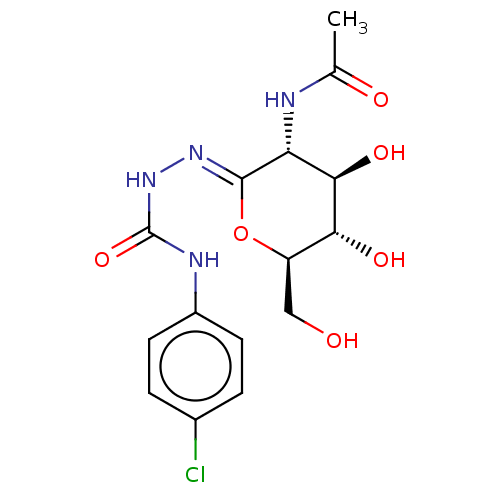
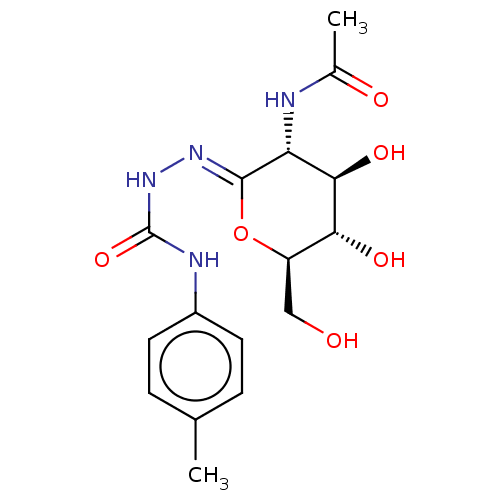
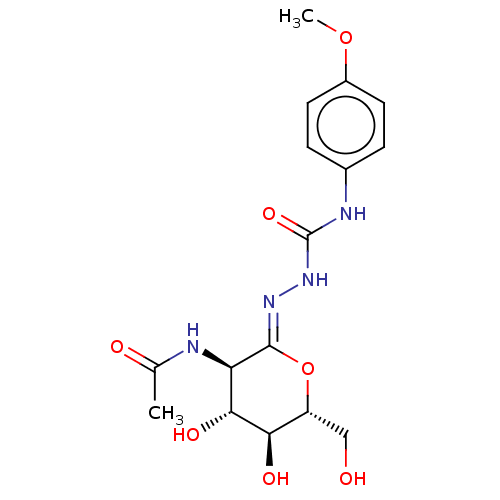
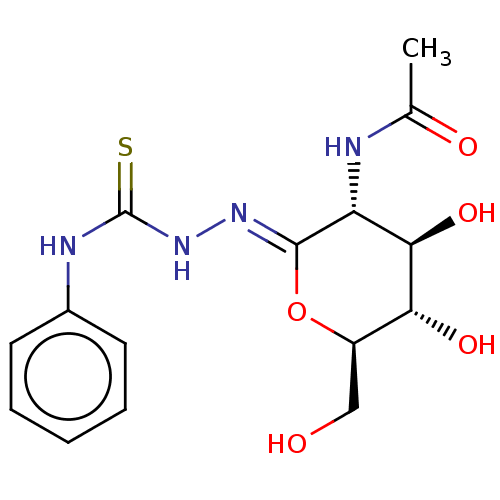
Affinity DataKi: 47nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 125nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 130nMAssay Description:Inhibition of HexA/HexB (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 154nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 170nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 205nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 332nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 413nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMAssay Description:Competitive inhibition of cow HexA/HexB (unknown origin) using pNP-GlcNAc as substrate assessed as inhibition constant by measuring liberation of nit...More data for this Ligand-Target Pair