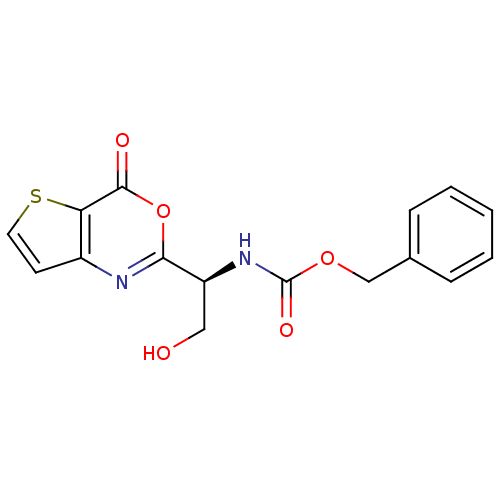
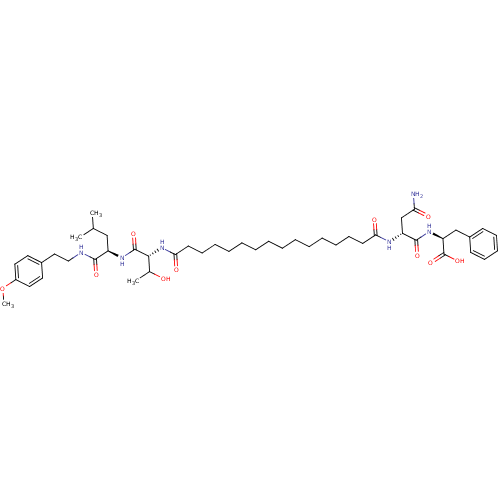
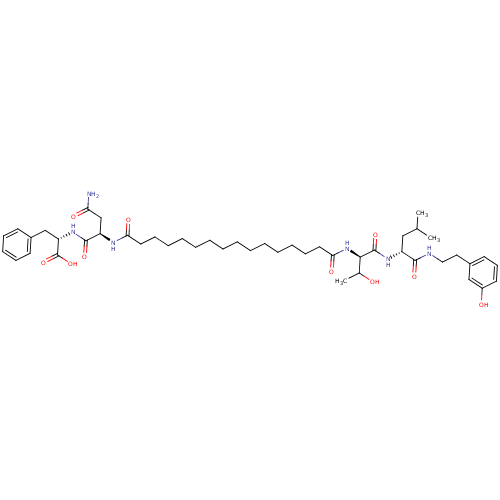
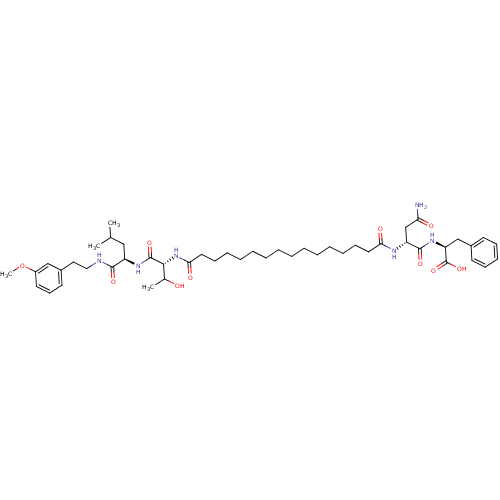
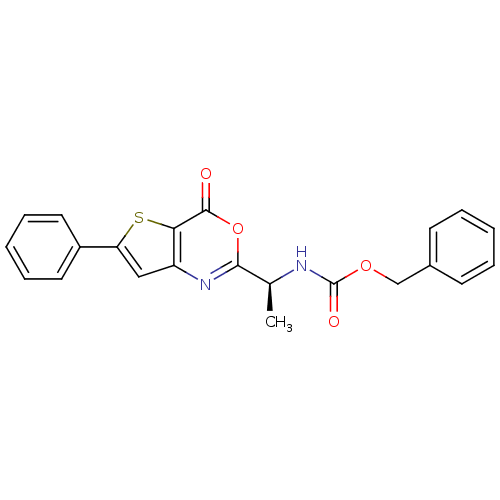
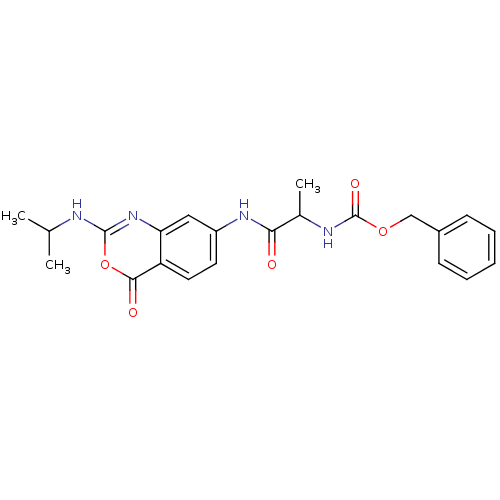
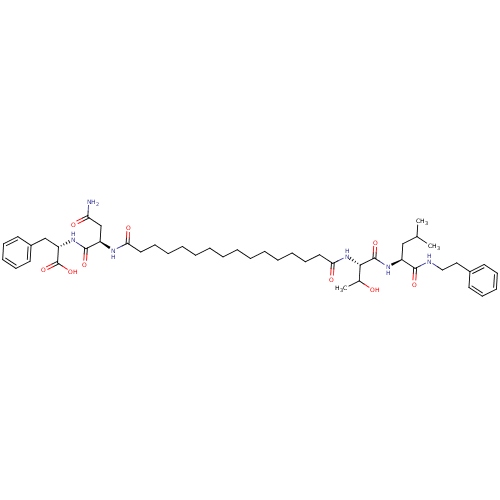
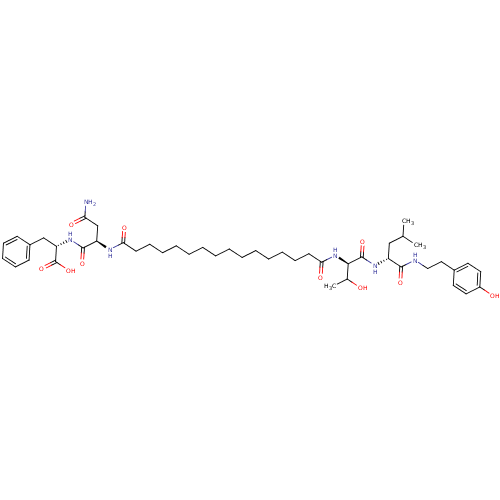
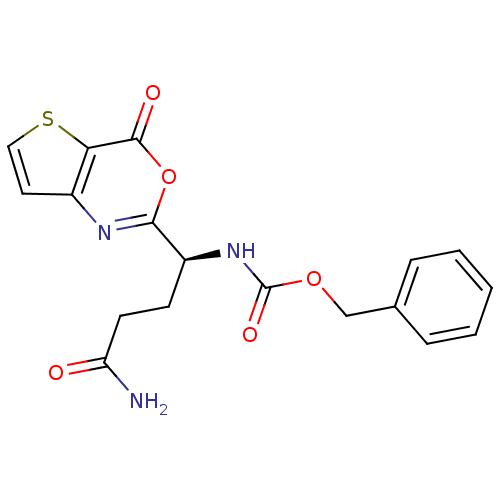
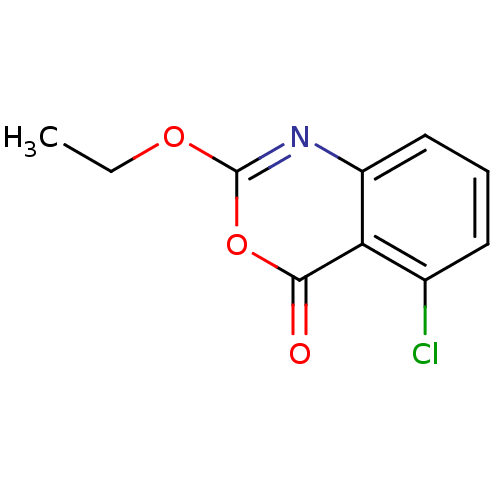
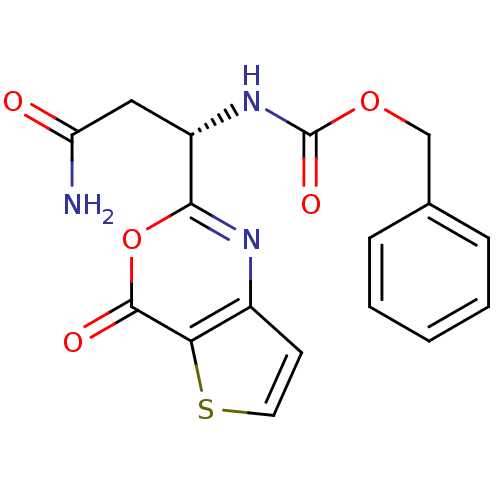
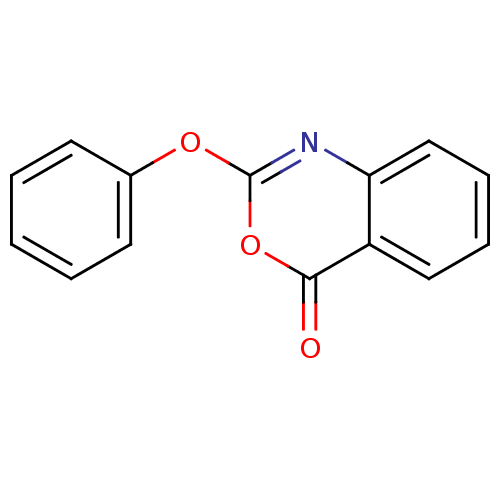
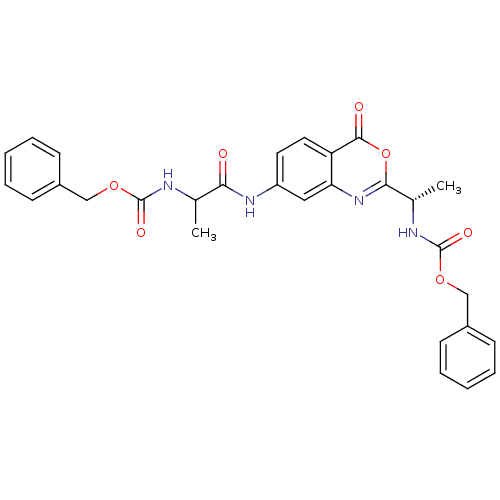
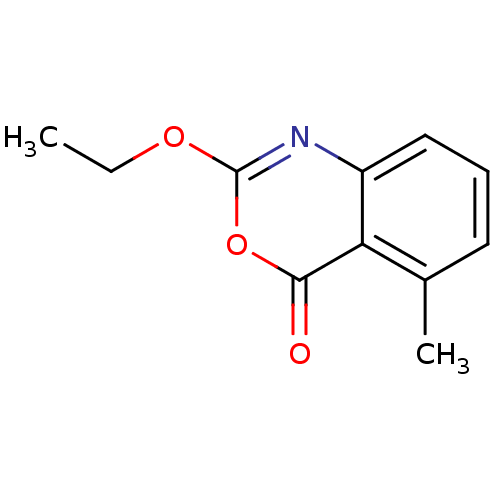
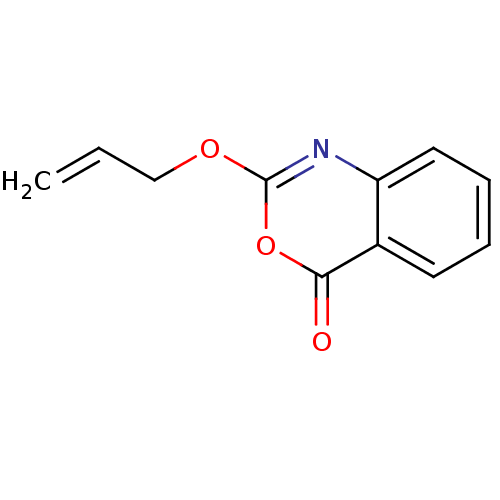
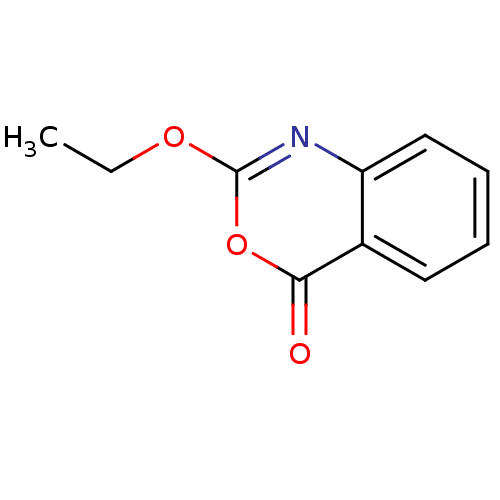
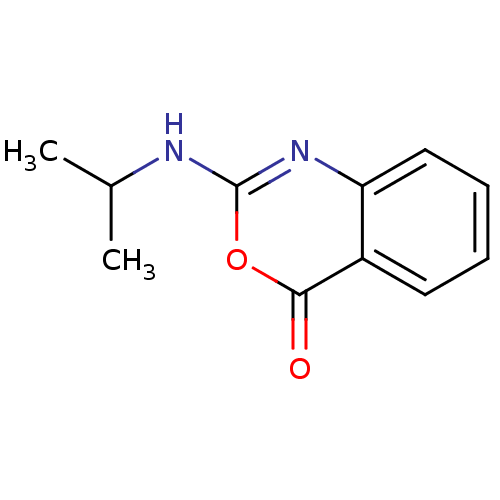
