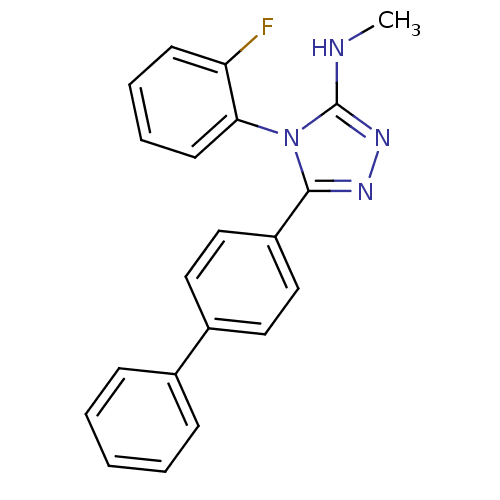
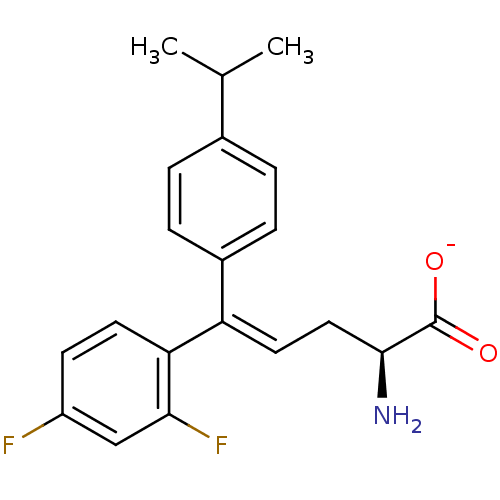
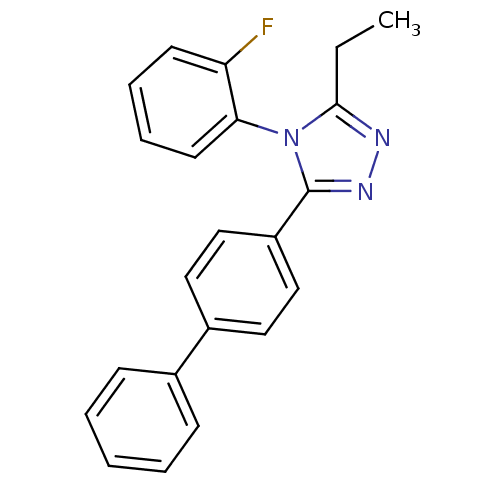
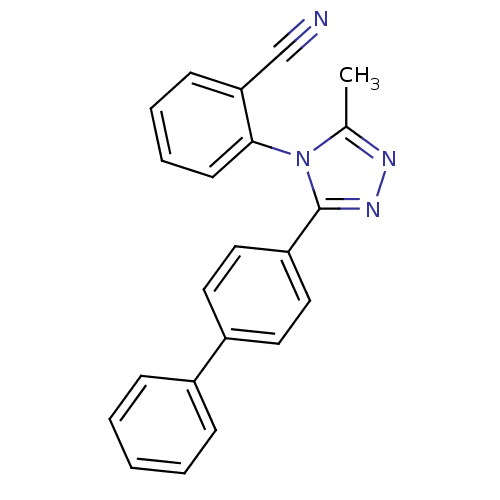
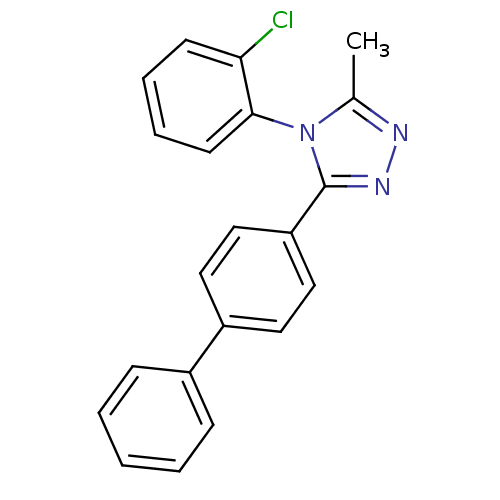
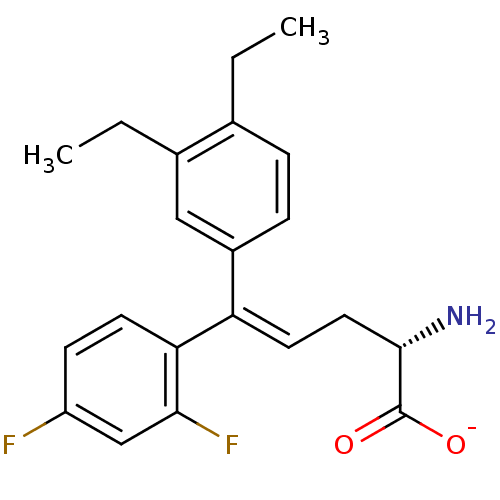
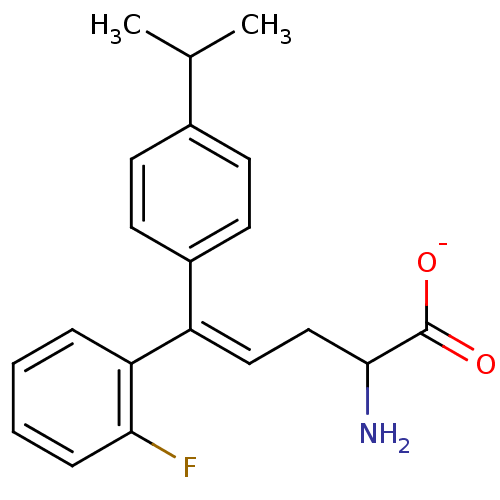
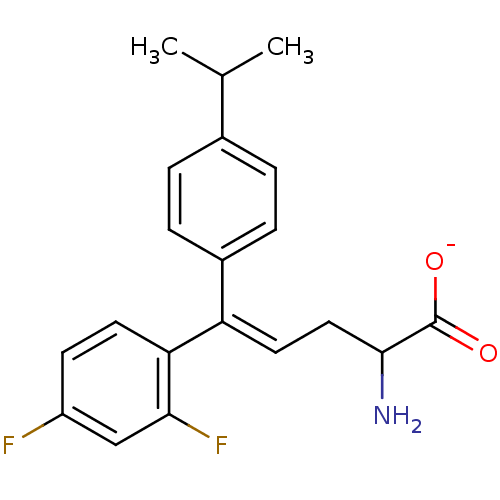
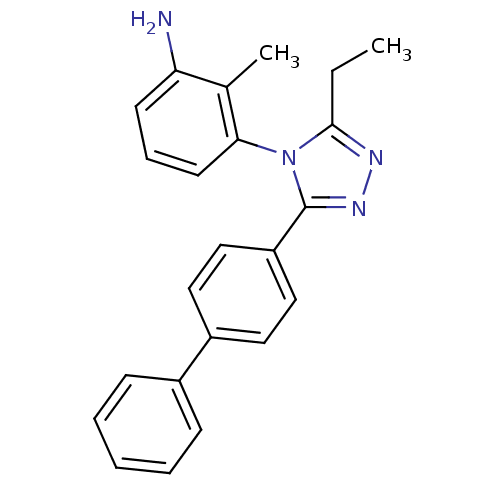
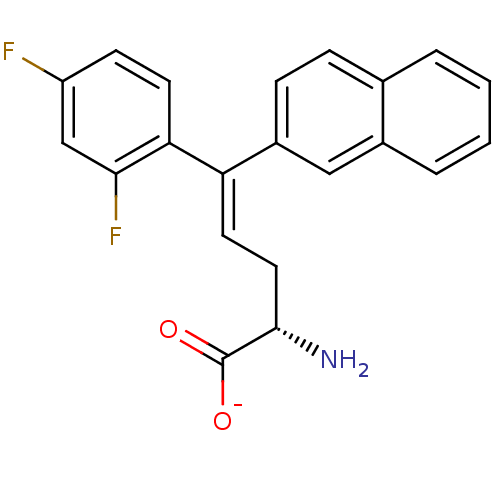
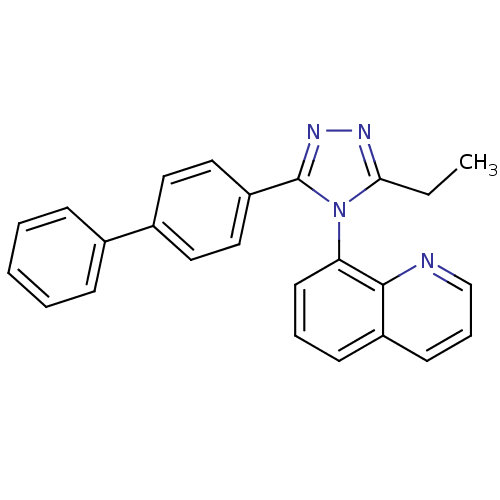
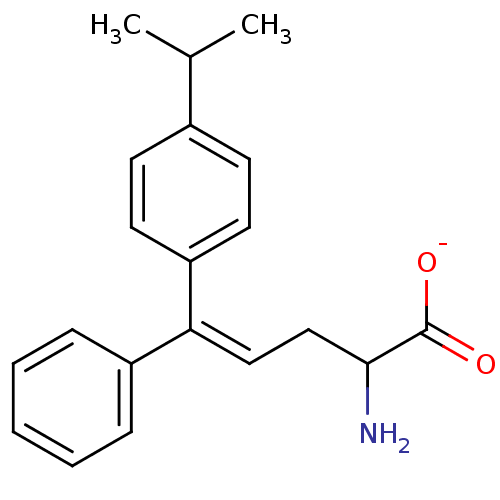
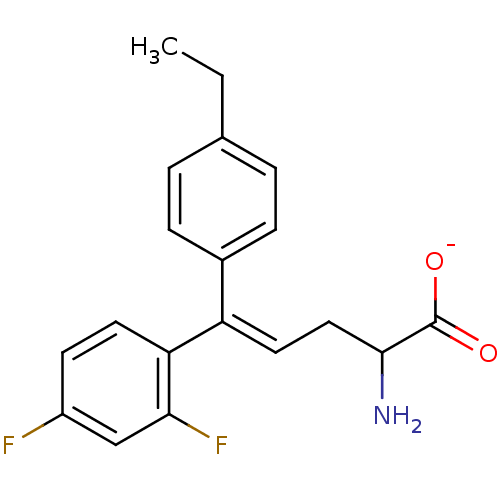
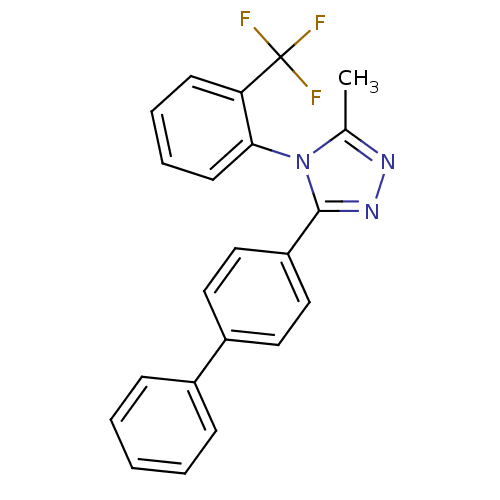
