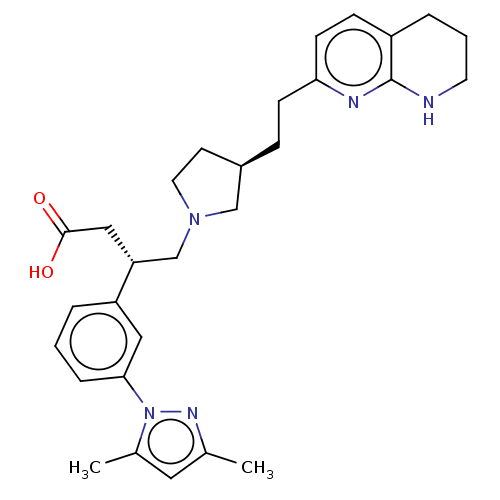
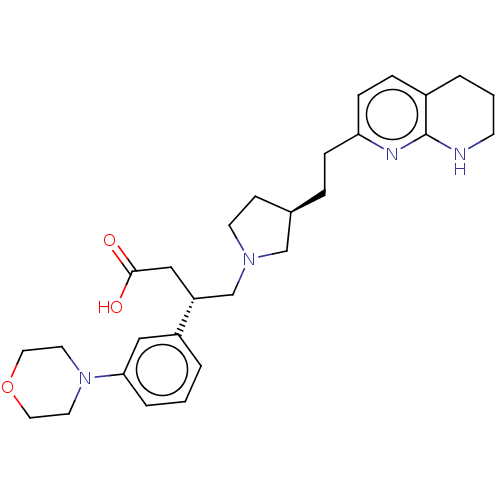
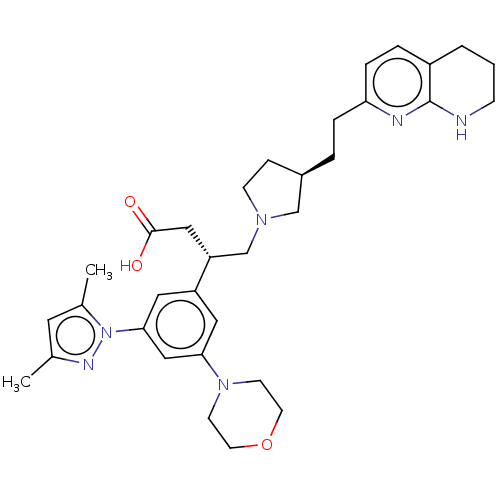
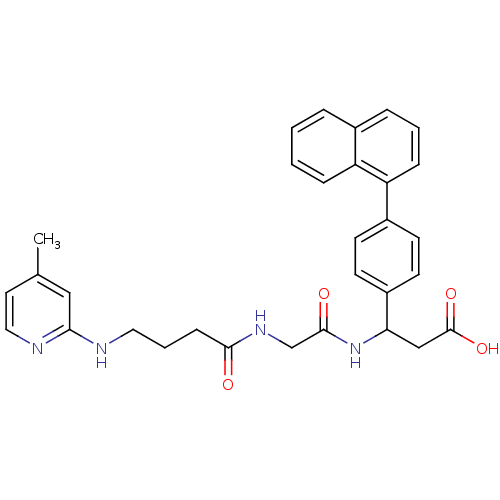
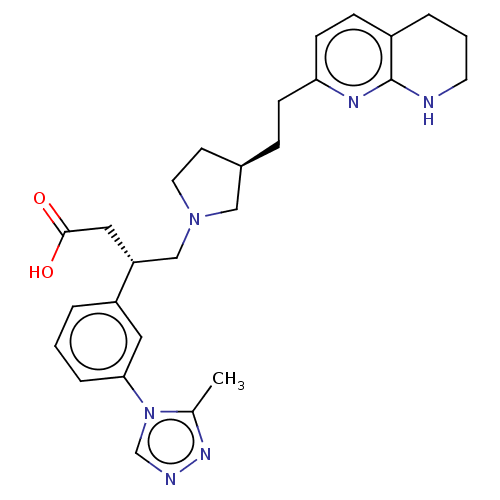
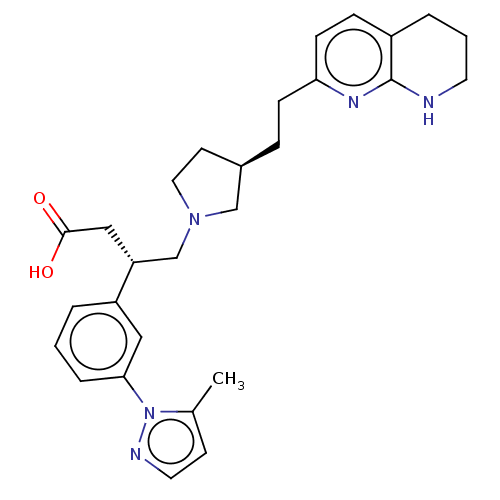
