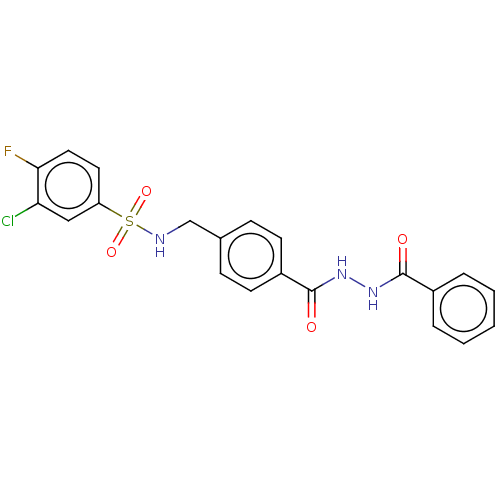
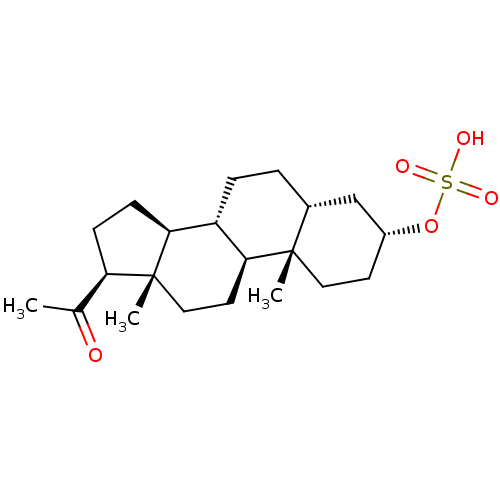
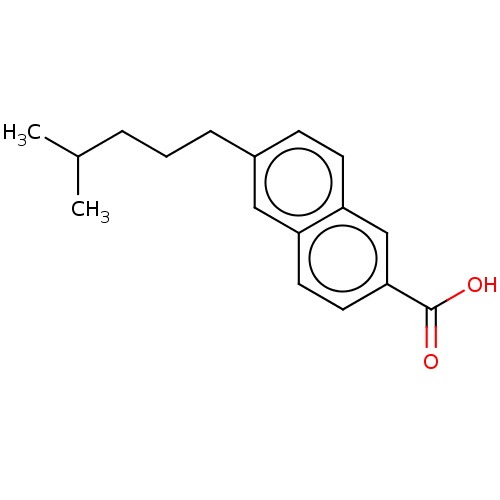
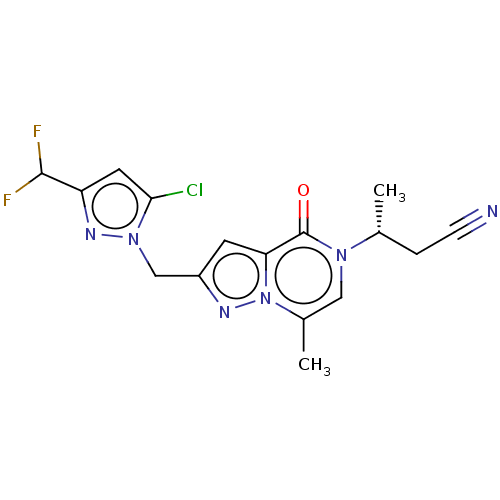
Affinity DataEC50: 430nMAssay Description:Positive allosteric modulation of recombinant human GluN1/GluN2D receptor stably expressed in HEK293 cells assessed as increase in glycine/L-glutamat...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of recombinant GluN1/GluN2D receptor (unknown origin) expressed in HEK293 cells assessed as inhibition of glutamate-evoked current by whol...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Inhibition of NMDA GluN2D receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataEC50: 2.80E+3nMAssay Description:Positive allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as increase in glycine-induced channe...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90E+3nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataEC50: 5.10E+3nMAssay Description:Positive allosteric modulator activity at GluN2D NMDA receptor (unknown origin) expressed in CHO cells co-expressing GluN1a in the presence of L-glut...More data for this Ligand-Target Pair
Affinity DataEC50: 6.20E+3nMAssay Description:Positive allosteric modulation of GluN1/GluN2D NMDAR (unknown origin) expressed in Dox-inducible cells by BD calcium indicator dye based-fluorescence...More data for this Ligand-Target Pair
Affinity DataEC50: 6.70E+3nMAssay Description:Positive allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as increase in glycine-induced channe...More data for this Ligand-Target Pair
Affinity DataEC50: 9.50E+3nMAssay Description:Positive allosteric modulation of GluN1/GluN2D receptor (unknown origin) assessed as increase in glutamate-induced calcium flux measured at time inte...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulation of recombinant human GluN1a/GluN2D expressed in CHO-T-REx cells assessed as inhibition of glutamate/glycine-induced re...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulation of recombinant human GluN1a/GluN2D expressed in CHO-T-REx cells assessed as inhibition of glutamate/glycine-induced re...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human GluN2D receptor by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human GluN2D receptor by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human GluN2D receptor by FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+4nMAssay Description:Positive allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as increase in glycine-induced channe...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulation of human GluN2D receptor expressed in xenopus laevis oocytes assessed as reduction in 3 uM glycine-induced channel cur...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataEC50: 2.90E+4nMAssay Description:Positive allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as increase in glycine-induced channe...More data for this Ligand-Target Pair
Affinity DataEC50: >3.00E+4nMAssay Description:Positive allosteric modulator activity at GluN1a/GluN2D (unknown origin) expressed in CHO cells in presence of glutamate by Ca2+ influx assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Negative allosteric modulation of human GluN2D receptor expressed in HEK cells assessed as reduction in glycine/glutamate-induced intracellular calci...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataIC50: 7.59E+4nMAssay Description:Negative allosteric modulation of human GluN2D receptor expressed in xenopus laevis oocytes assessed as reduction in 3 uM glycine-induced channel cur...More data for this Ligand-Target Pair
Affinity DataEC50: 7.80E+4nMAssay Description:Positive allosteric modulation of EGFP-fused human GluN2D receptor expressed in HEK293T cells assessed as increase in glycine-induced channel current...More data for this Ligand-Target Pair
Affinity DataIC50: 1.18E+5nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataIC50: 4.26E+5nMAssay Description:Negative allosteric modulation of GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glycine-induced chann...More data for this Ligand-Target Pair
Affinity DataIC50: 4.26E+5nMAssay Description:Negative allosteric modulation of GluN1a/GluN2D receptor (unknown origin) expressed in xenopus laevis oocytes assessed as reduction in glutamate/glyc...More data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)