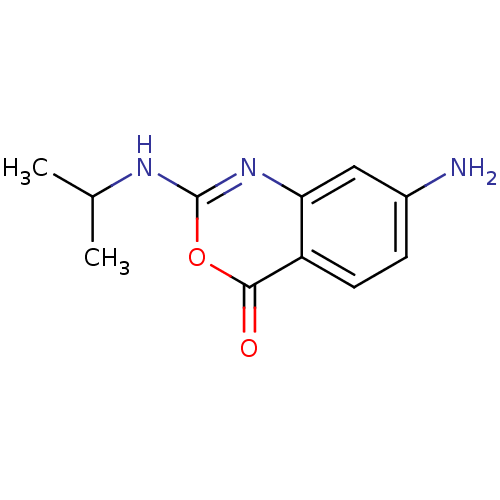
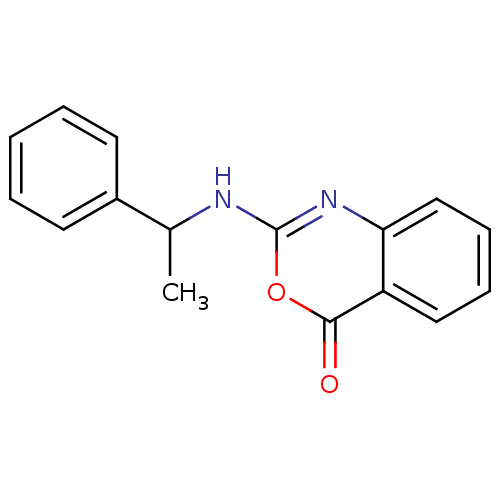
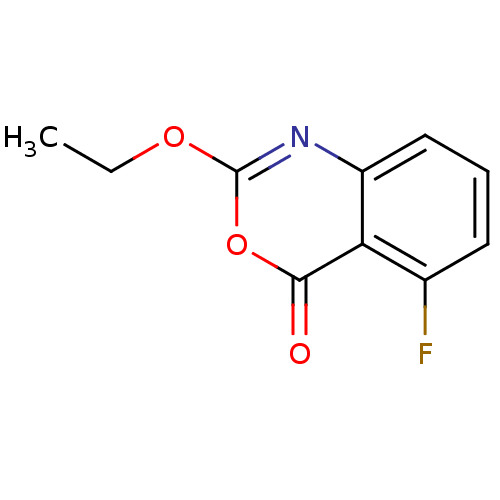
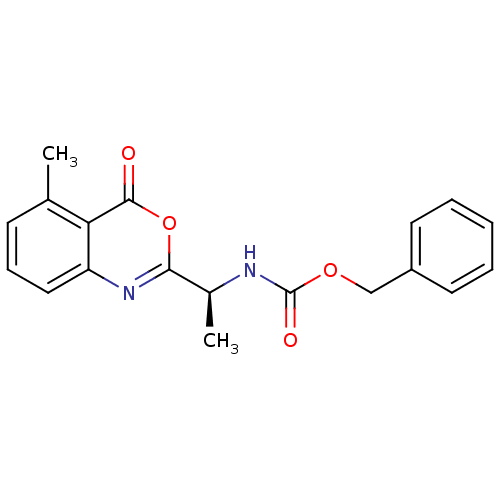
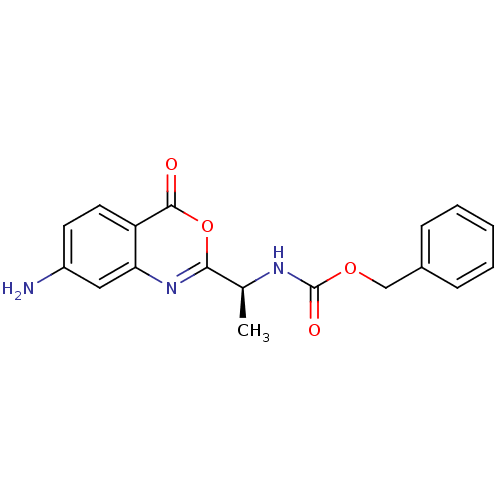
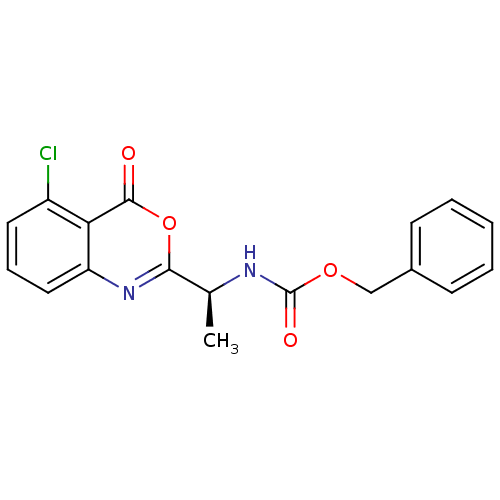
Affinity DataKi: 96nM IC50: 360nMpH: 5.52Assay Description:A zhang-poorman assay confirms a competitive inhibition mechanism or inhibit HIV-1 PR as a dimerization or mixed type inhibitor.More data for this Ligand-Target Pair
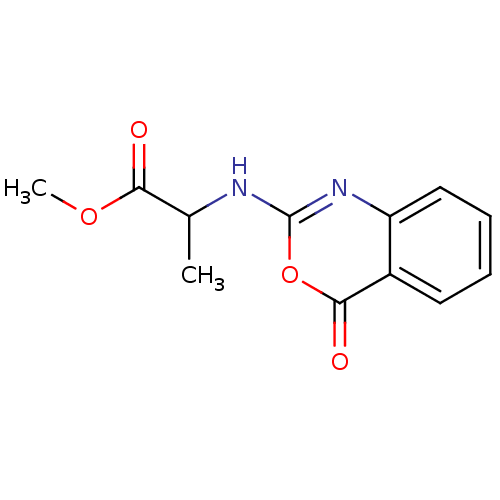
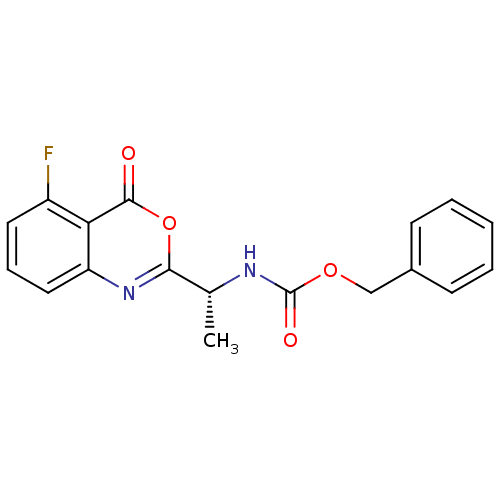
Affinity DataKi: 175nM IC50: 730nMpH: 5.52Assay Description:A zhang-poorman assay confirms a competitive