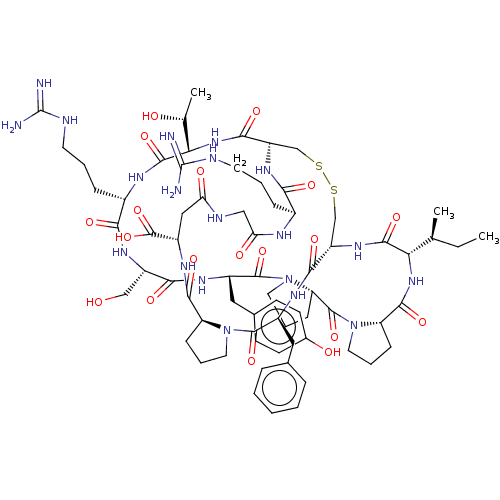
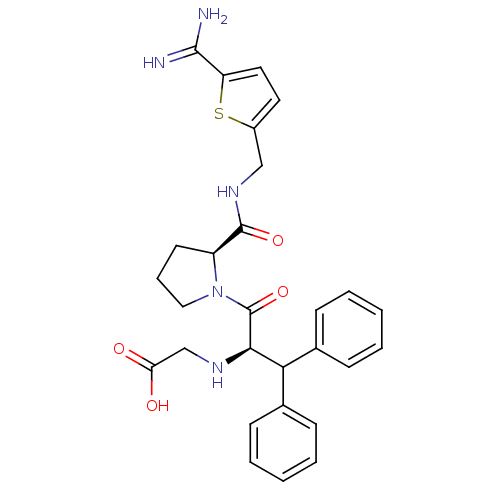
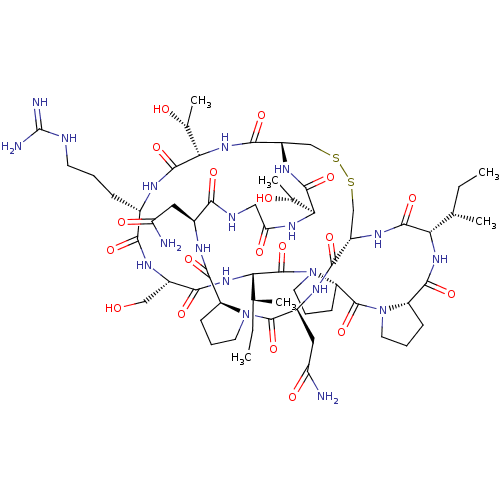
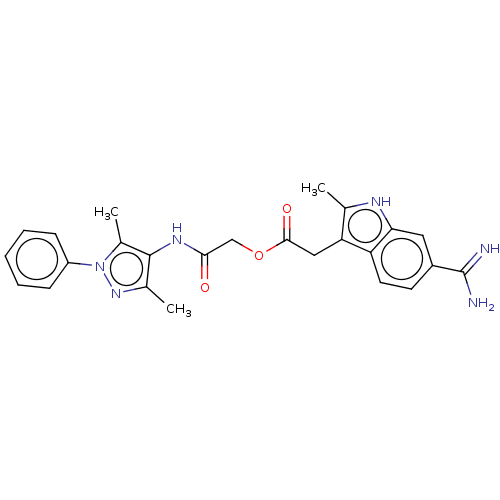
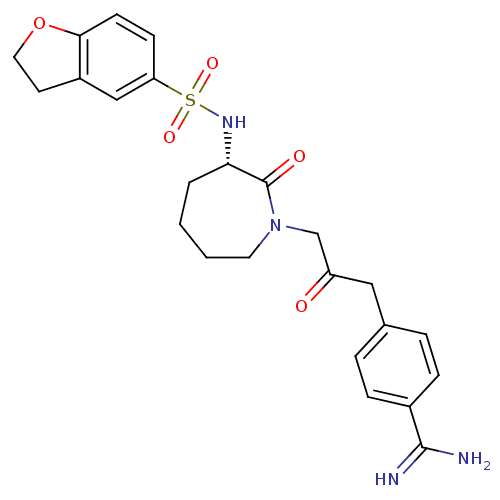
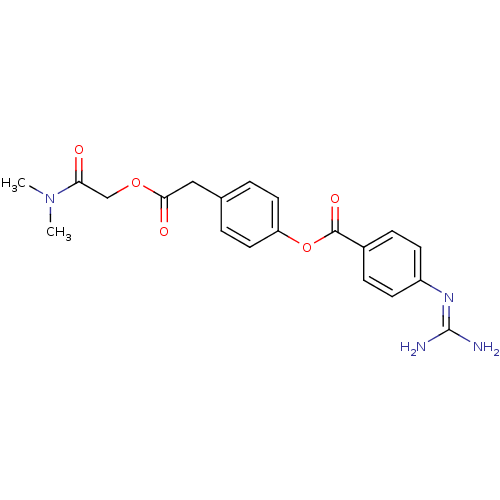
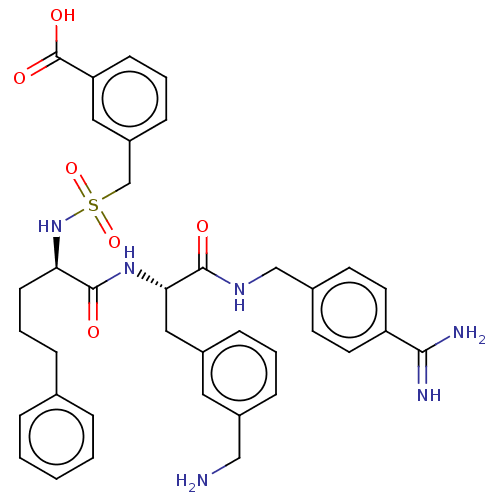
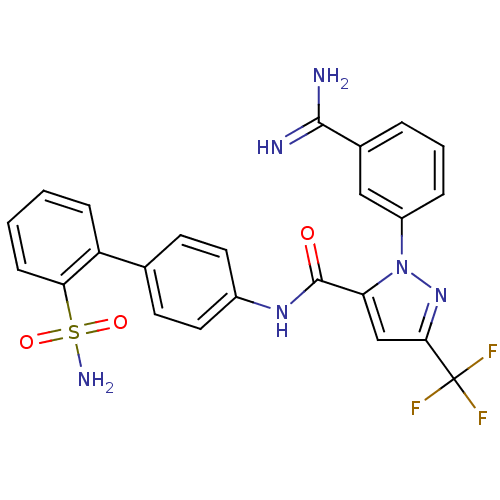
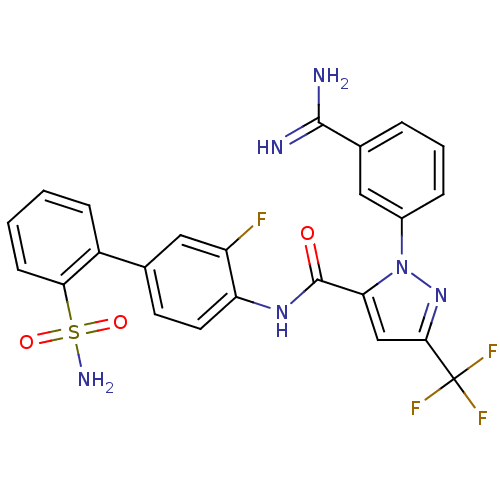
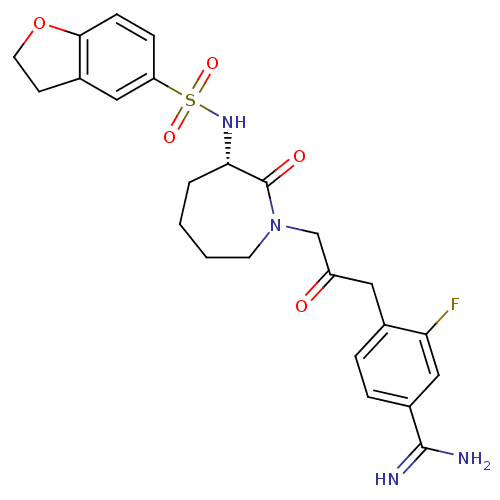
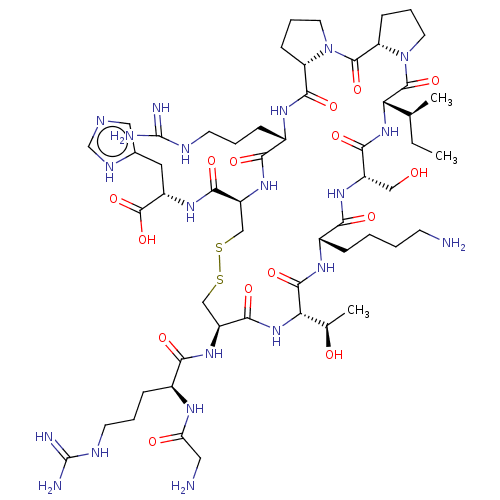
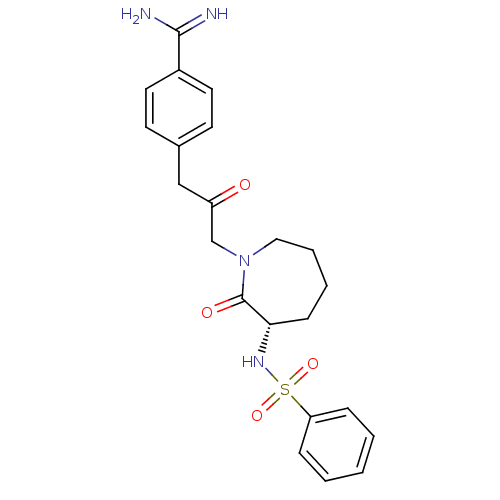
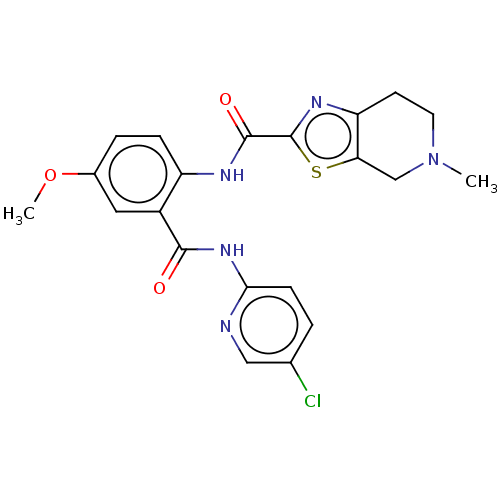
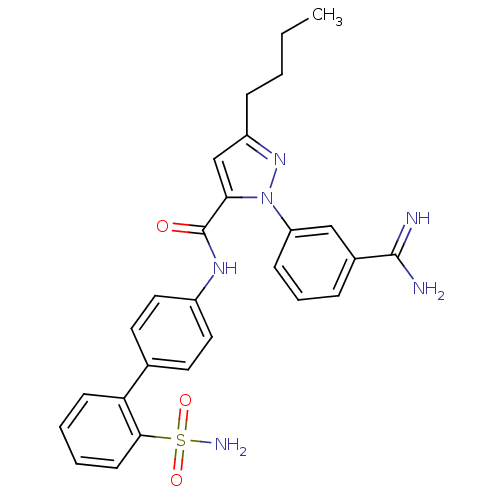
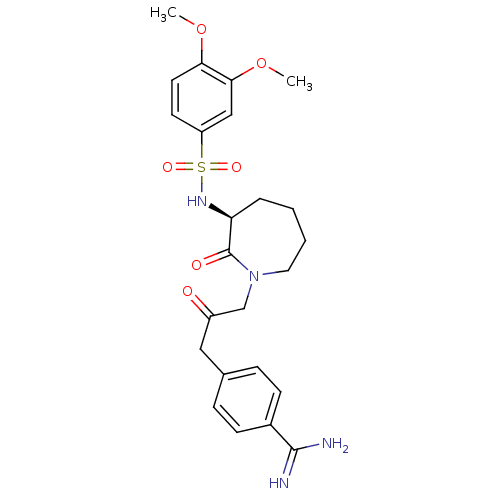
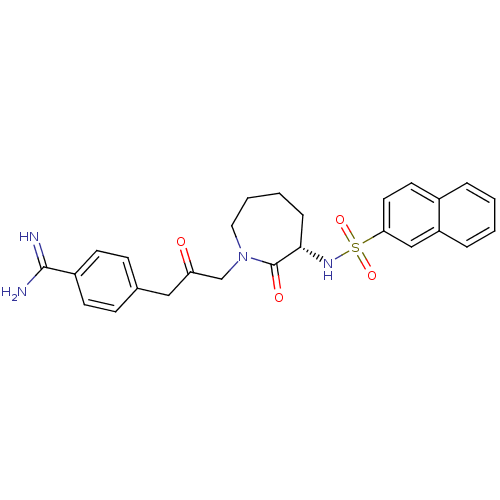
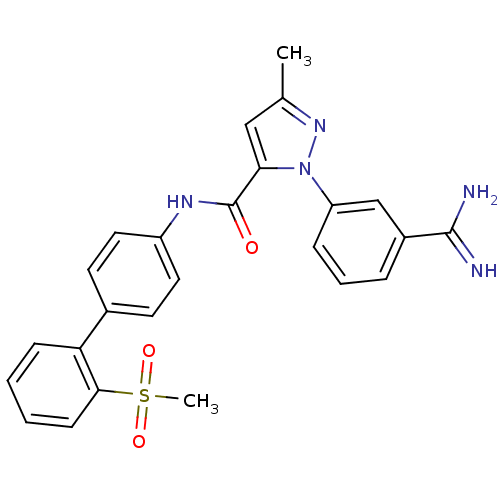
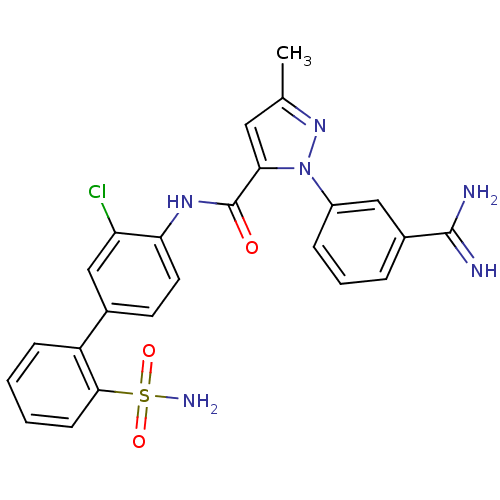
Affinity DataKi: 0.00230nMAssay Description:Inhibition of trypsin (unknown origin)More data for this Ligand-Target Pair
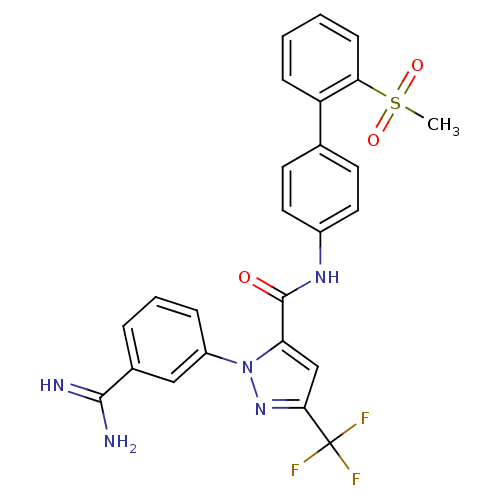
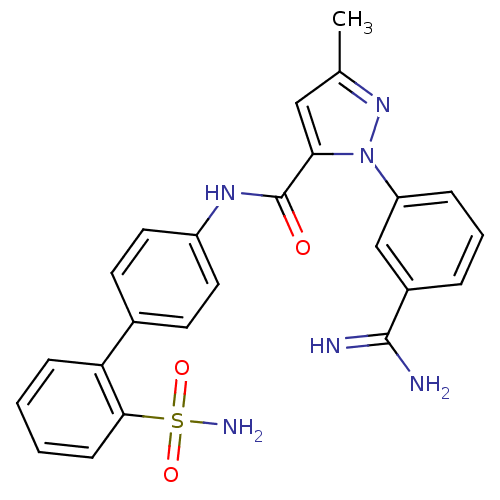
Affinity DataKi: 0.0300nMAssay Description:Inhibition of beta-trypsin (unknown origin) using Bz-FVRpNA substrate by spectrophotometry methodMore data for this Ligand-Target Pair
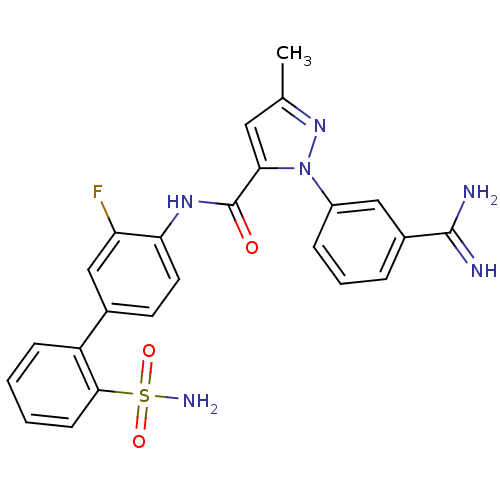
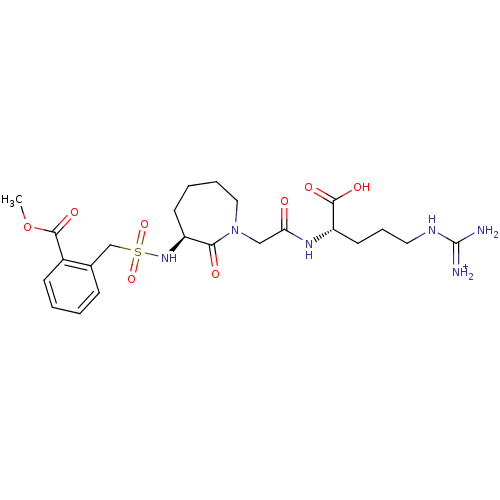
Affinity DataKi: 0.100nMAssay Description:Inhibition of trypsin (unknown origin)More data for this Ligand-Target Pair
