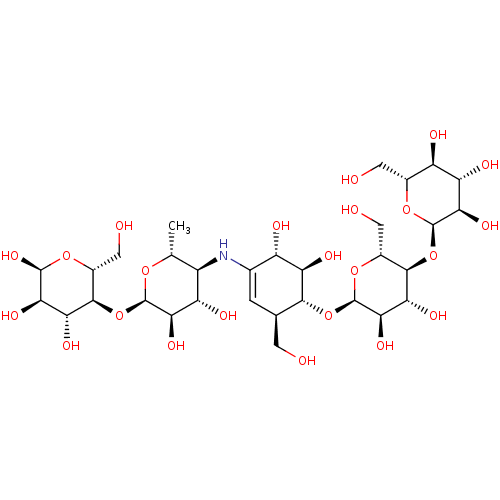
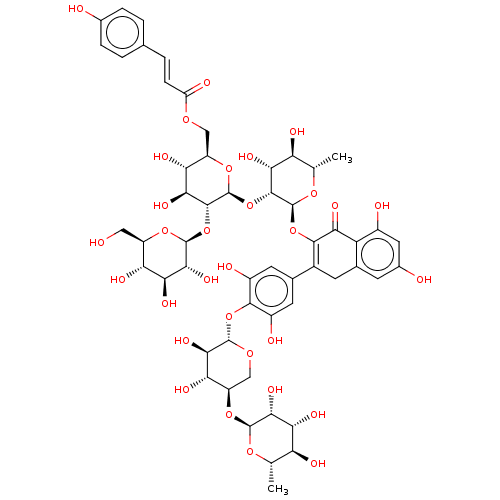
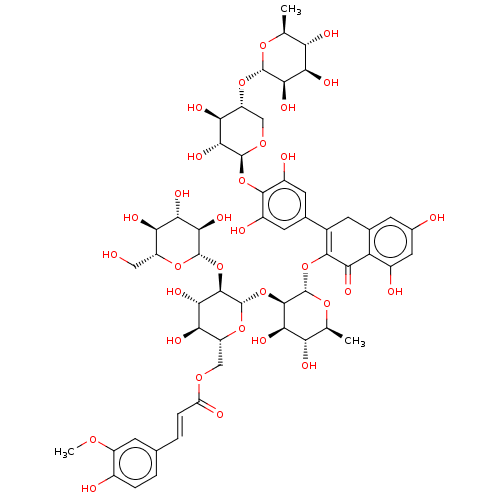
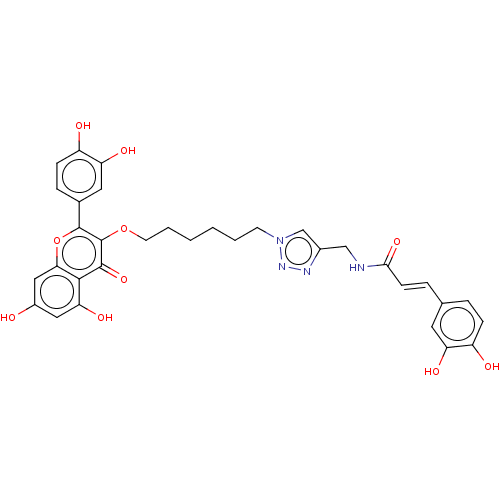
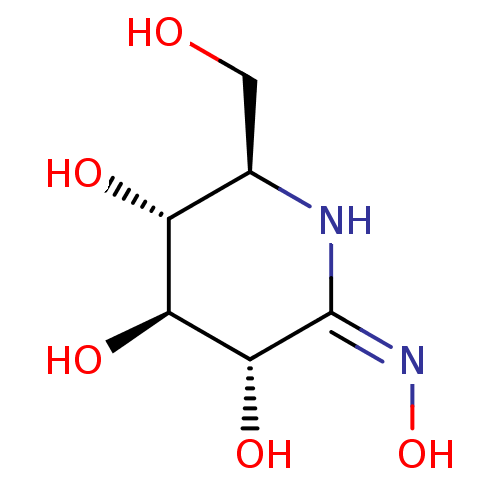
Affinity DataEC50: 0.0500nMAssay Description:Tested in vitro for amylase release from rat pancreatic aciniMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Tested in vitro for inhibition of amylase release from rat pancreatic aciniMore data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Tested in vitro for inhibition of amylase release from rat pancreatic aciniMore data for this Ligand-Target Pair
Affinity DataKi: 1nMpH: 7.0Assay Description:The release of 2-chloro-4-nitrophenol resulting from the HPA-catalyzed hydrolysis of 2-chloro-4-nitrophenyl a-maltotrioside (CNPG3) was monitored at ...More data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Tested in vitro for inhibition of amylase release from rat pancreatic aciniMore data for this Ligand-Target Pair
Affinity DataKi: 2.70nMpH: 7.0Assay Description:The release of 2-chloro-4-nitrophenol resulting from the HPA-catalyzed hydrolysis of 2-chloro-4-nitrophenyl a-maltotrioside (CNPG3) was monitored at ...More data for this Ligand-Target Pair
Affinity DataEC50: 3nMAssay Description:Tested in vitro for amylase release from rat pancreatic acini (percent of secretion)More data for this Ligand-Target Pair
Affinity DataEC50: 4nMAssay Description:Tested in vitro for amylase release from rat pancreatic acini (percent of secretion)More data for this Ligand-Target Pair
Affinity DataEC50: 6nMAssay Description:Tested in vitro for amylase release from rat pancreatic acini (percent of secretion)More data for this Ligand-Target Pair
Affinity DataKi: 7nMpH: 7.0Assay Description:The release of 2-chloro-4-nitrophenol resulting from the HPA-catalyzed hydrolysis of 2-chloro-4-nitrophenyl a-maltotrioside (CNPG3) was monitored at ...