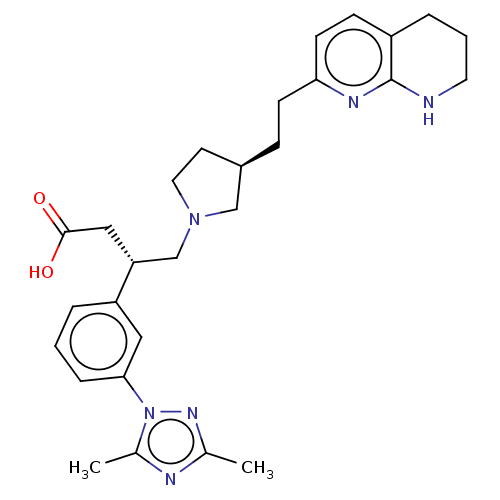
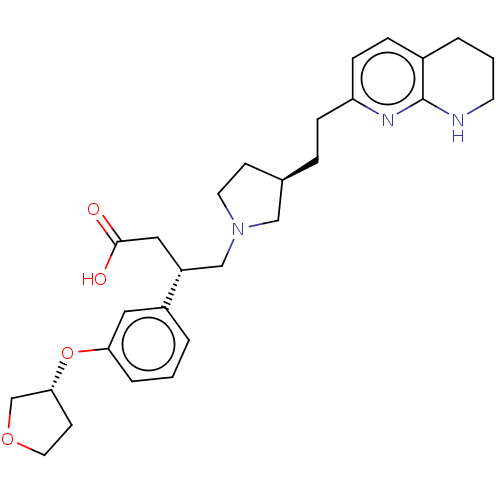
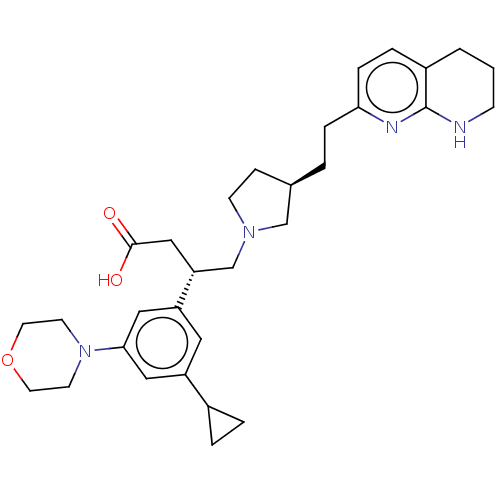
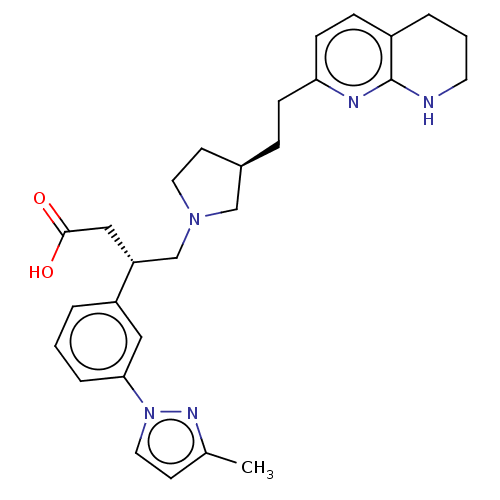
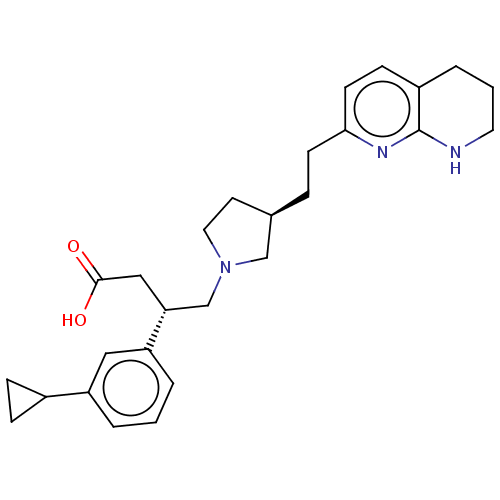
