Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Cyclin-A1/Cyclin-dependent kinase 2
Ligand
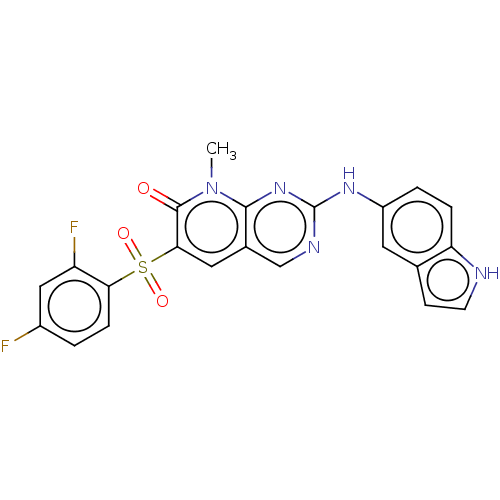
BDBM50135286
Substrate
n/a
Meas. Tech.
ChEMBL_1546467 (CHEMBL3748658)
IC50
>10000±n/a nM
Citation
 Reddy, MV; Akula, B; Jatiani, S; Vasquez-Del Carpio, R; Billa, VK; Mallireddigari, MR; Cosenza, SC; Venkata Subbaiah, DR; Bharathi, EV; Pallela, VR; Ramkumar, P; Jain, R; Aggarwal, AK; Reddy, EP Discovery of 2-(1H-indol-5-ylamino)-6-(2,4-difluorophenylsulfonyl)-8-methylpyrido[2,3-d]pyrimidin-7(8H)-one (7ao) as a potent selective inhibitor of Polo like kinase 2 (PLK2). Bioorg Med Chem 24:521-44 (2016) [PubMed] Article
Reddy, MV; Akula, B; Jatiani, S; Vasquez-Del Carpio, R; Billa, VK; Mallireddigari, MR; Cosenza, SC; Venkata Subbaiah, DR; Bharathi, EV; Pallela, VR; Ramkumar, P; Jain, R; Aggarwal, AK; Reddy, EP Discovery of 2-(1H-indol-5-ylamino)-6-(2,4-difluorophenylsulfonyl)-8-methylpyrido[2,3-d]pyrimidin-7(8H)-one (7ao) as a potent selective inhibitor of Polo like kinase 2 (PLK2). Bioorg Med Chem 24:521-44 (2016) [PubMed] Article More Info.:
Target
Name:
Cyclin-A1/Cyclin-dependent kinase 2
Synonyms:
CDK2/CycA | CDK2/Cyclin A1
Type:
n/a
Mol. Mass.:
n/a
Description:
ASSAY_ID of ChEMBL is 1979627
Components:
This complex has 2 components.
Component 1
Name:
Cyclin-dependent kinase 2
Synonyms:
CDK2 | CDK2-Kinase | CDK2_HUMAN | CDKN2 | Cell division protein kinase 2 | Cyclin-dependent kinase 2 (CDK2) | Protein cereblon/Cyclin-dependent kinase 2 | p33 protein kinase
Type:
Enzyme Subunit
Mol. Mass.:
33938.17
Organism:
Human
Description:
P24941
Residue:
298
Sequence:
MENFQKVEKIGEGTYGVVYKARNKLTGEVVALKKIRLDTETEGVPSTAIREISLLKELNHPNIVKLLDVIHTENKLYLVFEFLHQDLKKFMDASALTGIPLPLIKSYLFQLLQGLAFCHSHRVLHRDLKPQNLLINTEGAIKLADFGLARAFGVPVRTYTHEVVTLWYRAPEILLGCKYYSTAVDIWSLGCIFAEMVTRRALFPGDSEIDQLFRIFRTLGTPDEVVWPGVTSMPDYKPSFPKWARQDFSKVVPPLDEDGRSLLSQMLHYDPNKRISAKAALAHPFFQDVTKPVPHLRL
Component 2
Name:
Cyclin-A1
Synonyms:
CCNA1 | CCNA1_HUMAN | CDK2/Cyclin A1 | Cyclin-A1 | Cyclin-A1/Cyclin-dependent kinase 4
Type:
Enzyme
Mol. Mass.:
52343.37
Organism:
Human
Description:
P78396
Residue:
465
Sequence:
METGFPAIMYPGSFIGGWGEEYLSWEGPGLPDFVFQQQPVESEAMHCSNPKSGVVLATVARGPDACQILTRAPLGQDPPQRTVLGLLTANGQYRRTCGQGITRIRCYSGSENAFPPAGKKALPDCGVQEPPKQGFDIYMDELEQGDRDSCSVREGMAFEDVYEVDTGTLKSDLHFLLDFNTVSPMLVDSSLLSQSEDISSLGTDVINVTEYAEEIYQYLREAEIRHRPKAHYMKKQPDITEGMRTILVDWLVEVGEEYKLRAETLYLAVNFLDRFLSCMSVLRGKLQLVGTAAMLLASKYEEIYPPEVDEFVYITDDTYTKRQLLKMEHLLLKVLAFDLTVPTTNQFLLQYLRRQGVCVRTENLAKYVAELSLLEADPFLKYLPSLIAAAAFCLANYTVNKHFWPETLAAFTGYSLSEIVPCLSELHKAYLDIPHRPQQAIREKYKASKYLCVSLMEPPAVLLLQ