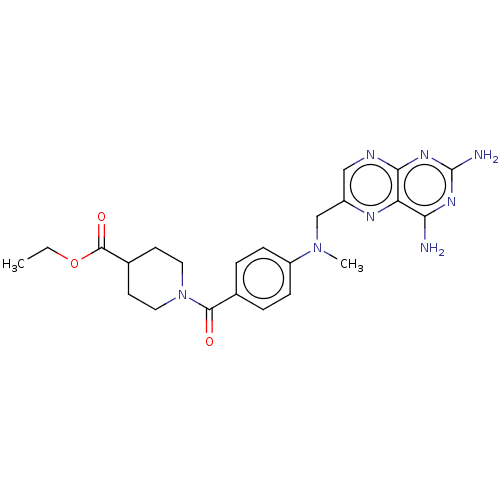
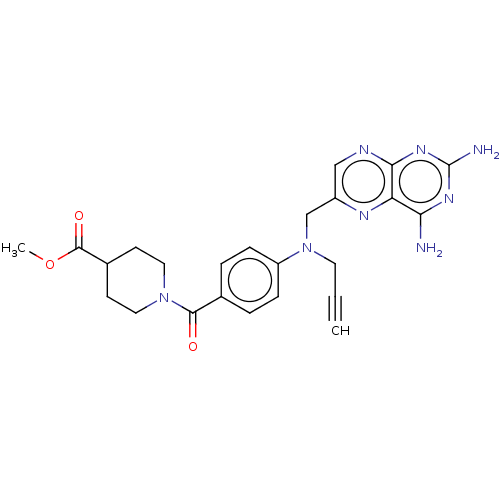
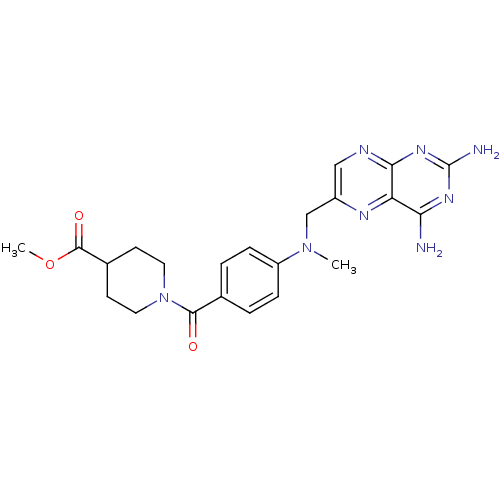
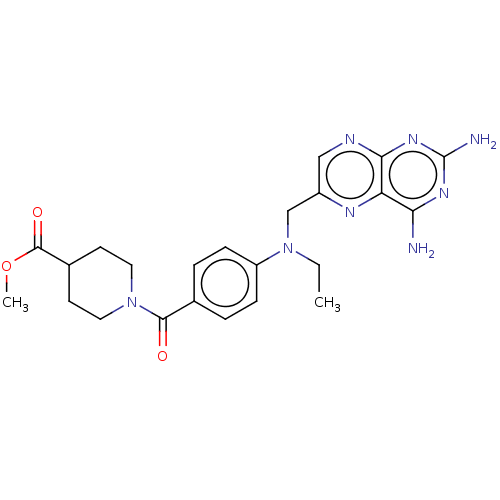
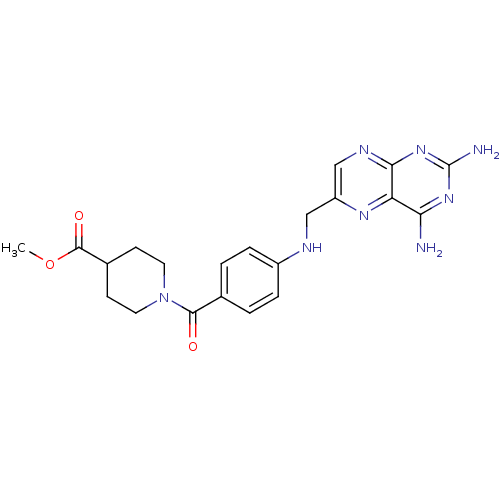
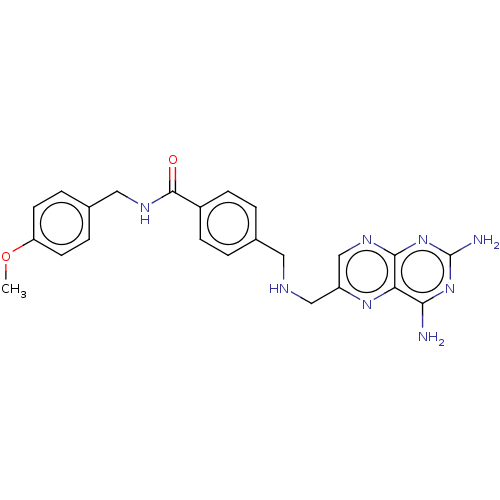
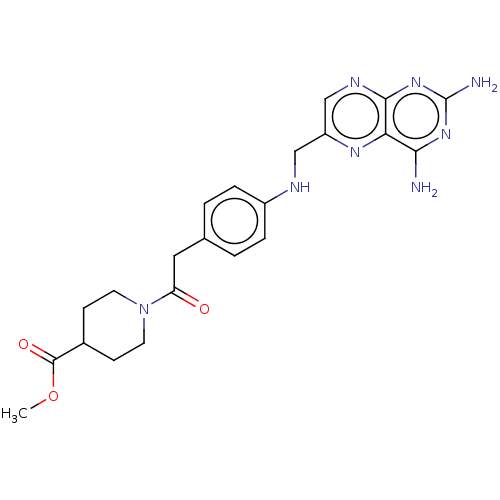
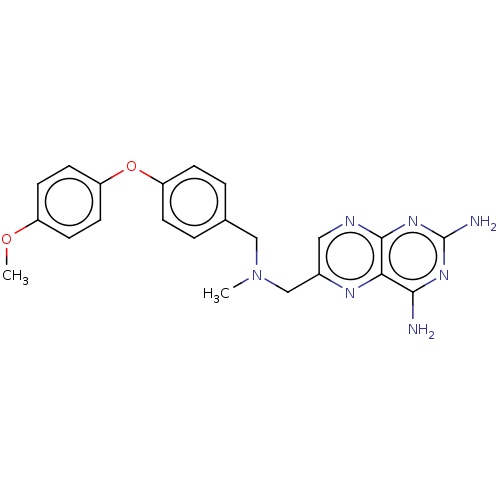
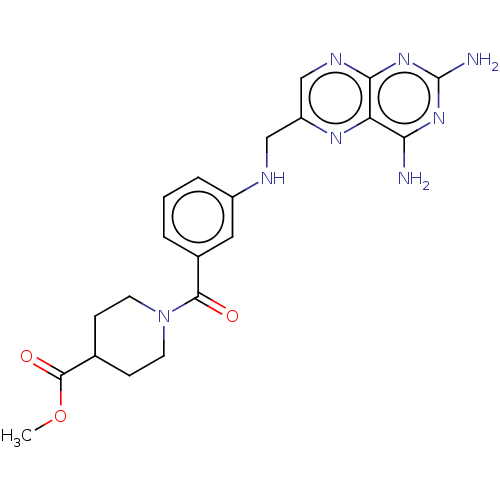
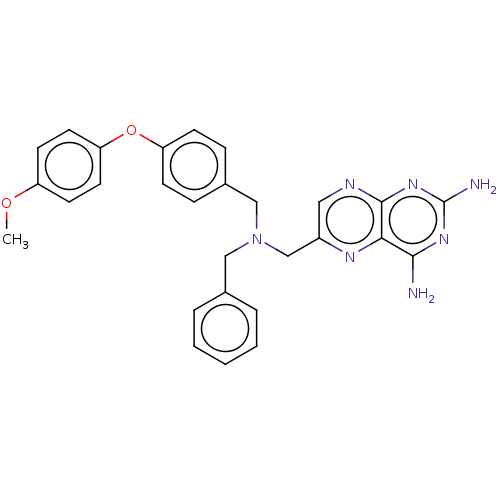
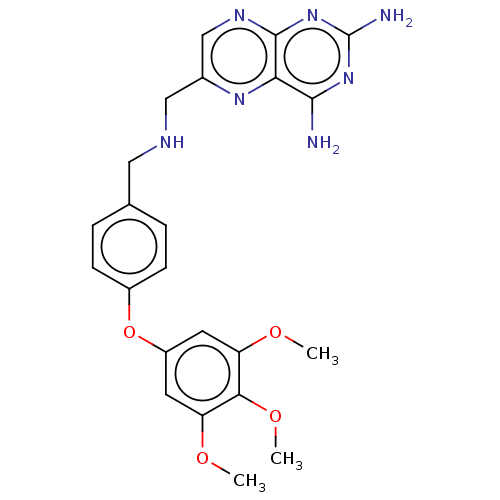
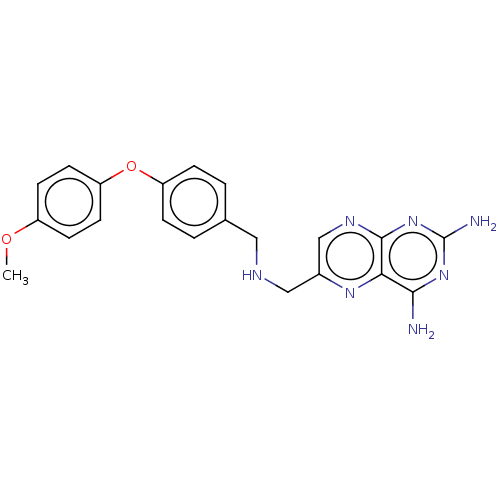
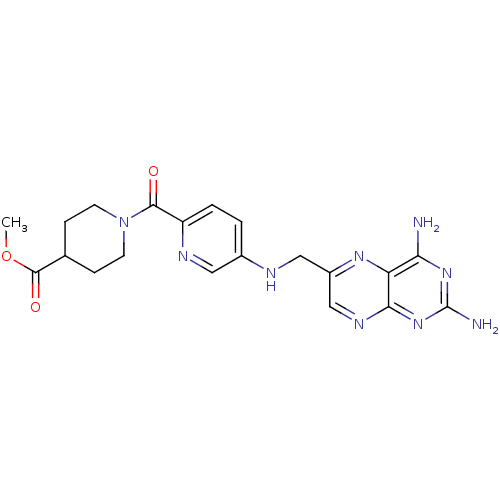
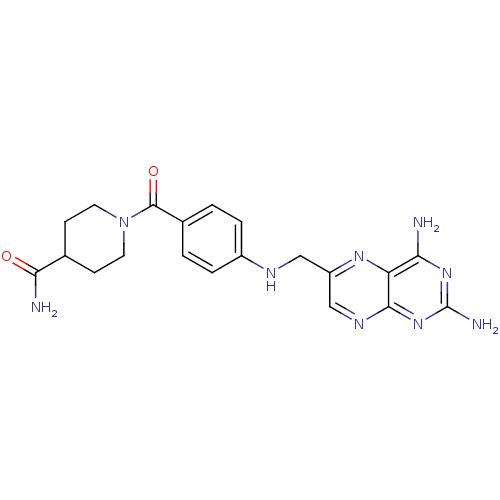