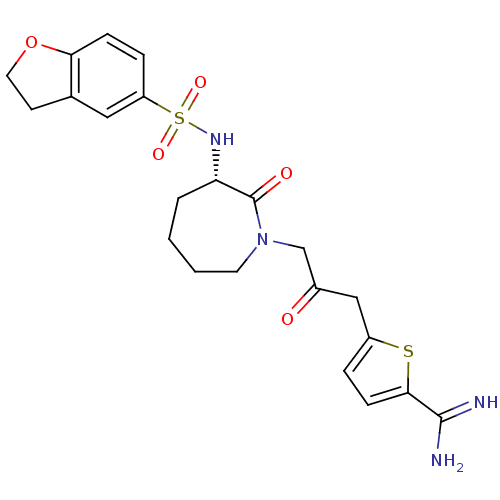
Affinity DataKi: 0.100nMAssay Description:Compound was tested for inhibition of Sunflower beta-trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Inhibition of beta-trypsin (unknown origin) using Bz-FVRpNA substrate by spectrophotometry methodMore data for this Ligand-Target Pair
Affinity DataKi: 0.160nMAssay Description:Inhibition of trypsin (unknown origin) using Bz-FVRpNA as substrate incubated for 30 mins measured for 7 mins by morrison plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:In vitro inhibition constant (Ki) against human trypsin was determinedMore data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Inhibition of trypsin (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 0.700nMAssay Description:Inhibitory constant against trypsinMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of trypsin (unknown origin) pre-incubated for 30 mins before N-Boc-FSR-AMC substrate addition and measured after 30 minsMore data for this Ligand-Target Pair
Affinity DataKi: 1.30nMAssay Description:Inhibitory constant against trypsinMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of trypsin (unknown origin) using Boc-Val-Pro-Arg-AMC as substrate preincubated with enzyme for 15 mins followed by addition of substrate ...More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibitory constant against trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:Inhibitory constant against trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Inhibitory constant against trypsinMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition of trypsin (unknown origin) using Boc-Val-Pro-Arg-AMC as substrate preincubated with enzyme for 15 mins followed by addition of substrate ...More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Inhibitory constant against trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Inhibitory constant against trypsin determined in VitroMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80nMAssay Description:Inhibition of trypsin (unknown origin) using Boc-Val-Pro-Arg-AMC as substrate preincubated with enzyme for 15 mins followed by addition of substrate ...More data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Inhibitory constant against trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 3.20nMAssay Description:Inhibitory constant against trypsin determined in VitroMore data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 3.40nMAssay Description:Binding affinity towards human trypsinMore data for this Ligand-Target Pair
Affinity DataIC50: 3.5nMAssay Description:Inhibition of trypsin (unknown origin) using Boc-Val-Pro-Arg-AMC as substrate preincubated with enzyme for 15 mins followed by addition of substrate ...More data for this Ligand-Target Pair
Affinity DataKi: 3.60nMAssay Description:Inhibition of trypsin (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 3.60nMAssay Description:Inhibition of trypsin (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 3.90nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 4.30nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 4.60nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 5.90nMAssay Description:Inhibitory constant against trypsin determined in VitroMore data for this Ligand-Target Pair
Affinity DataKi: 6.40nMAssay Description:Binding affinity towards human trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Inhibition of trypsin (unknown origin) using Bz-FVR-pNA as substrate preincubated for 30 mins followed by substrate addition and measured for 7 mins ...More data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Inhibition of trypsin (unknown origin) using Bz-FVRpNA as substrate incubated for 30 mins measured for 7 mins by morrison plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Inhibitory constant against trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Inhibitory constant against trypsin determined in VitroMore data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Inhibition of beta-trypsin (unknown origin) using Bz-FVRpNA substrate by spectrophotometry methodMore data for this Ligand-Target Pair
Affinity DataKi: 8.30nMAssay Description:Inhibitory constant against trypsin determined in VitroMore data for this Ligand-Target Pair
Affinity DataIC50: 9.10nMAssay Description:Inhibition of human trypsin preincubated for 30 mins followed by substrate addition and measured for 20 minsMore data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibitory constant against trypsin determined in VitroMore data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Inhibitory constant against trypsin determined in VitroMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Binding affinity towards human trypsinMore data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:In vitro activity against human trypsin.More data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Inhibition of human trypsin using S-2222 as substrate after 30 mins by spectrophotometric methodMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Inhibitory constant against trypsin determined in VitroMore data for this Ligand-Target Pair
Affinity DataKi: 16nMAssay Description:Binding affinity towards human trypsinMore data for this Ligand-Target Pair