Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Mitogen-activated protein kinase kinase kinase kinase 4
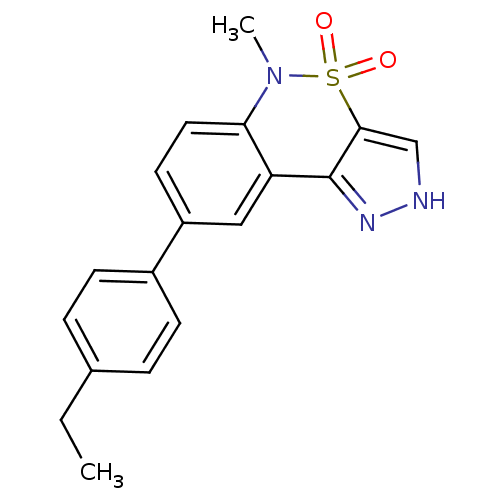
Ligand
BDBM50430289
Substrate
n/a
Meas. Tech.
ChEMBL_951385 (CHEMBL2349759)
IC50
1600±n/a nM
Citation
 Iwatani, M; Iwata, H; Okabe, A; Skene, RJ; Tomita, N; Hayashi, Y; Aramaki, Y; Hosfield, DJ; Hori, A; Baba, A; Miki, H Discovery and characterization of novel allosteric FAK inhibitors. Eur J Med Chem 61:49-60 (2013) [PubMed] Article
Iwatani, M; Iwata, H; Okabe, A; Skene, RJ; Tomita, N; Hayashi, Y; Aramaki, Y; Hosfield, DJ; Hori, A; Baba, A; Miki, H Discovery and characterization of novel allosteric FAK inhibitors. Eur J Med Chem 61:49-60 (2013) [PubMed] Article More Info.:
Target
Name:
Mitogen-activated protein kinase kinase kinase kinase 4
Synonyms:
HGK | HPK/GCK-like kinase HGK | KIAA0687 | M4K4_HUMAN | MAP4K4 | MAP4K4 (HGK) | MAPK/ERK kinase kinase kinase 4 | MEK kinase kinase 4 | MEKKK 4 | Mitogen-activated protein kinase kinase kinase kinase 4 (MAP4K4) | NIK | Nck-interacting kinase
Type:
Serine/threonine-protein kinase
Mol. Mass.:
142114.73
Organism:
Homo sapiens (Human)
Description:
O95819
Residue:
1239
Sequence:
MANDSPAKSLVDIDLSSLRDPAGIFELVEVVGNGTYGQVYKGRHVKTGQLAAIKVMDVTEDEEEEIKLEINMLKKYSHHRNIATYYGAFIKKSPPGHDDQLWLVMEFCGAGSITDLVKNTKGNTLKEDWIAYISREILRGLAHLHIHHVIHRDIKGQNVLLTENAEVKLVDFGVSAQLDRTVGRRNTFIGTPYWMAPEVIACDENPDATYDYRSDLWSCGITAIEMAEGAPPLCDMHPMRALFLIPRNPPPRLKSKKWSKKFFSFIEGCLVKNYMQRPSTEQLLKHPFIRDQPNERQVRIQLKDHIDRTRKKRGEKDETEYEYSGSEEEEEEVPEQEGEPSSIVNVPGESTLRRDFLRLQQENKERSEALRRQQLLQEQQLREQEEYKRQLLAERQKRIEQQKEQRRRLEEQQRREREARRQQEREQRRREQEEKRRLEELERRRKEEEERRRAEEEKRRVEREQEYIRRQLEEEQRHLEVLQQQLLQEQAMLLECRWREMEEHRQAERLQRQLQQEQAYLLSLQHDHRRPHPQHSQQPPPPQQERSKPSFHAPEPKAHYEPADRAREVEDRFRKTNHSSPEAQSKQTGRVLEPPVPSRSESFSNGNSESVHPALQRPAEPQVPVRTTSRSPVLSRRDSPLQGSGQQNSQAGQRNSTSIEPRLLWERVEKLVPRPGSGSSSGSSNSGSQPGSHPGSQSGSGERFRVRSSSKSEGSPSQRLENAVKKPEDKKEVFRPLKPADLTALAKELRAVEDVRPPHKVTDYSSSSEESGTTDEEDDDVEQEGADESTSGPEDTRAASSLNLSNGETESVKTMIVHDDVESEPAMTPSKEGTLIVRQTQSASSTLQKHKSSSSFTPFIDPRLLQISPSSGTTVTSVVGFSCDGMRPEAIRQDPTRKGSVVNVNPTNTRPQSDTPEIRKYKKRFNSEILCAALWGVNLLVGTESGLMLLDRSGQGKVYPLINRRRFQQMDVLEGLNVLVTISGKKDKLRVYYLSWLRNKILHNDPEVEKKQGWTTVGDLEGCVHYKVVKYERIKFLVIALKSSVEVYAWAPKPYHKFMAFKSFGELVHKPLLVDLTVEEGQRLKVIYGSCAGFHAVDVDSGSVYDIYLPTHIQCSIKPHAIIILPNTDGMELLVCYEDEGVYVNTYGRITKDVVLQWGEMPTSVAYIRSNQTMGWGEKAIEIRSVETGHLDGVFMHKRAQRLKFLCERNDKVFFASVRSGGSSQVYFMTLGRTSLLSW