Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
BMP-2-inducible protein kinase
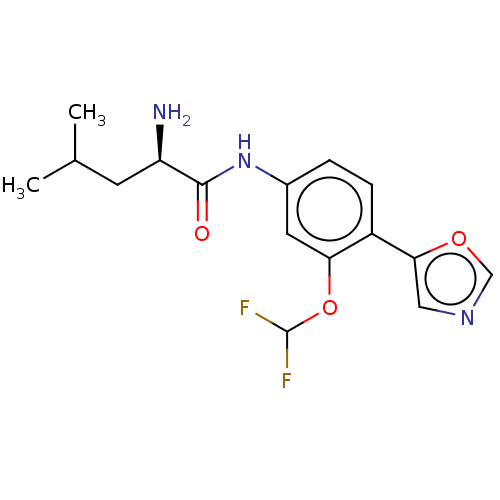
Ligand
BDBM50573682
Substrate
n/a
Meas. Tech.
ChEMBL_2120014 (CHEMBL4829161)
IC50
120±n/a nM
Citation
 Hartz, RA; Ahuja, VT; Nara, SJ; Kumar, CMV; Brown, JM; Bristow, LJ; Rajamani, R; Muckelbauer, JK; Camac, D; Kiefer, SE; Hunihan, L; Gulianello, M; Lewis, M; Easton, A; Lippy, JS; Surti, N; Pattipati, SN; Dokania, M; Elavazhagan, S; Dandapani, K; Hamman, BD; Allen, J; Kostich, W; Bronson, JJ; Macor, JE; Dzierba, CD Discovery, Structure-Activity Relationships, and In Vivo Evaluation of Novel Aryl Amides as Brain Penetrant Adaptor Protein 2-Associated Kinase 1 (AAK1) Inhibitors for the Treatment of Neuropathic Pain. J Med Chem 64:11090-11128 (2021) [PubMed] Article
Hartz, RA; Ahuja, VT; Nara, SJ; Kumar, CMV; Brown, JM; Bristow, LJ; Rajamani, R; Muckelbauer, JK; Camac, D; Kiefer, SE; Hunihan, L; Gulianello, M; Lewis, M; Easton, A; Lippy, JS; Surti, N; Pattipati, SN; Dokania, M; Elavazhagan, S; Dandapani, K; Hamman, BD; Allen, J; Kostich, W; Bronson, JJ; Macor, JE; Dzierba, CD Discovery, Structure-Activity Relationships, and In Vivo Evaluation of Novel Aryl Amides as Brain Penetrant Adaptor Protein 2-Associated Kinase 1 (AAK1) Inhibitors for the Treatment of Neuropathic Pain. J Med Chem 64:11090-11128 (2021) [PubMed] Article More Info.:
Target
Name:
BMP-2-inducible protein kinase
Synonyms:
BIKE | BMP-2-inducible protein kinase | BMP2K | BMP2K_HUMAN
Type:
PROTEIN
Mol. Mass.:
129169.50
Organism:
Homo sapiens (Human)
Description:
ChEMBL_774589
Residue:
1161
Sequence:
MKKFSRMPKSEGGSGGGAAGGGAGGAGAGAGCGSGGSSVGVRVFAVGRHQVTLEESLAEGGFSTVFLVRTHGGIRCALKRMYVNNMPDLNVCKREITIMKELSGHKNIVGYLDCAVNSISDNVWEVLILMEYCRAGQVVNQMNKKLQTGFTEPEVLQIFCDTCEAVARLHQCKTPIIHRDLKVENILLNDGGNYVLCDFGSATNKFLNPQKDGVNVVEEEIKKYTTLSYRAPEMINLYGGKPITTKADIWALGCLLYKLCFFTLPFGESQVAICDGNFTIPDNSRYSRNIHCLIRFMLEPDPEHRPDIFQVSYFAFKFAKKDCPVSNINNSSIPSALPEPMTASEAAARKSQIKARITDTIGPTETSIAPRQRPKANSATTATPSVLTIQSSATPVKVLAPGEFGNHRPKGALRPGNGPEILLGQGPPQQPPQQHRVLQQLQQGDWRLQQLHLQHRHPHQQQQQQQQQQQQQQQQQQQQQQQQQQQHHHHHHHHLLQDAYMQQYQHATQQQQMLQQQFLMHSVYQPQPSASQYPTMMPQYQQAFFQQQMLAQHQPSQQQASPEYLTSPQEFSPALVSYTSSLPAQVGTIMDSSYSANRSVADKEAIANFTNQKNISNPPDMSGWNPFGEDNFSKLTEEELLDREFDLLRSNRLEERASSDKNVDSLSAPHNHPPEDPFGSVPFISHSGSPEKKAEHSSINQENGTANPIKNGKTSPASKDQRTGKKTSVQGQVQKGNDESESDFESDPPSPKSSEEEEQDDEEVLQGEQGDFNDDDTEPENLGHRPLLMDSEDEEEEEKHSSDSDYEQAKAKYSDMSSVYRDRSGSGPTQDLNTILLTSAQLSSDVAVETPKQEFDVFGAVPFFAVRAQQPQQEKNEKNLPQHRFPAAGLEQEEFDVFTKAPFSKKVNVQECHAVGPEAHTIPGYPKSVDVFGSTPFQPFLTSTSKSESNEDLFGLVPFDEITGSQQQKVKQRSLQKLSSRQRRTKQDMSKSNGKRHHGTPTSTKKTLKPTYRTPERARRHKKVGRRDSQSSNEFLTISDSKENISVALTDGKDRGNVLQPEESLLDPFGAKPFHSPDLSWHPPHQGLSDIRADHNTVLPGRPRQNSLHGSFHSADVLKMDDFGAVPFTELVVQSITPHQSQQSQPVELDPFGAAPFPSKQ