-3-yl)-1-(2-propoxyethyl)pyrido[3,4-b]pyrazin-2(1H)-one. Bioorg Med Chem Lett 19:5209-13 (2009) [PubMed] Article
Hughes, RO; Walker, JK; Rogier, DJ; Heasley, SE; Blevis-Bal, RM; Benson, AG; Jacobsen, EJ; Cubbage, JW; Fobian, YM; Owen, DR; Freskos, JN; Molyneaux, JM; Brown, DL; Acker, BA; Maddux, TM; Tollefson, MB; Moon, JB; Mischke, BV; Rumsey, JM; Zheng, Y; MacInnes, A; Bond, BR; Yu, Y Optimization of the aminopyridopyrazinones class of PDE5 inhibitors: discovery of 3-[(trans-4-hydroxycyclohexyl)amino]-7-(6-methoxypyridin-3-yl)-1-(2-propoxyethyl)pyrido[3,4-b]pyrazin-2(1H)-one. Bioorg Med Chem Lett 19:5209-13 (2009) [PubMed] Article More Info.:
Target
Name:
cGMP-specific 3',5'-cyclic phosphodiesterase
Synonyms:
3',5'-cyclic phosphodiesterase | CGB-PDE | PDE5 | PDE5A | PDE5A_HUMAN | Phosphodiesterase 2 and 5 (PDE2 and PDE5) | Phosphodiesterase 5 (PDE5) | Phosphodiesterase 5A | Phosphodiesterase 5A (PDE5A) | cGMP-binding cGMP-specific phosphodiesterase | cGMP-specific 3',5'-cyclic phosphodiesterase
Type:
Protein
Mol. Mass.:
99975.83
Organism:
Homo sapiens (Human)
Description:
O76074
Residue:
875
Sequence:
MERAGPSFGQQRQQQQPQQQKQQQRDQDSVEAWLDDHWDFTFSYFVRKATREMVNAWFAERVHTIPVCKEGIRGHTESCSCPLQQSPRADNSAPGTPTRKISASEFDRPLRPIVVKDSEGTVSFLSDSEKKEQMPLTPPRFDHDEGDQCSRLLELVKDISSHLDVTALCHKIFLHIHGLISADRYSLFLVCEDSSNDKFLISRLFDVAEGSTLEEVSNNCIRLEWNKGIVGHVAALGEPLNIKDAYEDPRFNAEVDQITGYKTQSILCMPIKNHREEVVGVAQAINKKSGNGGTFTEKDEKDFAAYLAFCGIVLHNAQLYETSLLENKRNQVLLDLASLIFEEQQSLEVILKKIAATIISFMQVQKCTIFIVDEDCSDSFSSVFHMECEELEKSSDTLTREHDANKINYMYAQYVKNTMEPLNIPDVSKDKRFPWTTENTGNVNQQCIRSLLCTPIKNGKKNKVIGVCQLVNKMEENTGKVKPFNRNDEQFLEAFVIFCGLGIQNTQMYEAVERAMAKQMVTLEVLSYHASAAEEETRELQSLAAAVVPSAQTLKITDFSFSDFELSDLETALCTIRMFTDLNLVQNFQMKHEVLCRWILSVKKNYRKNVAYHNWRHAFNTAQCMFAALKAGKIQNKLTDLEILALLIAALSHDLDHRGVNNSYIQRSEHPLAQLYCHSIMEHHHFDQCLMILNSPGNQILSGLSIEEYKTTLKIIKQAILATDLALYIKRRGEFFELIRKNQFNLEDPHQKELFLAMLMTACDLSAITKPWPIQQRIAELVATEFFDQGDRERKELNIEPTDLMNREKKNKIPSMQVGFIDAICLQLYEALTHVSEDCFPLLDGCRKNRQKWQALAEQQEKMLINGESGQAKRN