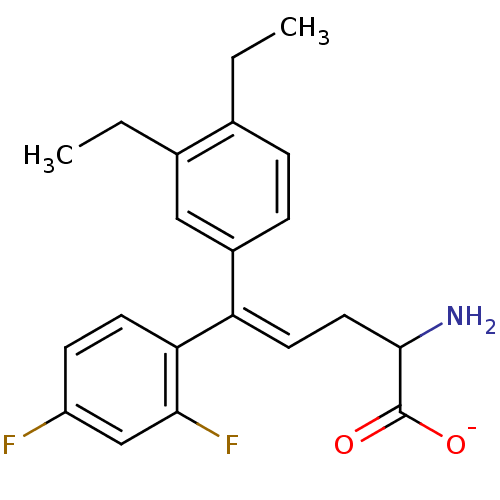
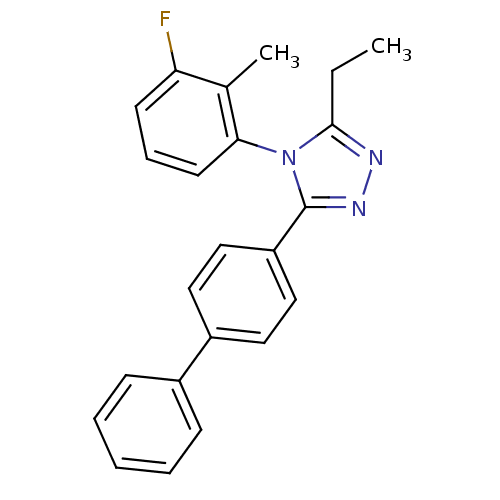
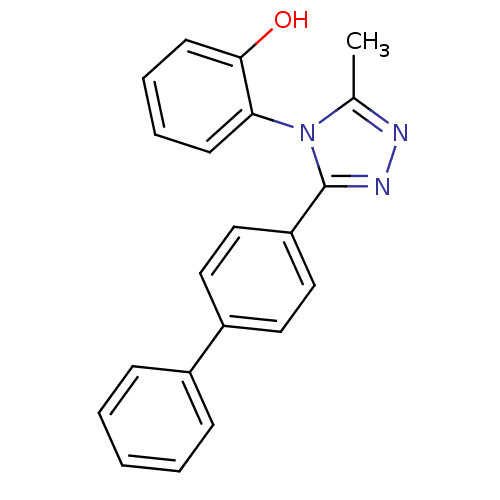
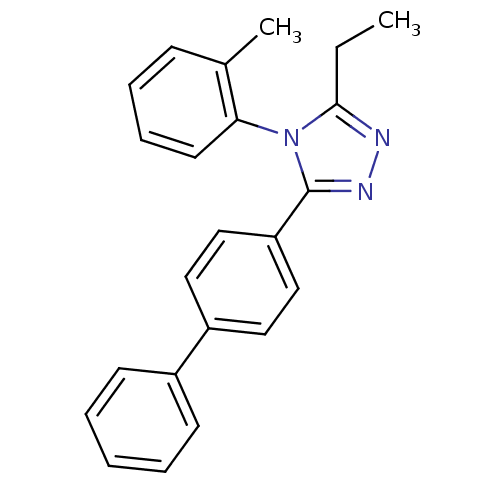
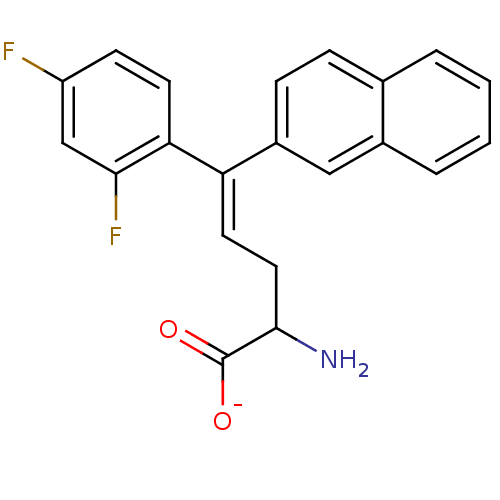
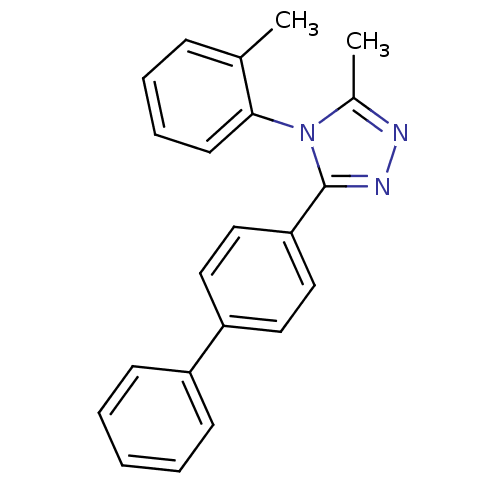
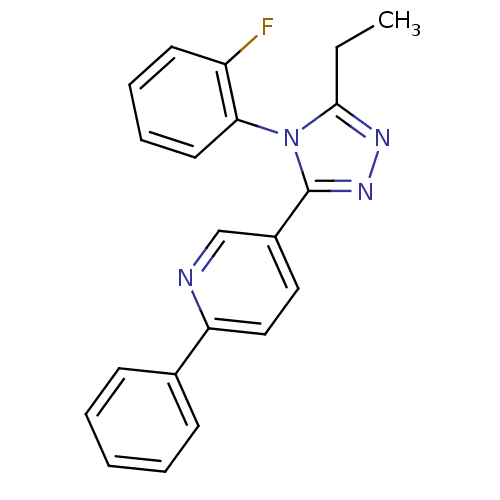
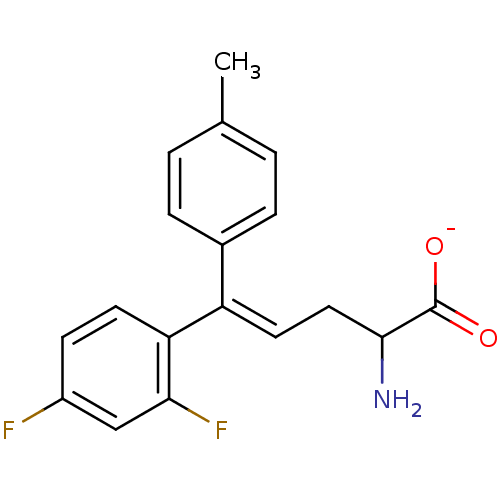
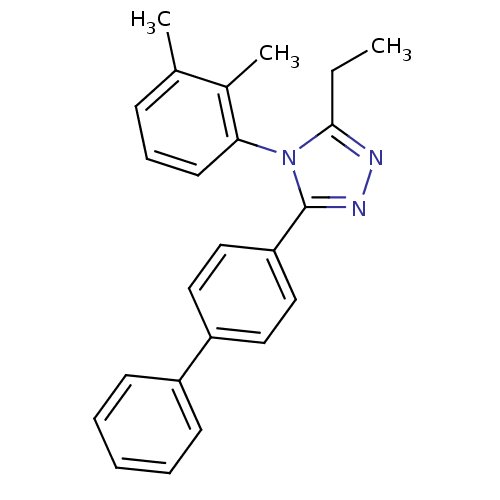
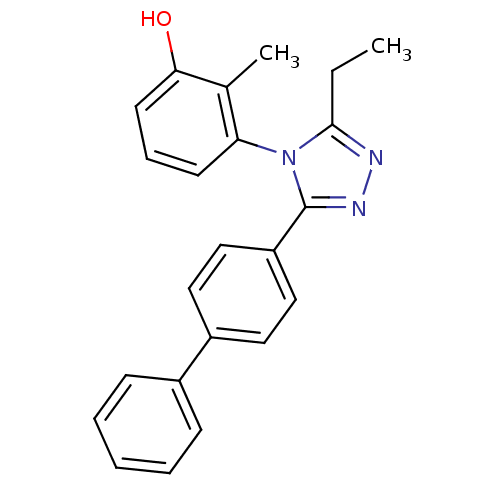
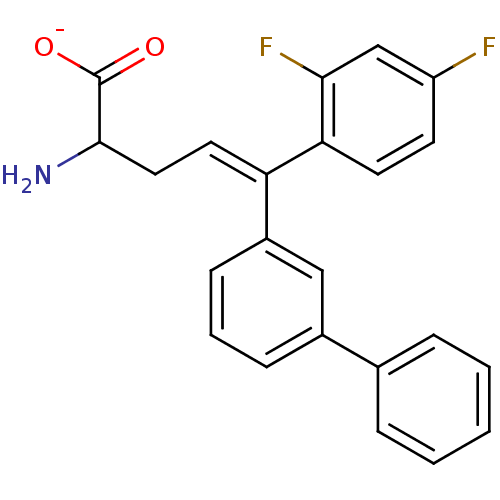
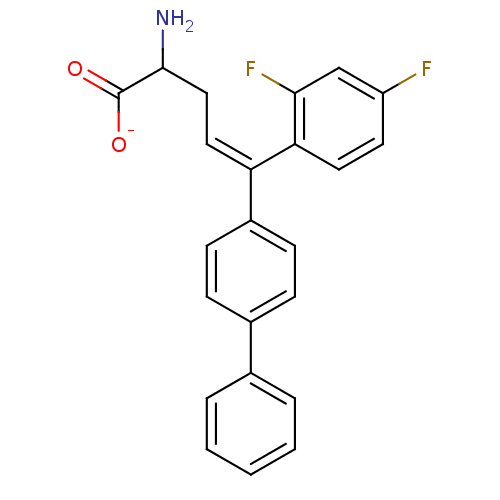
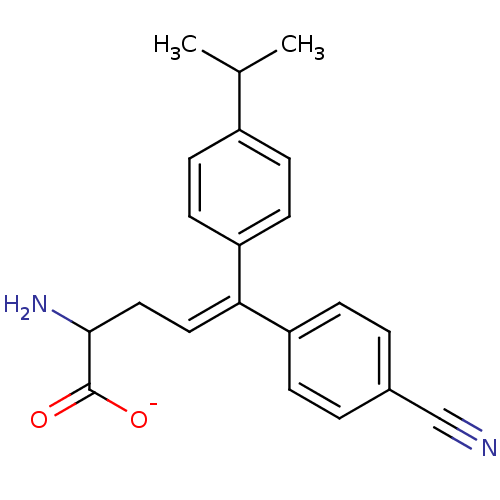
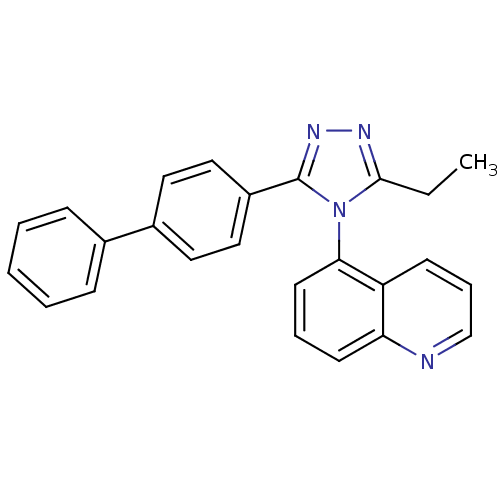
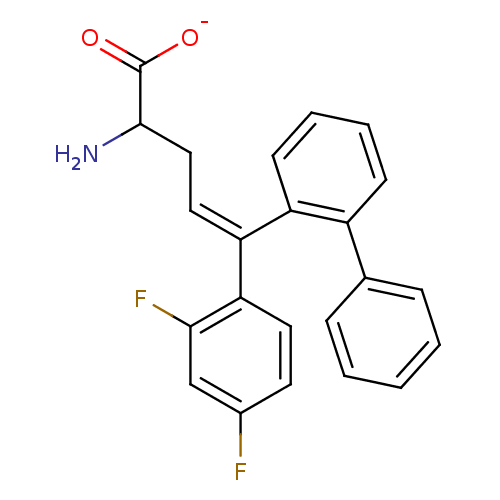