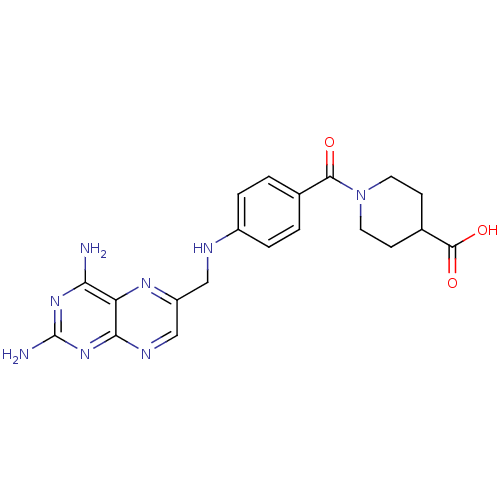
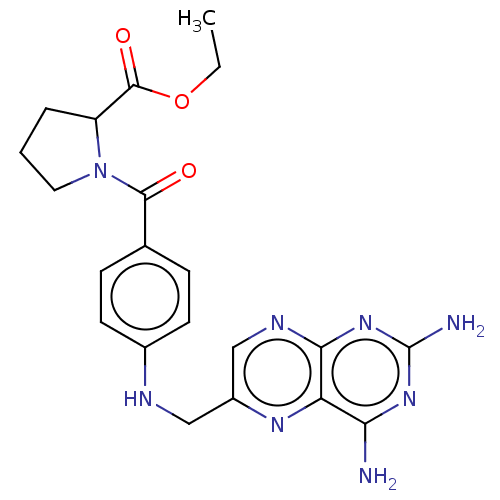
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
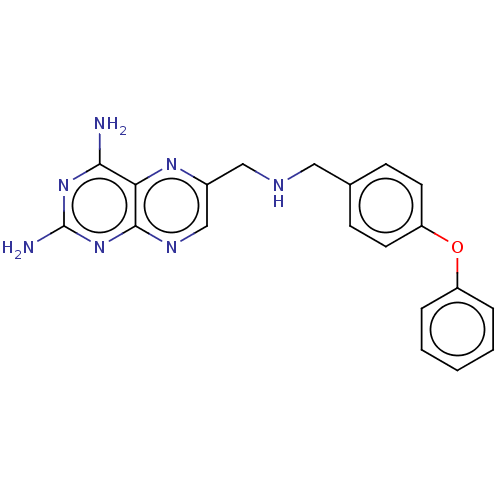
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
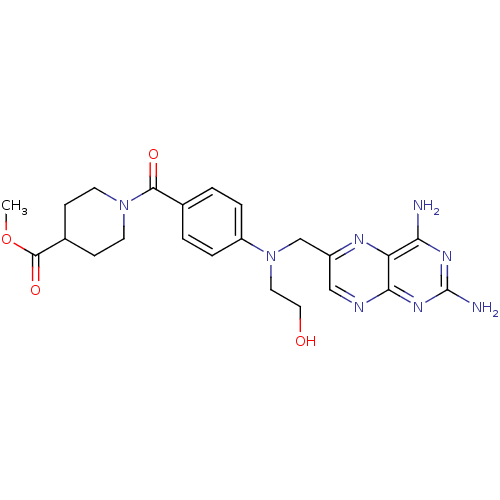
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 0.100nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
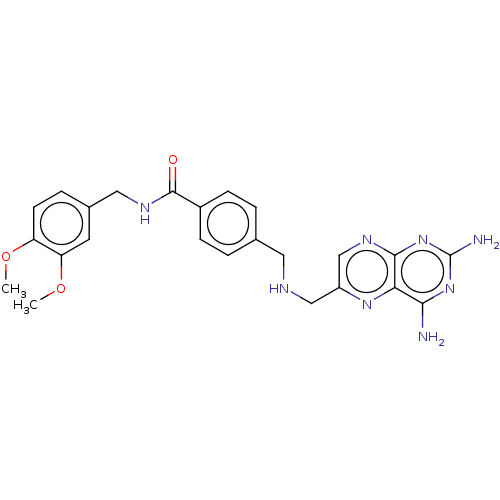
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 30nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 30nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 40nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 40nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 50nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 50nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 90nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 140nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 150nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 150nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 190nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 220nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 240nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 350nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 520nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 550nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 820nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 1.72E+3nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
TargetPteridine reductase, putative(Trypanosoma brucei brucei (strain 927/4 GUTat10.1))
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Heidelberg Institute For Theoretical Studies (Hits)
Curated by ChEMBL
Affinity DataIC50: 1.33E+5nMAssay Description:Inhibition of recombinant Trypanosoma brucei PTR1 expressed in Escherichia coli BL21 (DE3) cells using H2B as substrate in presence of cytochrome c a...More data for this Ligand-Target Pair
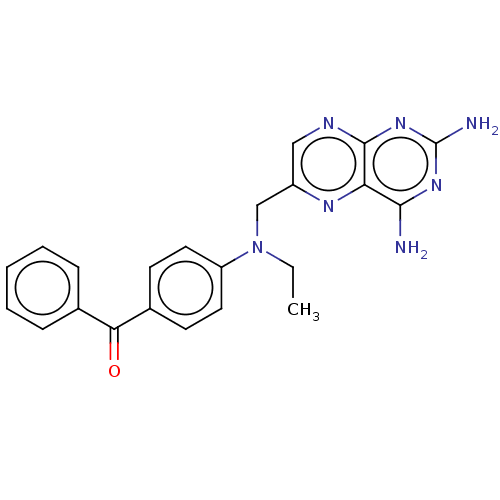
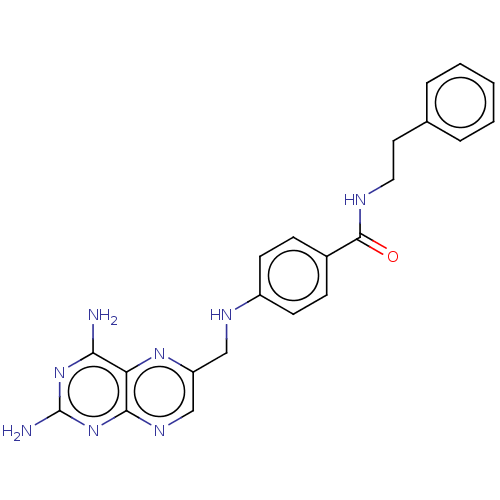
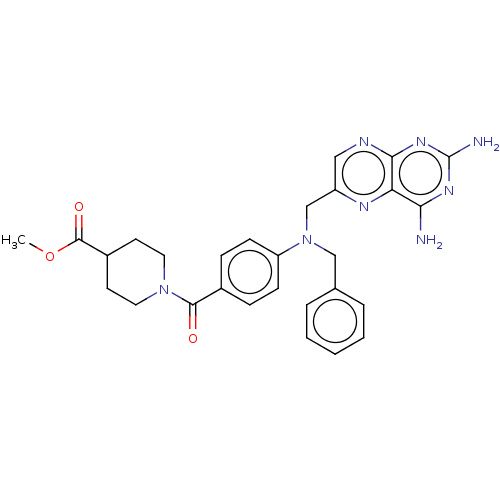
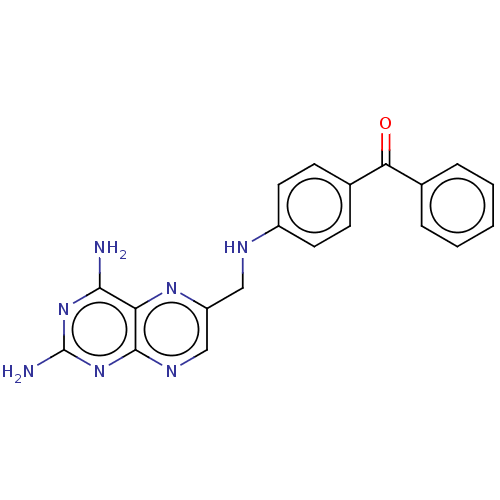
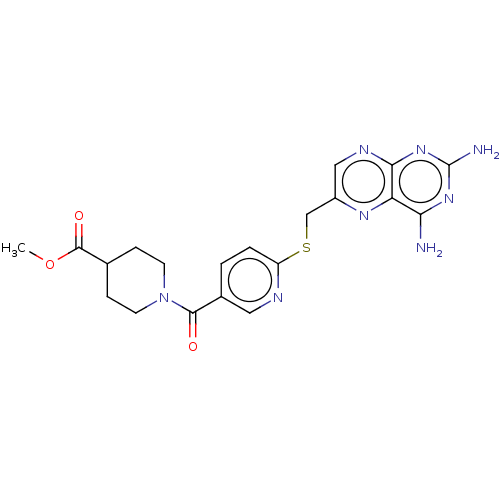
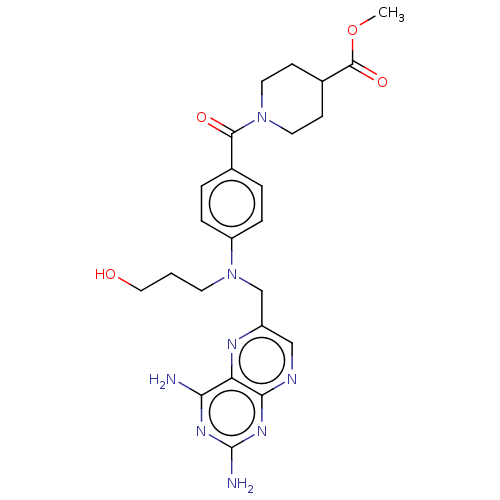
 3D Structure (crystal)
3D Structure (crystal)