Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Vascular endothelial growth factor receptor 3
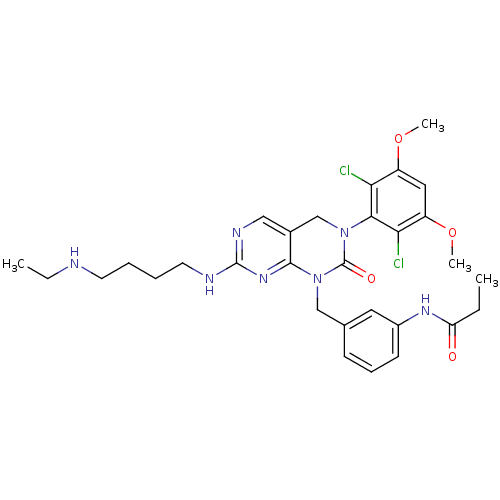
Ligand
BDBM81378
Substrate
n/a
Meas. Tech.
In Vitro Activity Kinase Assay
Kd
340±0.0 nM
Citation
 Zhou, W; Hur, W; McDermott, U; Dutt, A; Xian, W; Ficarro, SB; Zhang, J; Sharma, SV; Brugge, J; Meyerson, M; Settleman, J; Gray, NS A structure-guided approach to creating covalent FGFR inhibitors. Chem Biol 17:285-95 (2010) [PubMed] Article
Zhou, W; Hur, W; McDermott, U; Dutt, A; Xian, W; Ficarro, SB; Zhang, J; Sharma, SV; Brugge, J; Meyerson, M; Settleman, J; Gray, NS A structure-guided approach to creating covalent FGFR inhibitors. Chem Biol 17:285-95 (2010) [PubMed] Article More Info.:
Target
Name:
Vascular endothelial growth factor receptor 3
Synonyms:
FLT-4 | FLT4 | Fms-like tyrosine kinase 4 | Focal adhesion kinase 1/vascular endothelial growth factor receptor 3 | VEGFR-3 | VEGFR3 | VGFR3_HUMAN | Vascular endothelial growth factor receptor | Vascular endothelial growth factor receptor 3 (VEGFR-3) | Vascular endothelial growth factor receptor 3 (VEGFR3)
Type:
Protein
Mol. Mass.:
152749.58
Organism:
Homo sapiens (Human)
Description:
P35916-2
Residue:
1363
Sequence:
MQRGAALCLRLWLCLGLLDGLVSGYSMTPPTLNITEESHVIDTGDSLSISCRGQHPLEWAWPGAQEAPATGDKDSEDTGVVRDCEGTDARPYCKVLLLHEVHANDTGSYVCYYKYIKARIEGTTAASSYVFVRDFEQPFINKPDTLLVNRKDAMWVPCLVSIPGLNVTLRSQSSVLWPDGQEVVWDDRRGMLVSTPLLHDALYLQCETTWGDQDFLSNPFLVHITGNELYDIQLLPRKSLELLVGEKLVLNCTVWAEFNSGVTFDWDYPGKQAERGKWVPERRSQQTHTELSSILTIHNVSQHDLGSYVCKANNGIQRFRESTEVIVHENPFISVEWLKGPILEATAGDELVKLPVKLAAYPPPEFQWYKDGKALSGRHSPHALVLKEVTEASTGTYTLALWNSAAGLRRNISLELVVNVPPQIHEKEASSPSIYSRHSRQALTCTAYGVPLPLSIQWHWRPWTPCKMFAQRSLRRRQQQDLMPQCRDWRAVTTQDAVNPIESLDTWTEFVEGKNKTVSKLVIQNANVSAMYKCVVSNKVGQDERLIYFYVTTIPDGFTIESKPSEELLEGQPVLLSCQADSYKYEHLRWYRLNLSTLHDAHGNPLLLDCKNVHLFATPLAASLEEVAPGARHATLSLSIPRVAPEHEGHYVCEVQDRRSHDKHCHKKYLSVQALEAPRLTQNLTDLLVNVSDSLEMQCLVAGAHAPSIVWYKDERLLEEKSGVDLADSNQKLSIQRVREEDAGRYLCSVCNAKGCVNSSASVAVEGSEDKGSMEIVILVGTGVIAVFFWVLLLLIFCNMRRPAHADIKTGYLSIIMDPGEVPLEEQCEYLSYDASQWEFPRERLHLGRVLGYGAFGKVVEASAFGIHKGSSCDTVAVKMLKEGATASEHRALMSELKILIHIGNHLNVVNLLGACTKPQGPLMVIVEFCKYGNLSNFLRAKRDAFSPCAEKSPEQRGRFRAMVELARLDRRRPGSSDRVLFARFSKTEGGARRASPDQEAEDLWLSPLTMEDLVCYSFQVARGMEFLASRKCIHRDLAARNILLSESDVVKICDFGLARDIYKDPDYVRKGSARLPLKWMAPESIFDKVYTTQSDVWSFGVLLWEIFSLGASPYPGVQINEEFCQRLRDGTRMRAPELATPAIRRIMLNCWSGDPKARPAFSELVEILGDLLQGRGLQEEEEVCMAPRSSQSSEEGSFSQVSTMALHIAQADAEDSPPSLQRHSLAARYYNWVSFPGCLARGAETRGSSRMKTFEEFPMTPTTYKGSVDNQTDSGMVLASEEFEQIESRHRQESGFSCKGPGQNVAVTRAHPDSQGRRRRPERGARGGQVFYNSEYGELSEPSEEDHCSPSARVTFFTDNSY