Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Histone acetyltransferase KAT2A
Ligand
BDBM220447
Substrate
n/a
Meas. Tech.
ChEMBL_1670375 (CHEMBL4020263)
Kd
>1000±n/a nM
Citation
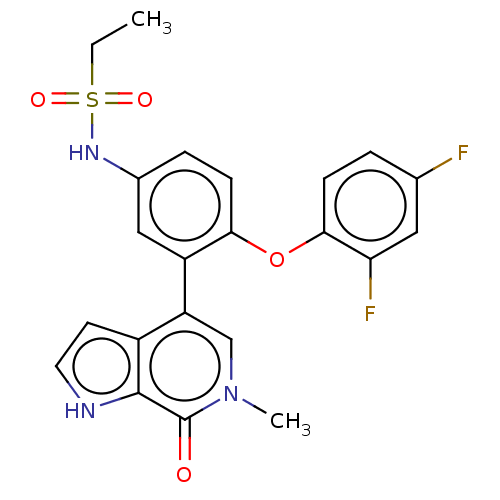
 McDaniel, KF; Wang, L; Soltwedel, T; Fidanze, SD; Hasvold, LA; Liu, D; Mantei, RA; Pratt, JK; Sheppard, GS; Bui, MH; Faivre, EJ; Huang, X; Li, L; Lin, X; Wang, R; Warder, SE; Wilcox, D; Albert, DH; Magoc, TJ; Rajaraman, G; Park, CH; Hutchins, CW; Shen, JJ; Edalji, RP; Sun, CC; Martin, R; Gao, W; Wong, S; Fang, G; Elmore, SW; Shen, Y; Kati, WM Discovery of N-(4-(2,4-Difluorophenoxy)-3-(6-methyl-7-oxo-6,7-dihydro-1H-pyrrolo[2,3-c]pyridin-4-yl)phenyl)ethanesulfonamide (ABBV-075/Mivebresib), a Potent and Orally Available Bromodomain and Extraterminal Domain (BET) Family Bromodomain Inhibitor. J Med Chem 60:8369-8384 (2017) [PubMed] Article
McDaniel, KF; Wang, L; Soltwedel, T; Fidanze, SD; Hasvold, LA; Liu, D; Mantei, RA; Pratt, JK; Sheppard, GS; Bui, MH; Faivre, EJ; Huang, X; Li, L; Lin, X; Wang, R; Warder, SE; Wilcox, D; Albert, DH; Magoc, TJ; Rajaraman, G; Park, CH; Hutchins, CW; Shen, JJ; Edalji, RP; Sun, CC; Martin, R; Gao, W; Wong, S; Fang, G; Elmore, SW; Shen, Y; Kati, WM Discovery of N-(4-(2,4-Difluorophenoxy)-3-(6-methyl-7-oxo-6,7-dihydro-1H-pyrrolo[2,3-c]pyridin-4-yl)phenyl)ethanesulfonamide (ABBV-075/Mivebresib), a Potent and Orally Available Bromodomain and Extraterminal Domain (BET) Family Bromodomain Inhibitor. J Med Chem 60:8369-8384 (2017) [PubMed] Article More Info.:
Target
Name:
Histone acetyltransferase KAT2A
Synonyms:
GCN5 | GCN5 | GCN5L2 | General control of amino acid synthesis protein 5-like 2 | HGCN5 | Histone acetyltransferase GCN5 | Histone acetyltransferase KAT2A | Histone acetyltransferase KAT2A/KAT2B | HsGCN5 | KAT2A | KAT2A_HUMAN | Lysine acetyltransferase 2A | STAF97
Type:
PROTEIN
Mol. Mass.:
93956.22
Organism:
Homo sapiens (Human)
Description:
ChEMBL_100876
Residue:
837
Sequence:
MAEPSQAPTPAPAAQPRPLQSPAPAPTPTPAPSPASAPIPTPTPAPAPAPAAAPAGSTGTGGPGVGSGGAGSGGDPARPGLSQQQRASQRKAQVRGLPRAKKLEKLGVFSACKANETCKCNGWKNPKPPTAPRMDLQQPAANLSELCRSCEHPLADHVSHLENVSEDEINRLLGMVVDVENLFMSVHKEEDTDTKQVYFYLFKLLRKCILQMTRPVVEGSLGSPPFEKPNIEQGVLNFVQYKFSHLAPRERQTMFELSKMFLLCLNYWKLETPAQFRQRSQAEDVATYKVNYTRWLCYCHVPQSCDSLPRYETTHVFGRSLLRSIFTVTRRQLLEKFRVEKDKLVPEKRTLILTHFPKFLSMLEEEIYGANSPIWESGFTMPPSEGTQLVPRPASVSAAVVPSTPIFSPSMGGGSNSSLSLDSAGAEPMPGEKRTLPENLTLEDAKRLRVMGDIPMELVNEVMLTITDPAAMLGPETSLLSANAARDETARLEERRGIIEFHVIGNSLTPKANRRVLLWLVGLQNVFSHQLPRMPKEYIARLVFDPKHKTLALIKDGRVIGGICFRMFPTQGFTEIVFCAVTSNEQVKGYGTHLMNHLKEYHIKHNILYFLTYADEYAIGYFKKQGFSKDIKVPKSRYLGYIKDYEGATLMECELNPRIPYTELSHIIKKQKEIIKKLIERKQAQIRKVYPGLSCFKEGVRQIPVESVPGIRETGWKPLGKEKGKELKDPDQLYTTLKNLLAQIKSHPSAWPFMEPVKKSEAPDYYEVIRFPIDLKTMTERLRSRYYVTRKLFVADLQRVIANCREYNPPDSEYCRCASALEKFFYFKLKEGGLIDK