Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Epoxide hydrolase B
Ligand
BDBM25742
Substrate
BDBM25726
Meas. Tech.
Mtb EHB Inhibition Assay
pH
7±n/a
Temperature
303.15±n/a K
IC50
2060±n/a nM
Citation
 Biswal, BK; Morisseau, C; Garen, G; Cherney, MM; Garen, C; Niu, C; Hammock, BD; James, MN The molecular structure of epoxide hydrolase B from Mycobacterium tuberculosis and its complex with a urea-based inhibitor. J Mol Biol 381:897-912 (2008) [PubMed] Article
Biswal, BK; Morisseau, C; Garen, G; Cherney, MM; Garen, C; Niu, C; Hammock, BD; James, MN The molecular structure of epoxide hydrolase B from Mycobacterium tuberculosis and its complex with a urea-based inhibitor. J Mol Biol 381:897-912 (2008) [PubMed] Article Target
Name:
Epoxide hydrolase B
Synonyms:
EPHB_MYCTO | Epoxide Hydrolase B (EHB) | Mtb EHB
Type:
Enzyme
Mol. Mass.:
39285.78
Organism:
Mycobacterium tuberculosis
Description:
The His6-tagged Mtb EHB was expressed in E. coli and purified.
Residue:
356
Sequence:
MSQVHRILNCRGTRIHAVADSPPDQQGPLVVLLHGFPESWYSWRHQIPALAGAGYRVVAIDQRGYGRSSKYRVQKAYRIKELVGDVVGVLDSYGAEQAFVVGHDWGAPVAWTFAWLHPDRCAGVVGISVPFAGRGVIGLPGSPFGERRPSDYHLELAGPGRVWYQDYFAVQDGIITEIEEDLRGWLLGLTYTVSGEGMMAATKAAVDAGVDLESMDPIDVIRAGPLCMAEGARLKDAFVYPETMPAWFTEADLDFYTGEFERSGFGGPLSFYHNIDNDWHDLADQQGKPLTPPALFIGGQYDVGTIWGAQAIERAHEVMPNYRGTHMIADVGHWIQQEAPEETNRLLLDFLGGLRP
Inhibitor
Name:
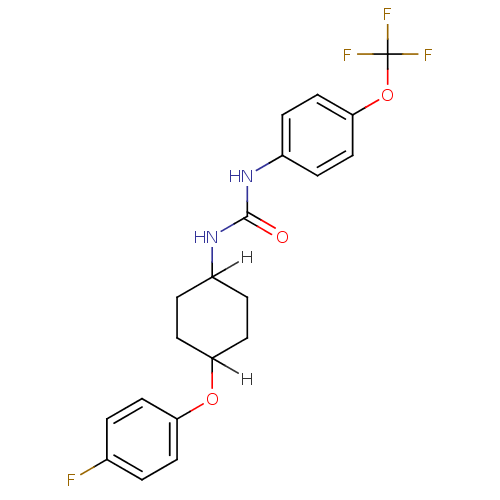
BDBM25742
Synonyms:
1-[4-(4-fluorophenoxy)cyclohexyl]-3-[4-(trifluoromethoxy)phenyl]urea | Urea-based compound, 23
Type:
Small organic molecule
Emp. Form.:
C20H20F4N2O3
Mol. Mass.:
412.378
SMILES:
Fc1ccc(OC2CCC(CC2)NC(=O)Nc2ccc(OC(F)(F)F)cc2)cc1 |(13.37,-2.95,;12.06,-2.13,;10.72,-2.9,;9.39,-2.13,;9.39,-.59,;7.9,-.2,;7.9,1.34,;6.86,2.47,;5.01,1.58,;3.75,1.86,;4.78,.7,;6.61,1.63,;2.42,1.09,;1.08,1.86,;1.08,3.4,;-.25,1.08,;-1.58,1.85,;-2.92,1.08,;-4.25,1.85,;-4.25,3.4,;-5.58,4.17,;-6.92,3.39,;-8.25,4.16,;-8.25,2.62,;-6.92,1.85,;-2.92,4.17,;-1.58,3.4,;10.72,.18,;12.15,-.58,)|
Substrate
Name:
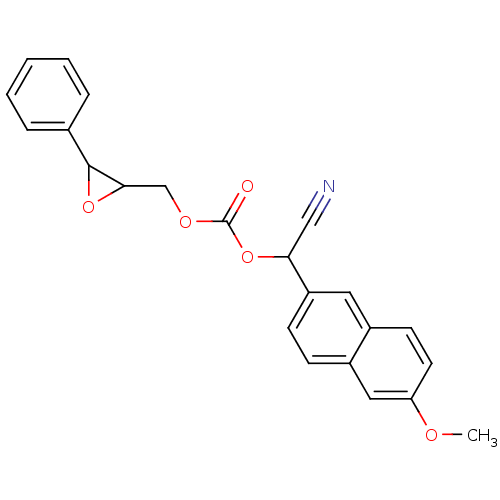
BDBM25726
Synonyms:
2-(6-methoxynaphthalen-2-yl)-2-({[(3-phenyloxiran-2-yl)methoxy]carbonyl}oxy)acetonitrile | alpha-cyanocarbonate epoxide | cyano(6-methoxy-naphthalen-2-yl)methyl trans-[(3-phenyloxiran-2-yl)methyl] carbonate
Type:
Small organic molecule
Emp. Form.:
C23H19NO5
Mol. Mass.:
389.4007
SMILES:
COc1ccc2cc(ccc2c1)C(OC(=O)OCC1OC1c1ccccc1)C#N