Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
E3 ubiquitin-protein ligase TRIM33
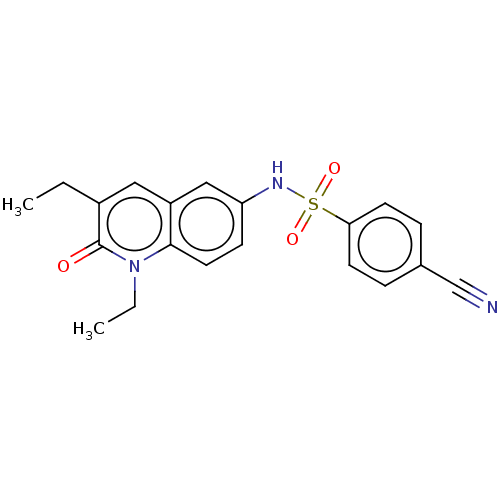
Ligand
BDBM50249797
Substrate
n/a
Meas. Tech.
ChEMBL_1701838 (CHEMBL4053071)
IC50
>100000±n/a nM
Citation
 Igoe, N; Bayle, ED; Fedorov, O; Tallant, C; Savitsky, P; Rogers, C; Owen, DR; Deb, G; Somervaille, TC; Andrews, DM; Jones, N; Cheasty, A; Ryder, H; Brennan, PE; M�ller, S; Knapp, S; Fish, PV Design of a Biased Potent Small Molecule Inhibitor of the Bromodomain and PHD Finger-Containing (BRPF) Proteins Suitable for Cellular and in Vivo Studies. J Med Chem 60:668-680 (2017) [PubMed] Article
Igoe, N; Bayle, ED; Fedorov, O; Tallant, C; Savitsky, P; Rogers, C; Owen, DR; Deb, G; Somervaille, TC; Andrews, DM; Jones, N; Cheasty, A; Ryder, H; Brennan, PE; M�ller, S; Knapp, S; Fish, PV Design of a Biased Potent Small Molecule Inhibitor of the Bromodomain and PHD Finger-Containing (BRPF) Proteins Suitable for Cellular and in Vivo Studies. J Med Chem 60:668-680 (2017) [PubMed] Article More Info.:
Target
Name:
E3 ubiquitin-protein ligase TRIM33
Synonyms:
6.3.2.- | E3 ubiquitin-protein ligase TRIM33 | Ectodermin homolog | KIAA1113 | KIAA1113 | Protein Rfg7 | RET-fused gene 7 protein | RFG7 | TIF1-gamma | TIF1G | TRI33_HUMAN | TRIM33 | Transcription intermediary factor 1-gamma | Tripartite motif-containing protein 33
Type:
PROTEIN
Mol. Mass.:
122534.89
Organism:
Homo sapiens (Human)
Description:
ChEMBL_105347
Residue:
1127
Sequence:
MAENKGGGEAESGGGGSGSAPVTAGAAGPAAQEAEPPLTAVLVEEEEEEGGRAGAEGGAAGPDDGGVAAASSGSAQAASSPAASVGTGVAGGAVSTPAPAPASAPAPGPSAGPPPGPPASLLDTCAVCQQSLQSRREAEPKLLPCLHSFCLRCLPEPERQLSVPIPGGSNGDIQQVGVIRCPVCRQECRQIDLVDNYFVKDTSEAPSSSDEKSEQVCTSCEDNASAVGFCVECGEWLCKTCIEAHQRVKFTKDHLIRKKEDVSESVGASGQRPVFCPVHKQEQLKLFCETCDRLTCRDCQLLEHKEHRYQFLEEAFQNQKGAIENLLAKLLEKKNYVHFAATQVQNRIKEVNETNKRVEQEIKVAIFTLINEINKKGKSLLQQLENVTKERQMKLLQQQNDITGLSRQVKHVMNFTNWAIASGSSTALLYSKRLITFQLRHILKARCDPVPAANGAIRFHCDPTFWAKNVVNLGNLVIESKPAPGYTPNVVVGQVPPGTNHISKTPGQINLAQLRLQHMQQQVYAQKHQQLQQMRMQQPPAPVPTTTTTTQQHPRQAAPQMLQQQPPRLISVQTMQRGNMNCGAFQAHQMRLAQNAARIPGIPRHSGPQYSMMQPHLQRQHSNPGHAGPFPVVSVHNTTINPTSPTTATMANANRGPTSPSVTAIELIPSVTNPENLPSLPDIPPIQLEDAGSSSLDNLLSRYISGSHLPPQPTSTMNPSPGPSALSPGSSGLSNSHTPVRPPSTSSTGSRGSCGSSGRTAEKTSLSFKSDQVKVKQEPGTEDEICSFSGGVKQEKTEDGRRSACMLSSPESSLTPPLSTNLHLESELDALASLENHVKIEPADMNESCKQSGLSSLVNGKSPIRSLMHRSARIGGDGNNKDDDPNEDWCAVCQNGGDLLCCEKCPKVFHLTCHVPTLLSFPSGDWICTFCRDIGKPEVEYDCDNLQHSKKGKTAQGLSPVDQRKCERLLLYLYCHELSIEFQEPVPASIPNYYKIIKKPMDLSTVKKKLQKKHSQHYQIPDDFVADVRLIFKNCERFNEMMKVVQVYADTQEINLKADSEVAQAGKAVALYFEDKLTEIYSDRTFAPLPEFEQEEDDGEVTEDSDEDFIQPRRKRLKSDERPVHIK