Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Dipeptidyl peptidase 4
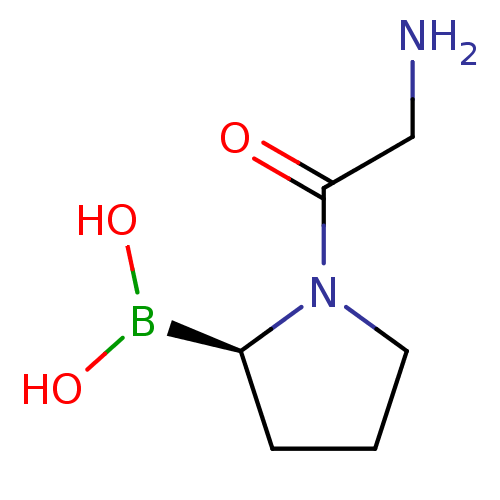
Ligand
BDBM50050511
Substrate
n/a
Meas. Tech.
ChEMBL_53379 (CHEMBL665783)
IC50
16000±n/a nM
Citation
 Coutts, SJ; Kelly, TA; Snow, RJ; Kennedy, CA; Barton, RW; Adams, J; Krolikowski, DA; Freeman, DM; Campbell, SJ; Ksiazek, JF; Bachovchin, WW Structure-activity relationships of boronic acid inhibitors of dipeptidyl peptidase IV. 1. Variation of the P2 position of Xaa-boroPro dipeptides. J Med Chem 39:2087-94 (1996) [PubMed] Article
Coutts, SJ; Kelly, TA; Snow, RJ; Kennedy, CA; Barton, RW; Adams, J; Krolikowski, DA; Freeman, DM; Campbell, SJ; Ksiazek, JF; Bachovchin, WW Structure-activity relationships of boronic acid inhibitors of dipeptidyl peptidase IV. 1. Variation of the P2 position of Xaa-boroPro dipeptides. J Med Chem 39:2087-94 (1996) [PubMed] Article More Info.:
Target
Name:
Dipeptidyl peptidase 4
Synonyms:
ADABP | ADCP2 | Adenosine deaminase complexing protein 2 | CD26 | CD_antigen=CD26 | DPP IV | DPP4 | DPP4_HUMAN | DPPIV | Dipeptidyl peptidase 4 (DDP-IV) | Dipeptidyl peptidase 4 (DPP IV) | Dipeptidyl peptidase 4 (DPP-4) | Dipeptidyl peptidase 4 (DPP4) | Dipeptidyl peptidase 4 (DPPIV) | Dipeptidyl peptidase 4 membrane form | Dipeptidyl peptidase 4 soluble form | Dipeptidyl peptidase IV | Dipeptidyl peptidase IV (DDP-4) | Dipeptidyl peptidase IV (DDP-IV) | Dipeptidyl peptidase IV (DPP IV) | Dipeptidyl peptidase IV membrane form | Dipeptidyl peptidase IV soluble form | Dipeptidyl peptidase-IV (DPP-4) | Dipeptidyl peptidase-IV (DPP-IV) | T-cell activation antigen CD26 | TP103
Type:
Enzyme
Mol. Mass.:
88271.01
Organism:
Human
Description:
P27487
Residue:
766
Sequence:
MKTPWKVLLGLLGAAALVTIITVPVVLLNKGTDDATADSRKTYTLTDYLKNTYRLKLYSLRWISDHEYLYKQENNILVFNAEYGNSSVFLENSTFDEFGHSINDYSISPDGQFILLEYNYVKQWRHSYTASYDIYDLNKRQLITEERIPNNTQWVTWSPVGHKLAYVWNNDIYVKIEPNLPSYRITWTGKEDIIYNGITDWVYEEEVFSAYSALWWSPNGTFLAYAQFNDTEVPLIEYSFYSDESLQYPKTVRVPYPKAGAVNPTVKFFVVNTDSLSSVTNATSIQITAPASMLIGDHYLCDVTWATQERISLQWLRRIQNYSVMDICDYDESSGRWNCLVARQHIEMSTTGWVGRFRPSEPHFTLDGNSFYKIISNEEGYRHICYFQIDKKDCTFITKGTWEVIGIEALTSDYLYYISNEYKGMPGGRNLYKIQLSDYTKVTCLSCELNPERCQYYSVSFSKEAKYYQLRCSGPGLPLYTLHSSVNDKGLRVLEDNSALDKMLQNVQMPSKKLDFIILNETKFWYQMILPPHFDKSKKYPLLLDVYAGPCSQKADTVFRLNWATYLASTENIIVASFDGRGSGYQGDKIMHAINRRLGTFEVEDQIEAARQFSKMGFVDNKRIAIWGWSYGGYVTSMVLGSGSGVFKCGIAVAPVSRWEYYDSVYTERYMGLPTPEDNLDHYRNSTVMSRAENFKQVEYLLIHGTADDNVHFQQSAQISKALVDVGVDFQAMWYTDEDHGIASSTAHQHIYTHMSHFIKQCFSLP