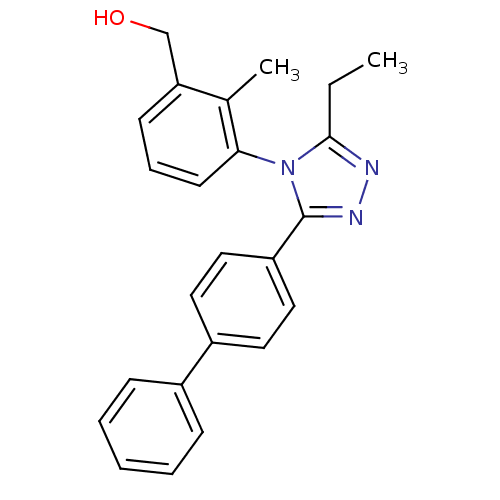
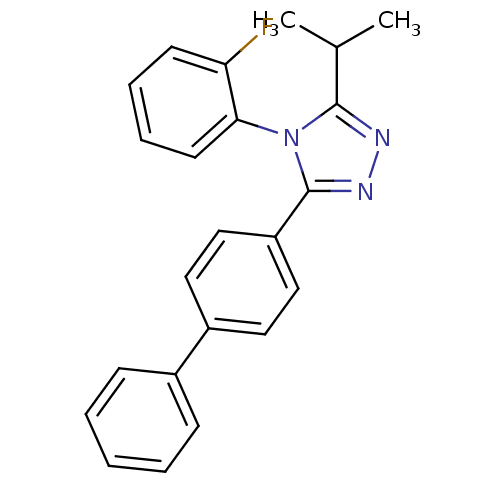
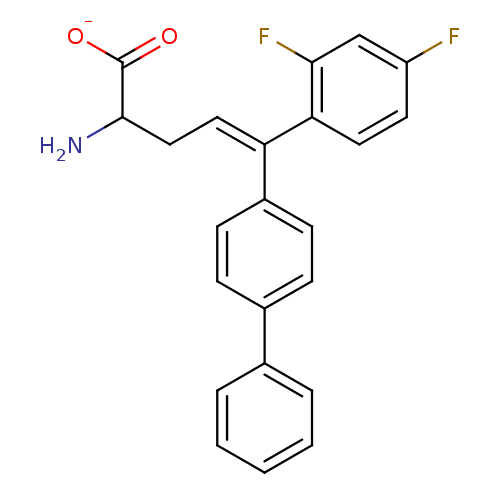
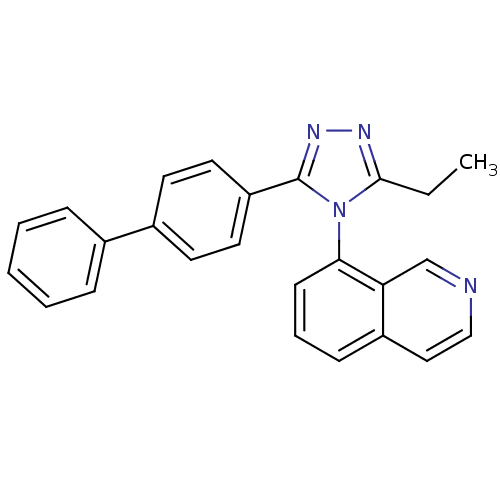
Affinity DataIC50: 110nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
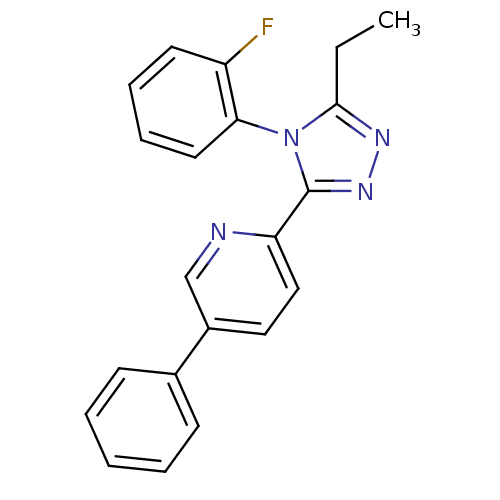
Affinity DataIC50: 150nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
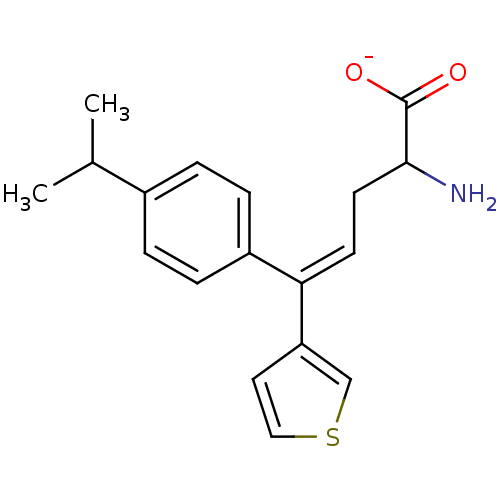
Affinity DataIC50: 230nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
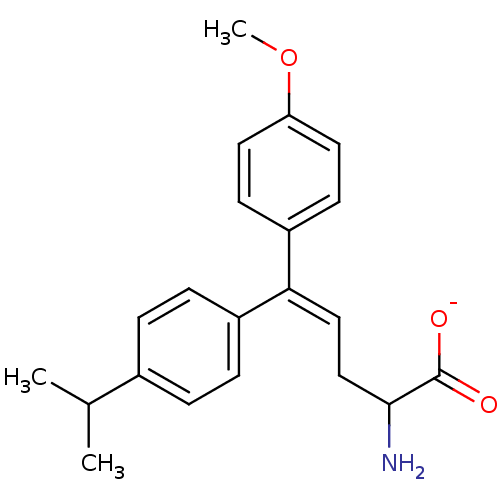
Affinity DataIC50: 230nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Displacement of [3H]glycine from Wistar rat brain GlyT2 after 10 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 260nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 283nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 330nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 340nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 365nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Displacement of [3H]glycine from Wistar rat brain GlyT2 after 10 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 460nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 660nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 660nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 680nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 770nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 780nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 790nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 890nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 990nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.34E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.74E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 2.03E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 2.03E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 2.06E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 2.64E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+3nMAssay Description:Displacement of [3H]glycine from Wistar rat brain GlyT2 after 10 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 2.75E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Displacement of [3H]glycine from GlyT2 in Wistar rat brainstem cells after 10 mins by scintillation counting analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of [3H]glycine uptake at GlyT2 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Displacement of [3H]glycine from Wistar rat brain GlyT2 after 10 mins by scintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 3.66E+3nMAssay Description:In vitro glycine reuptake inhibitory activity against glycine transporter type-2 (GlyT-2) transporter cells after incubation at 30 degrees for 30 min...More data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+3nMAssay Description:Displacement of [3H]glycine from Wistar rat brain GlyT2 after 10 mins by scintillation countingMore data for this Ligand-Target Pair