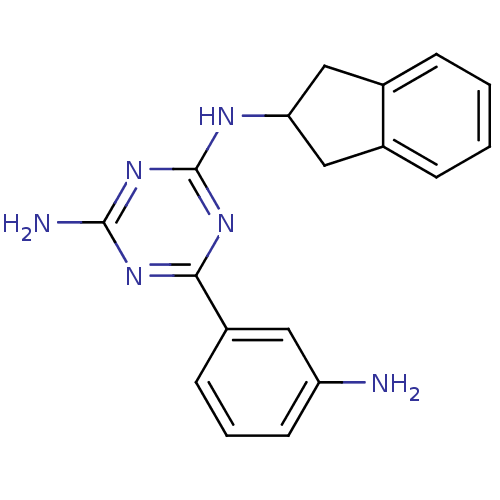
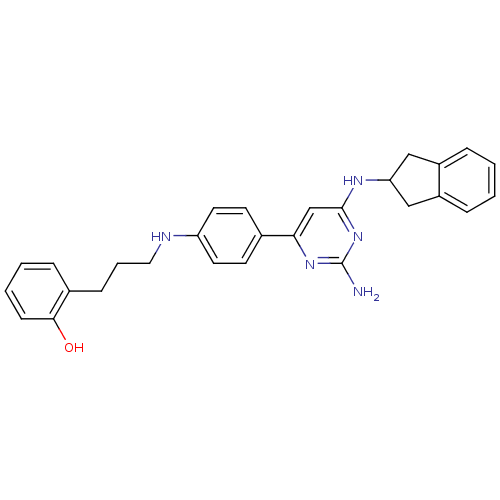
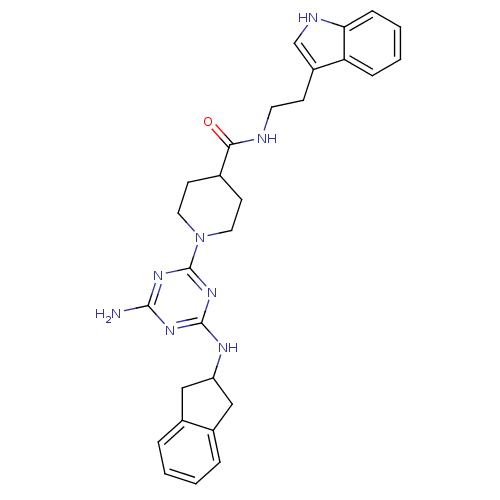
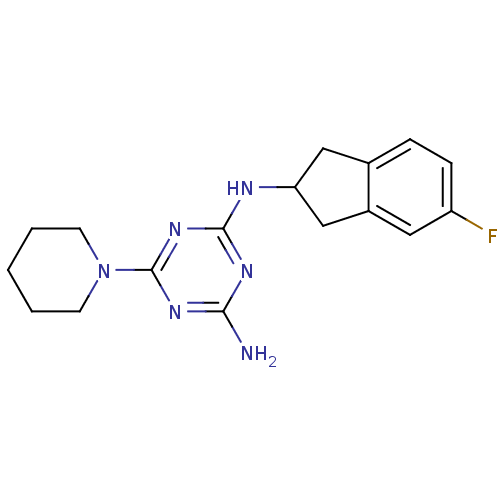
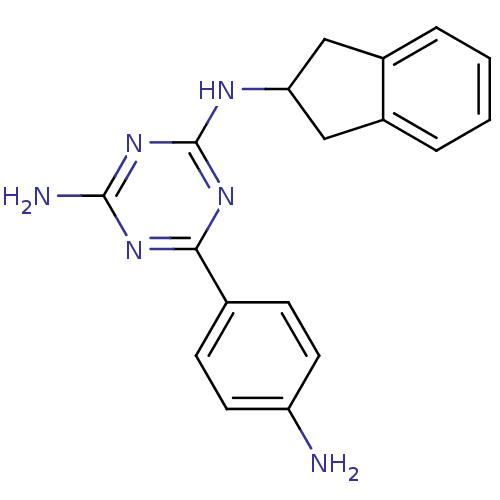
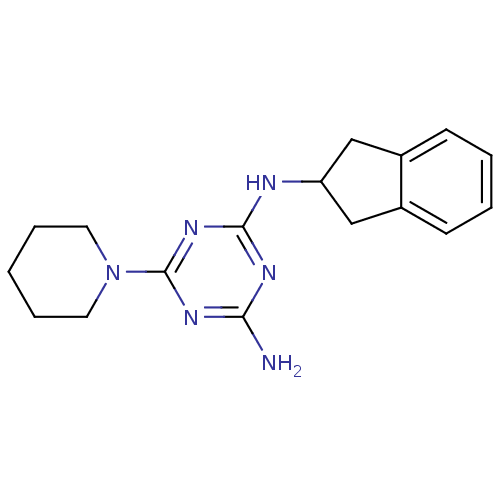
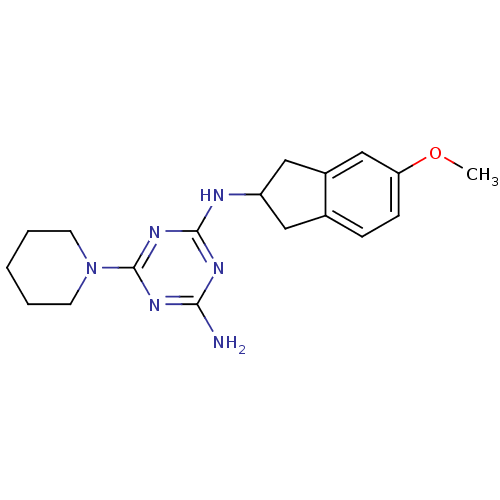
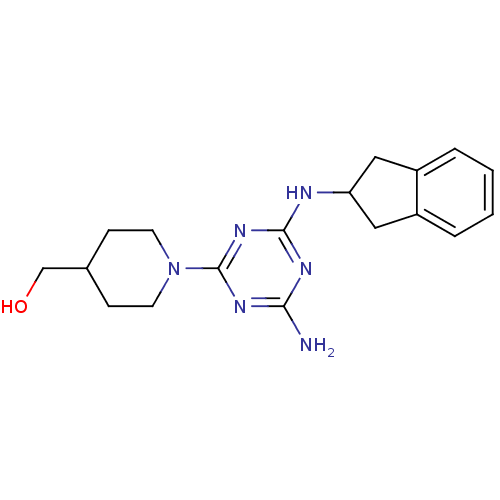
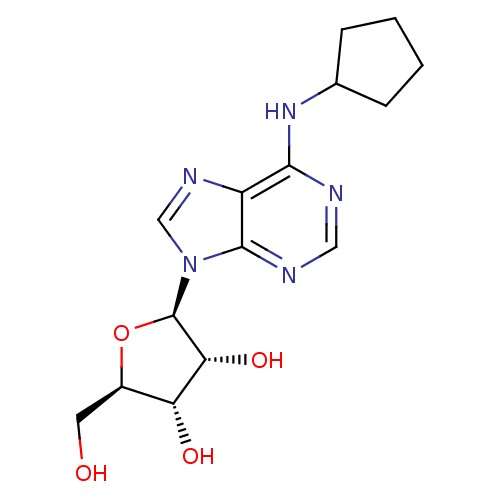
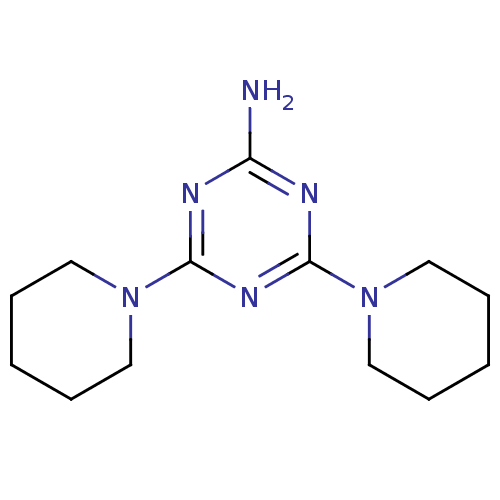
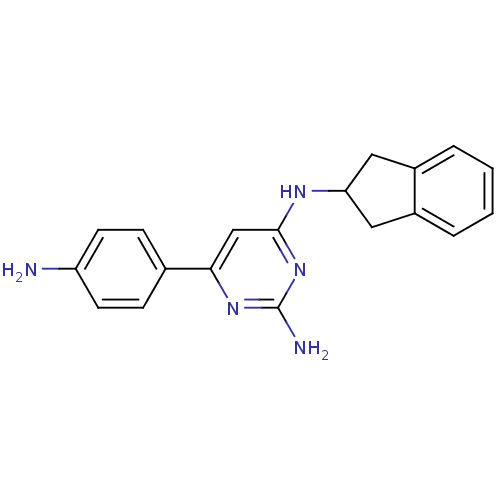
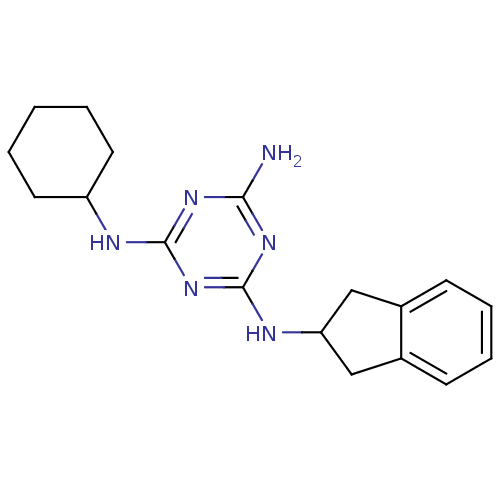
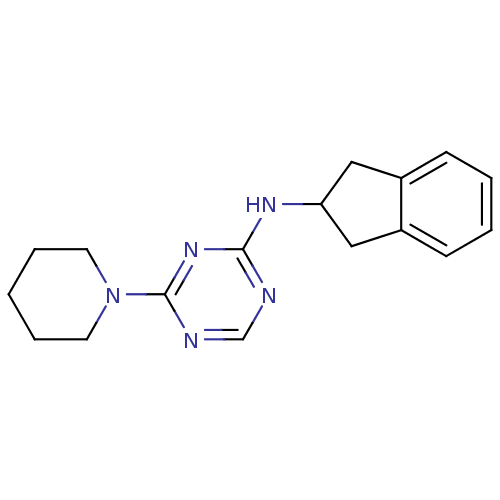
