Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
cAMP-dependent protein kinase catalytic subunit alpha
Ligand
BDBM13530
Substrate
n/a
Meas. Tech.
ChEMBL_587098 (CHEMBL1051309)
Kd
>10000±n/a nM
Citation
 Karaman, MW; Herrgard, S; Treiber, DK; Gallant, P; Atteridge, CE; Campbell, BT; Chan, KW; Ciceri, P; Davis, MI; Edeen, PT; Faraoni, R; Floyd, M; Hunt, JP; Lockhart, DJ; Milanov, ZV; Morrison, MJ; Pallares, G; Patel, HK; Pritchard, S; Wodicka, LM; Zarrinkar, PP A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol 26:127-32 (2008) [PubMed] Article
Karaman, MW; Herrgard, S; Treiber, DK; Gallant, P; Atteridge, CE; Campbell, BT; Chan, KW; Ciceri, P; Davis, MI; Edeen, PT; Faraoni, R; Floyd, M; Hunt, JP; Lockhart, DJ; Milanov, ZV; Morrison, MJ; Pallares, G; Patel, HK; Pritchard, S; Wodicka, LM; Zarrinkar, PP A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol 26:127-32 (2008) [PubMed] Article More Info.:
Target
Name:
cAMP-dependent protein kinase catalytic subunit alpha
Synonyms:
KAPCA_HUMAN | PKA C-alpha | PKACA | PRKACA | cAMP-dependent protein kinase (PKA) | cAMP-dependent protein kinase catalytic (PKA) | cAMP-dependent protein kinase catalytic subunit alpha | cAMP-dependent protein kinase catalytic subunit alpha (PKA) | cAMP-dependent protein kinase catalytic subunit alpha (PKACA) | cAMP-dependent protein kinase catalytic subunit alpha (PKAc) | cAMP-dependent protein kinase catalytic subunit alpha isoform 1 | cAMP-dependent protein kinase, alpha-catalytic subunit
Type:
Enzyme Catalytic Subunit
Mol. Mass.:
40598.73
Organism:
Homo sapiens (Human)
Description:
n/a
Residue:
351
Sequence:
MGNAAAAKKGSEQESVKEFLAKAKEDFLKKWESPAQNTAHLDQFERIKTLGTGSFGRVMLVKHKETGNHYAMKILDKQKVVKLKQIEHTLNEKRILQAVNFPFLVKLEFSFKDNSNLYMVMEYVPGGEMFSHLRRIGRFSEPHARFYAAQIVLTFEYLHSLDLIYRDLKPENLLIDQQGYIQVTDFGFAKRVKGRTWTLCGTPEYLAPEIILSKGYNKAVDWWALGVLIYEMAAGYPPFFADQPIQIYEKIVSGKVRFPSHFSSDLKDLLRNLLQVDLTKRFGNLKNGVNDIKNHKWFATTDWIAIYQRKVEAPFIPKFKGPGDTSNFDDYEEEEIRVSINEKCGKEFSEF
Inhibitor
Name:
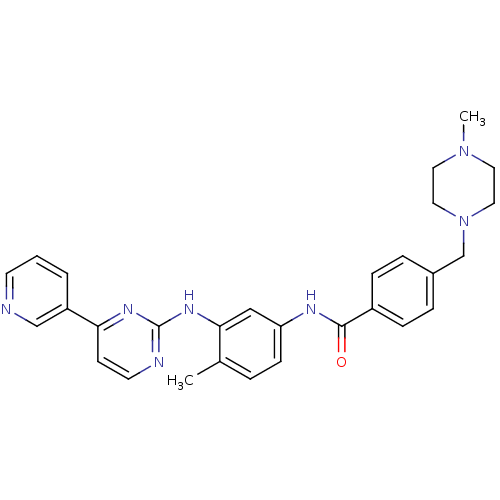
BDBM13530
Synonyms:
4-[(4-methylpiperazin-1-yl)methyl]-N-[4-methyl-3-[(4-pyridin-3-ylpyrimidin-2-yl)amino]phenyl]benzamide | CHEMBL941 | Gleevec | Imatinib | Imatinib, 21 | N-(4-methyl-3-{[4-(pyridin-3-yl)pyrimidin-2-yl]amino}phenyl)-4-[(4-methylpiperazin-1-yl)methyl]benzamide | STI-571 | STI571 | US10906896, Cpd imatinib | US11649218, Example Imatinib | US11725005, Compound imatinib | US9255107, Imatinib | cid_5291 | imatinib-CD3 | med.21724, Compound 6
Type:
Small organic molecule
Emp. Form.:
C29H31N7O
Mol. Mass.:
493.6027
SMILES:
CN1CCN(Cc2ccc(cc2)C(=O)Nc2ccc(C)c(Nc3nccc(n3)-c3cccnc3)c2)CC1