inhibition mechanism or inhibit HIV-1 PR as a dimerization or mixed type inhibitor.More data for this Ligand-Target Pair
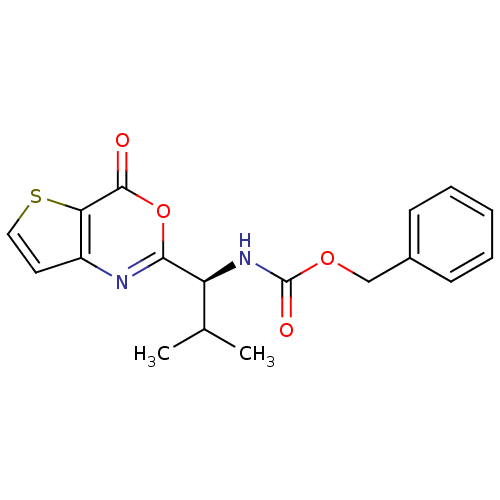
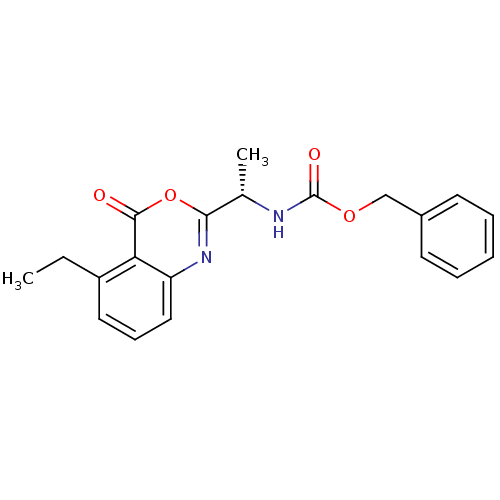
Affinity DataKi: 220nM IC50: 350nMpH: 5.52Assay Description:The competitive assay requires two inhibitors to act by a purely competitive mechanism, whereas the binding site of on the inhibitors has been establ...More data for this Ligand-Target Pair
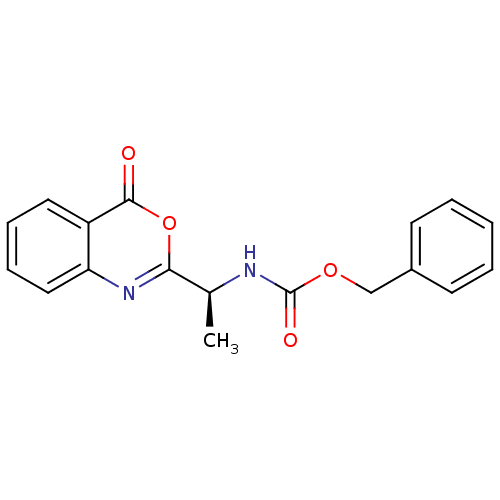
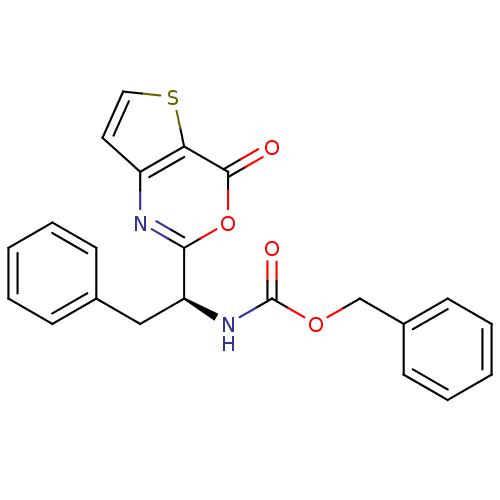
Affinity DataIC50: 480nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataKi: 600nM IC50: 2.90E+3nMpH: 5.52Assay Description:A zhang-poorman