Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Trifunctional purine biosynthetic protein adenosine-3
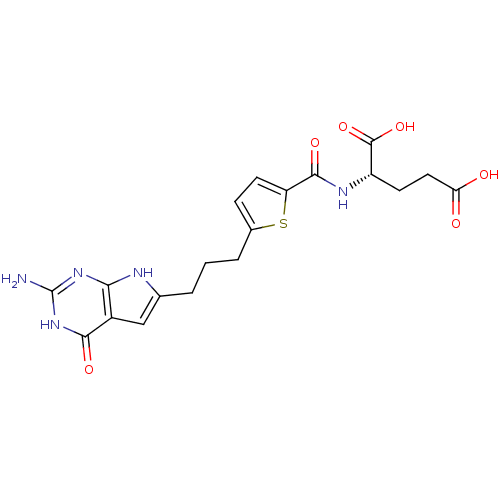
Ligand
BDBM50354833
Substrate
n/a
Meas. Tech.
ChEMBL_1928983 (CHEMBL4432159)
Ki
68±n/a nM
Citation
 Golani, LK; Wallace-Povirk, A; Deis, SM; Wong, J; Ke, J; Gu, X; Raghavan, S; Wilson, MR; Li, X; Polin, L; de Waal, PW; White, K; Kushner, J; O'Connor, C; Hou, Z; Xu, HE; Melcher, K; Dann, CE; Matherly, LH; Gangjee, A Tumor Targeting with Novel 6-Substituted Pyrrolo [2,3-d] Pyrimidine Antifolates with Heteroatom Bridge Substitutions via Cellular Uptake by Folate Receptor ? and the Proton-Coupled Folate Transporter and Inhibition of de Novo Purine Nucleotide Biosynthesis. J Med Chem 59:7856-76 (2016) [PubMed] Article
Golani, LK; Wallace-Povirk, A; Deis, SM; Wong, J; Ke, J; Gu, X; Raghavan, S; Wilson, MR; Li, X; Polin, L; de Waal, PW; White, K; Kushner, J; O'Connor, C; Hou, Z; Xu, HE; Melcher, K; Dann, CE; Matherly, LH; Gangjee, A Tumor Targeting with Novel 6-Substituted Pyrrolo [2,3-d] Pyrimidine Antifolates with Heteroatom Bridge Substitutions via Cellular Uptake by Folate Receptor ? and the Proton-Coupled Folate Transporter and Inhibition of de Novo Purine Nucleotide Biosynthesis. J Med Chem 59:7856-76 (2016) [PubMed] Article More Info.:
Target
Name:
Trifunctional purine biosynthetic protein adenosine-3
Synonyms:
GAR Tfase | GAR transformylase | GART | Glycinamide ribonucleotide formyltransferase (GARFTase) | Glycinamide ribonucleotide transformylase (GAR Tfase) | PGFT | PRGS | PUR2_HUMAN | Thymidylate synthase/GAR transformylase/AICAR transformylase
Type:
Protein
Mol. Mass.:
107768.47
Organism:
Human
Description:
P22102
Residue:
1010
Sequence:
MAARVLIIGSGGREHTLAWKLAQSHHVKQVLVAPGNAGTACSEKISNTAISISDHTALAQFCKEKKIEFVVVGPEAPLAAGIVGNLRSAGVQCFGPTAEAAQLESSKRFAKEFMDRHGIPTAQWKAFTKPEEACSFILSADFPALVVKASGLAAGKGVIVAKSKEEACKAVQEIMQEKAFGAAGETIVIEELLDGEEVSCLCFTDGKTVAPMPPAQDHKRLLEGDGGPNTGGMGAYCPAPQVSNDLLLKIKDTVLQRTVDGMQQEGTPYTGILYAGIMLTKNGPKVLEFNCRFGDPECQVILPLLKSDLYEVIQSTLDGLLCTSLPVWLENHTALTVVMASKGYPGDYTKGVEITGFPEAQALGLEVFHAGTALKNGKVVTHGGRVLAVTAIRENLISALEEAKKGLAAIKFEGAIYRKDVGFRAIAFLQQPRSLTYKESGVDIAAGNMLVKKIQPLAKATSRSGCKVDLGGFAGLFDLKAAGFKDPLLASGTDGVGTKLKIAQLCNKHDTIGQDLVAMCVNDILAQGAEPLFFLDYFSCGKLDLSVTEAVVAGIAKACGKAGCALLGGETAEMPDMYPPGEYDLAGFAVGAMERDQKLPHLERITEGDVVVGIASSGLHSNGFSLVRKIVAKSSLQYSSPAPDGCGDQTLGDLLLTPTRIYSHSLLPVLRSGHVKAFAHITGGGLLENIPRVLPEKLGVDLDAQTWRIPRVFSWLQQEGHLSEEEMARTFNCGVGAVLVVSKEQTEQILRDIQQHKEEAWVIGSVVARAEGSPRVKVKNLIESMQINGSVLKNGSLTNHFSFEKKKARVAVLISGTGSNLQALIDSTREPNSSAQIDIVISNKAAVAGLDKAERAGIPTRVINHKLYKNRVEFDSAIDLVLEEFSIDIVCLAGFMRILSGPFVQKWNGKMLNIHPSLLPSFKGSNAHEQALETGVTVTGCTVHFVAEDVDAGQIILQEAVPVKRGDTVATLSERVKLAEHKIFPAALQLVASGTVQLGENGKICWVKEE