Reaction Details Report a problem with these data
Report a problem with these data
 Report a problem with these data
Report a problem with these dataTarget
Histone acetyltransferase KAT2A
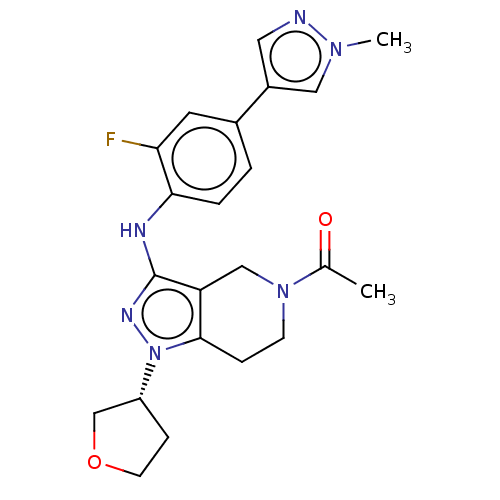
Ligand
BDBM50200384
Substrate
n/a
Meas. Tech.
ChEBML_1622061
IC50
>20000±n/a nM
Citation
 Crawford, TD; Romero, FA; Lai, KW; Tsui, V; Taylor, AM; de Leon Boenig, G; Noland, CL; Murray, J; Ly, J; Choo, EF; Hunsaker, TL; Chan, EW; Merchant, M; Kharbanda, S; Gascoigne, KE; Kaufman, S; Beresini, MH; Liao, J; Liu, W; Chen, KX; Chen, Z; Conery, AR; C�t�, A; Jayaram, H; Jiang, Y; Kiefer, JR; Kleinheinz, T; Li, Y; Maher, J; Pardo, E; Poy, F; Spillane, KL; Wang, F; Wang, J; Wei, X; Xu, Z; Xu, Z; Yen, I; Zawadzke, L; Zhu, X; Bellon, S; Cummings, R; Cochran, AG; Albrecht, BK; Magnuson, S Discovery of a Potent and Selective in Vivo Probe (GNE-272) for the Bromodomains of CBP/EP300. J Med Chem 59:10549-10563 (2016) [PubMed] Article
Crawford, TD; Romero, FA; Lai, KW; Tsui, V; Taylor, AM; de Leon Boenig, G; Noland, CL; Murray, J; Ly, J; Choo, EF; Hunsaker, TL; Chan, EW; Merchant, M; Kharbanda, S; Gascoigne, KE; Kaufman, S; Beresini, MH; Liao, J; Liu, W; Chen, KX; Chen, Z; Conery, AR; C�t�, A; Jayaram, H; Jiang, Y; Kiefer, JR; Kleinheinz, T; Li, Y; Maher, J; Pardo, E; Poy, F; Spillane, KL; Wang, F; Wang, J; Wei, X; Xu, Z; Xu, Z; Yen, I; Zawadzke, L; Zhu, X; Bellon, S; Cummings, R; Cochran, AG; Albrecht, BK; Magnuson, S Discovery of a Potent and Selective in Vivo Probe (GNE-272) for the Bromodomains of CBP/EP300. J Med Chem 59:10549-10563 (2016) [PubMed] Article More Info.:
Target
Name:
Histone acetyltransferase KAT2A
Synonyms:
GCN5 | GCN5 | GCN5L2 | General control of amino acid synthesis protein 5-like 2 | HGCN5 | Histone acetyltransferase GCN5 | Histone acetyltransferase KAT2A | Histone acetyltransferase KAT2A/KAT2B | HsGCN5 | KAT2A | KAT2A_HUMAN | Lysine acetyltransferase 2A | STAF97
Type:
PROTEIN
Mol. Mass.:
93956.22
Organism:
Homo sapiens (Human)
Description:
ChEMBL_100876
Residue:
837
Sequence:
MAEPSQAPTPAPAAQPRPLQSPAPAPTPTPAPSPASAPIPTPTPAPAPAPAAAPAGSTGTGGPGVGSGGAGSGGDPARPGLSQQQRASQRKAQVRGLPRAKKLEKLGVFSACKANETCKCNGWKNPKPPTAPRMDLQQPAANLSELCRSCEHPLADHVSHLENVSEDEINRLLGMVVDVENLFMSVHKEEDTDTKQVYFYLFKLLRKCILQMTRPVVEGSLGSPPFEKPNIEQGVLNFVQYKFSHLAPRERQTMFELSKMFLLCLNYWKLETPAQFRQRSQAEDVATYKVNYTRWLCYCHVPQSCDSLPRYETTHVFGRSLLRSIFTVTRRQLLEKFRVEKDKLVPEKRTLILTHFPKFLSMLEEEIYGANSPIWESGFTMPPSEGTQLVPRPASVSAAVVPSTPIFSPSMGGGSNSSLSLDSAGAEPMPGEKRTLPENLTLEDAKRLRVMGDIPMELVNEVMLTITDPAAMLGPETSLLSANAARDETARLEERRGIIEFHVIGNSLTPKANRRVLLWLVGLQNVFSHQLPRMPKEYIARLVFDPKHKTLALIKDGRVIGGICFRMFPTQGFTEIVFCAVTSNEQVKGYGTHLMNHLKEYHIKHNILYFLTYADEYAIGYFKKQGFSKDIKVPKSRYLGYIKDYEGATLMECELNPRIPYTELSHIIKKQKEIIKKLIERKQAQIRKVYPGLSCFKEGVRQIPVESVPGIRETGWKPLGKEKGKELKDPDQLYTTLKNLLAQIKSHPSAWPFMEPVKKSEAPDYYEVIRFPIDLKTMTERLRSRYYVTRKLFVADLQRVIANCREYNPPDSEYCRCASALEKFFYFKLKEGGLIDK