assay confirms a competitive inhibition mechanism or inhibit HIV-1 PR as a dimerization or mixed type inhibitor.More data for this Ligand-Target Pair
Affinity DataKi: 650nM IC50: 2.50E+3nMpH: 5.52Assay Description:A zhang-poorman assay confirms a competitive inhibition mechanism or inhibit HIV-1 PR as a dimerization or mixed type inhibitor.More data for this Ligand-Target Pair
Affinity DataIC50: 650nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataKi: 720nM IC50: 2.70E+3nMpH: 5.52Assay Description:A zhang-poorman assay confirms a competitive inhibition mechanism or inhibit HIV-1 PR as a dimerization or mixed type inhibitor.More data for this Ligand-Target Pair
Affinity DataKi: 1.13E+3nM IC50: 4.40E+3nMpH: 5.52Assay Description:The competitive assay requires two inhibitors to act by a purely competitive mechanism, whereas the binding site of on the inhibitors has been establ...More data for this Ligand-Target Pair
Affinity DataKi: 1.29E+3nM IC50: 4.90E+3nMpH: 5.52Assay Description:The competitive assay requires two inhibitors to act by a purely competitive mechanism, whereas the binding site of on the inhibitors has been establ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nM IC50: 5.90E+3nMpH: 5.52Assay Description:The competitive assay requires two inhibitors to act by a purely competitive mechanism, whereas the binding site of on the inhibitors has been establ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMpH: 5.52Assay Description:A zhang-poorman assay confirms a competitive inhibition mechanism or inhibit HIV-1 PR as a dimerization or mixed type inhibitor.More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMpH: 5.52Assay Description:The competitive assay requires two inhibitors to act by a purely competitive mechanism, whereas the binding site of on the inhibitors has been establ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+3nMpH: 5.52Assay Description:A zhang-poorman assay confirms a competitive inhibition mechanism or inhibit HIV-1 PR as a dimerization or mixed type inhibitor.More data for this Ligand-Target Pair
Affinity DataIC50: 6.30E+3nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+4nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 7.50E+4nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+5nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+5nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+5nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibitory concentration for peptidolytic activity against herpes simplex type-1 (HSV-1) protease at 10 uM.More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibitory activity against herpes simplex type 1 protease (HSV-1 pr).More data for this Ligand-Target Pair