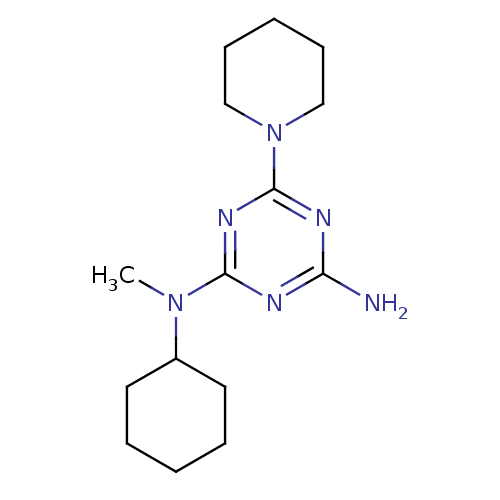
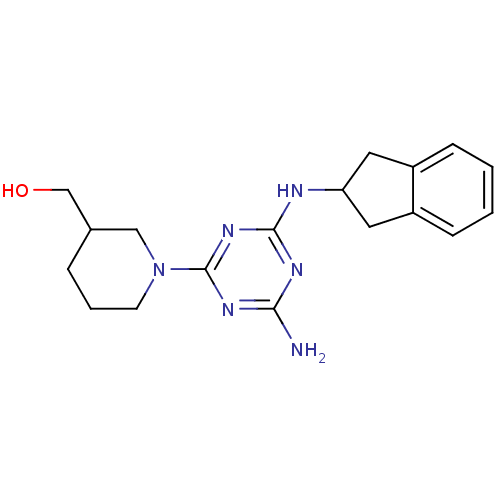
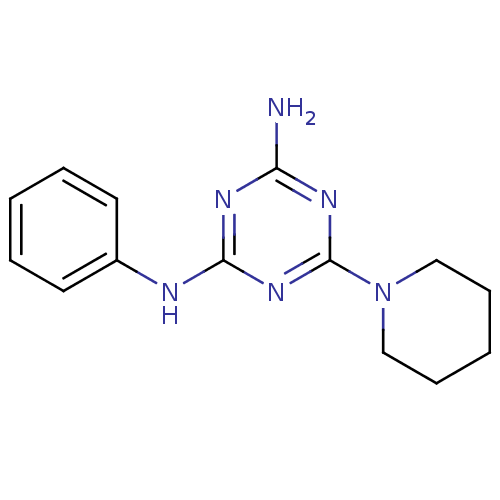
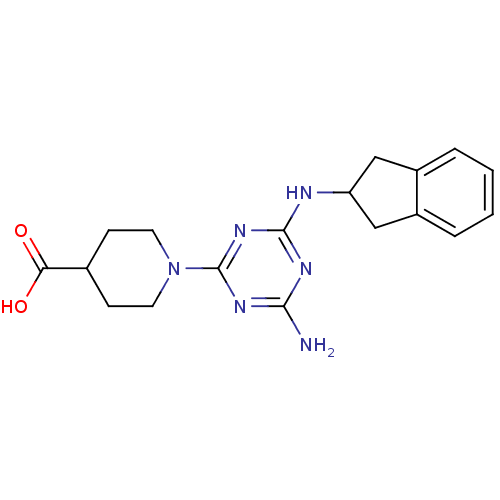
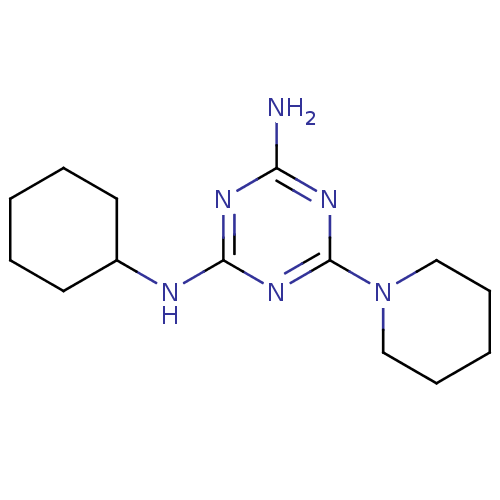
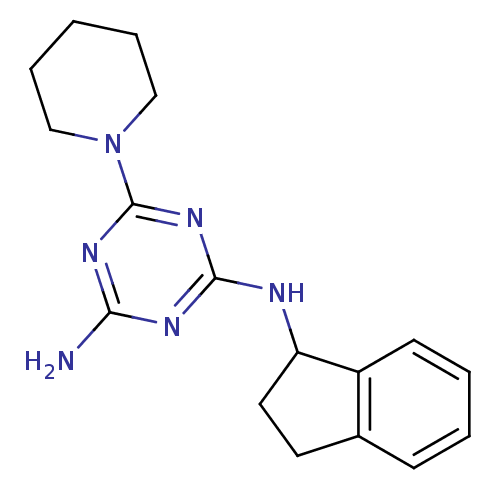
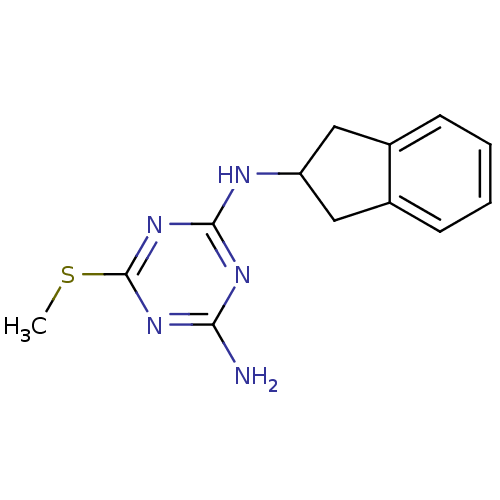
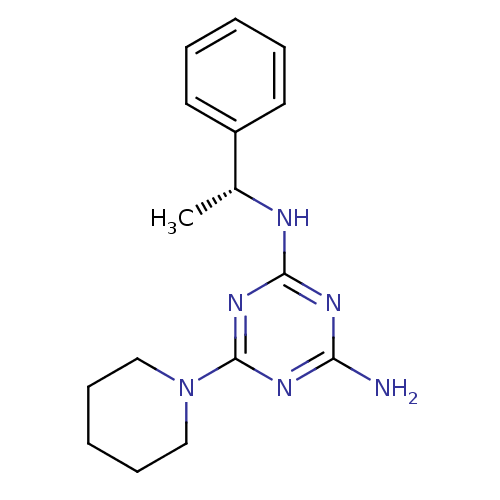
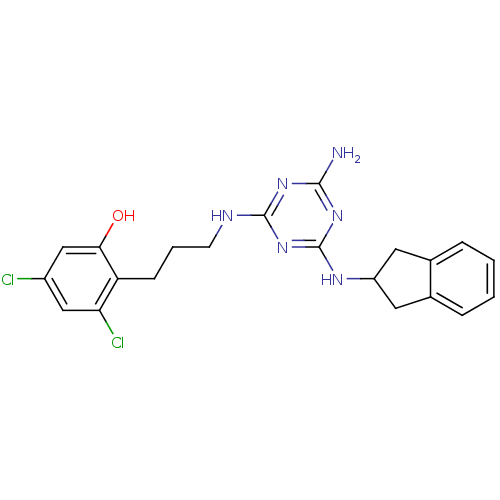
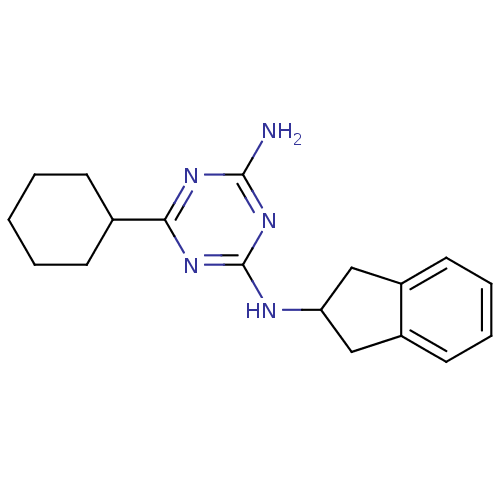
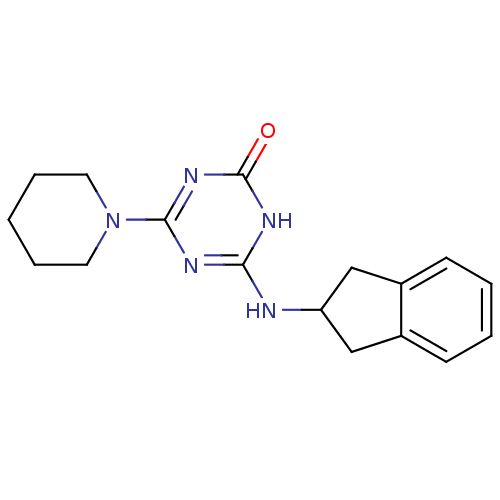
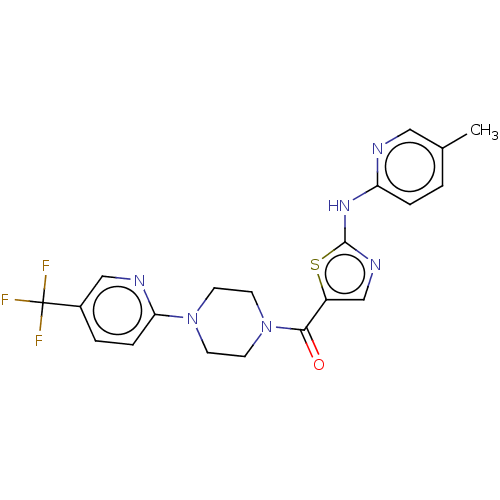
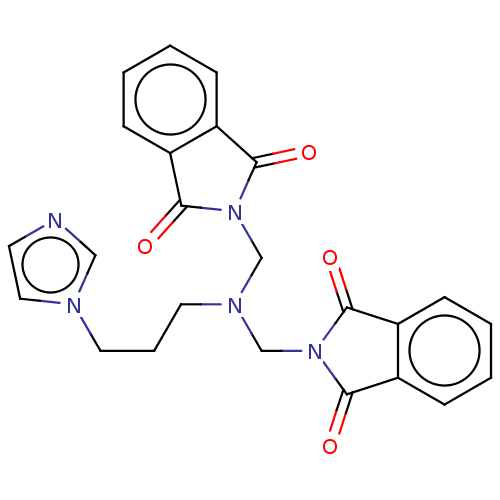
Affinity DataKi: 1.00E+4nMAssay Description:Binding affinity to Bacillus subtilis ErmC' methyltransferase assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: 1.25E+4nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nMAssay Description:Binding affinity to Bacillus subtilis ErmC' methyltransferase assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 3.70E+4nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >5.00E+4nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: 7.50E+4nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: 7.50E+4nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: 7.50E+4nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+4nMAssay Description:Inhibition of Bacillus subtilis ErmC' methyltransferaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Bacillus subtilis ErmC' methyltransferaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+5nMAssay Description:Inhibitory activity against ErmC methylaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+5nMAssay Description:Inhibition of Bacillus subtilis ErmC' methyltransferaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+5nMAssay Description:Inhibition of Bacillus subtilis ErmC' methyltransferaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+5nMAssay Description:Inhibition of Bacillus subtilis ErmC' methyltransferaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of Bacillus subtilis ErmC' methyltransferaseMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+5nMAssay Description:Inhibition of Bacillus subtilis ErmC' methyltransferaseMore data for this Ligand-Target Pair