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Tested in vitro for inhibition of amylase release from rat pancreatic aciniMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Competitive inhibition of human pancreatic alpha-amylase by double reciprocal plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.10nM ΔG°: -47.0kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 9.10nM ΔG°: -46.7kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibition of human pancreatic alpha-amylaseMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibition of human pancreatic alpha-amylaseMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Inhibition of human pancreatic alpha-amylase expressed in Pichia pastoris using amylase as substrate preincubated with substrate for 10 mins followed...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Inhibition of human pancreatic alpha-amylase expressed in Pichia pastoris using amylase as substrate preincubated with substrate for 10 mins followed...More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Tested in vitro for amylase release from rat pancreatic acini (percent of secretion)More data for this Ligand-Target Pair
Affinity DataEC50: 20nMAssay Description:Tested in vitro for amylase release from rat pancreatic acini (percent of secretion)More data for this Ligand-Target Pair
Affinity DataKi: 21.3nM ΔG°: -44.5kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataEC50: 22nMAssay Description:Tested in vitro for amylase release from rat pancreatic acini (% of amylase secretion at 1 mM)More data for this Ligand-Target Pair
Affinity DataKi: 42nMAssay Description:Inhibition of human pancreatic alpha-amylase expressed in Pichia pastoris using amylase as substrate preincubated with substrate for 10 mins followed...More data for this Ligand-Target Pair
Affinity DataKi: 42.4nM ΔG°: -42.8kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Tested in vitro for amylase release from rat pancreatic acini (percent of secretion)More data for this Ligand-Target Pair
Affinity DataKi: 75nMAssay Description:Inhibition of human pancreatic alpha-amylaseMore data for this Ligand-Target Pair
Affinity DataKi: 79.3nM ΔG°: -41.2kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 93.3nM ΔG°: -40.8kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 730nM ΔG°: -35.6kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 1.25E+3nMAssay Description:Inhibition of human pancreatic alpha-amylase expressed in Pichia pastoris using amylase as substrate preincubated with substrate for 10 mins followed...More data for this Ligand-Target Pair
Affinity DataKi: 2.24E+3nM ΔG°: -32.8kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataIC50: 2.59E+3nMAssay Description:Inhibition of human pancreatic alpha-amylase expressed in Pichia pastoris using amylase as substrate preincubated with substrate for 10 mins followed...More data for this Ligand-Target Pair
Affinity DataKi: 3.60E+3nMAssay Description:Competitive inhibition of human pancreatic alpha-amylase by double reciprocal plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 6.10E+3nMAssay Description:Competitive inhibition of human pancreatic alpha-amylase by double reciprocal plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.39E+4nM ΔG°: -28.2kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 2.50E+4nMAssay Description:Inhibition of human pancreatic alpha-amylase assessed as hydrolysis of G3F by Dixon plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 4.68E+4nM ΔG°: -25.1kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 1.28E+5nM ΔG°: -22.6kJ/moleT: 2°CAssay Description:Inhibitor concentrations varied from 0.25 to 5 times the Ki value in 50 mM sodium phosphate, 100 mM sodium chloride buffer, pH 7.0, at 30 °C. 2-Chlor...More data for this Ligand-Target Pair
Affinity DataKi: 1.80E+6nMAssay Description:Inhibition of human pancreatic alpha-amylase assessed as hydrolysis of G3F by Dixon plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+7nMAssay Description:Inhibition of human pancreatic alpha-amylase assessed as hydrolysis of G3F by Dixon plot analysisMore data for this Ligand-Target Pair
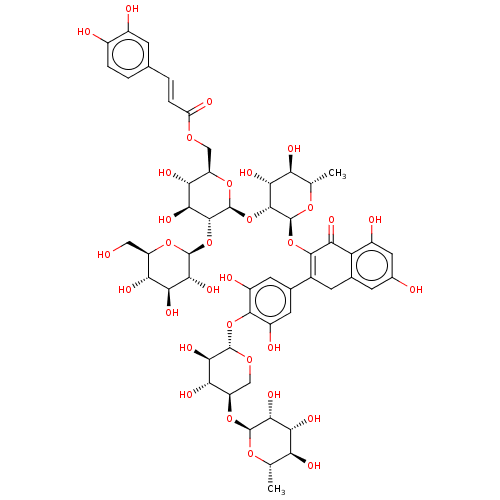
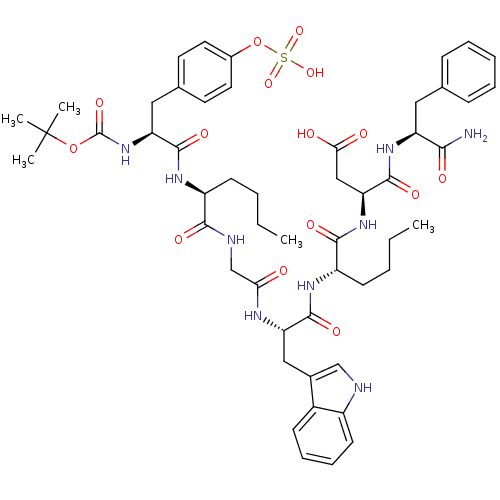
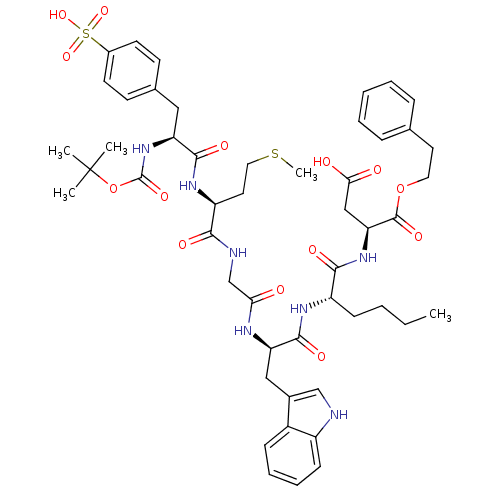
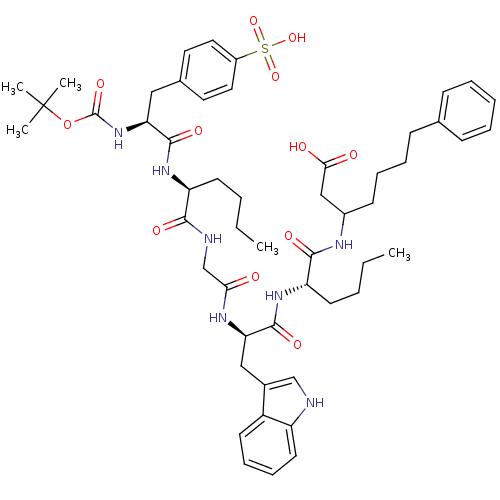


 3D Structure (crystal)
3D Structure (crystal)