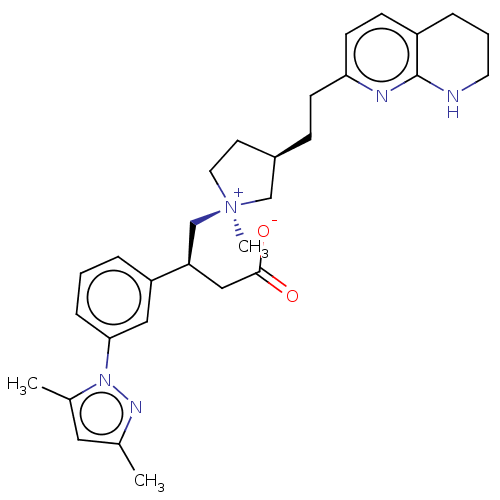
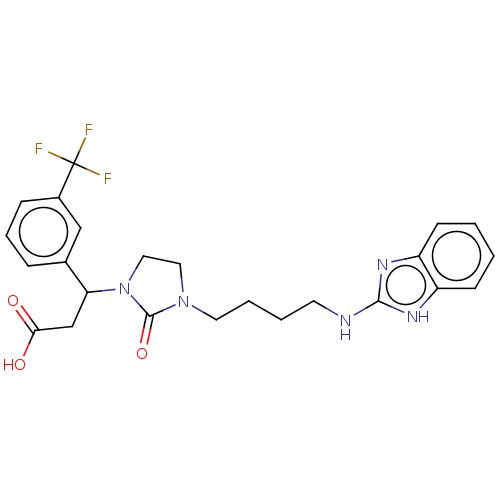
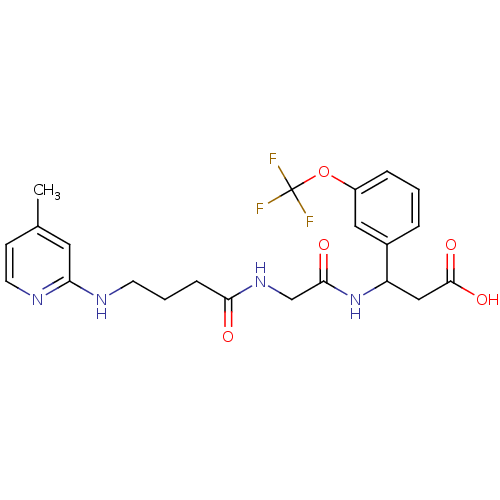
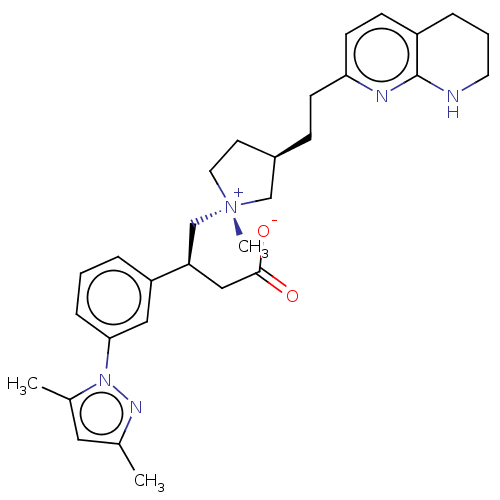
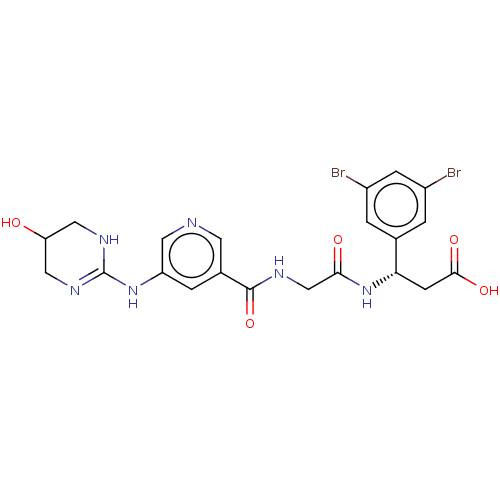
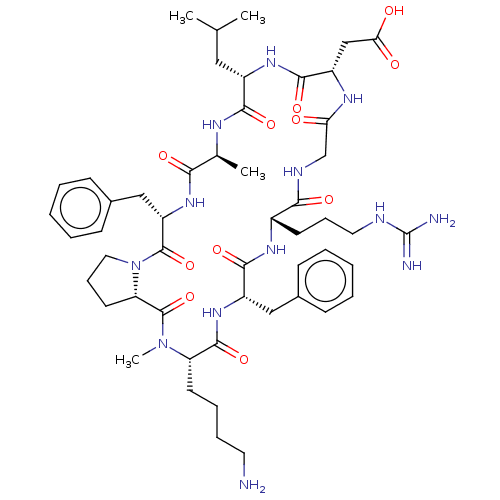
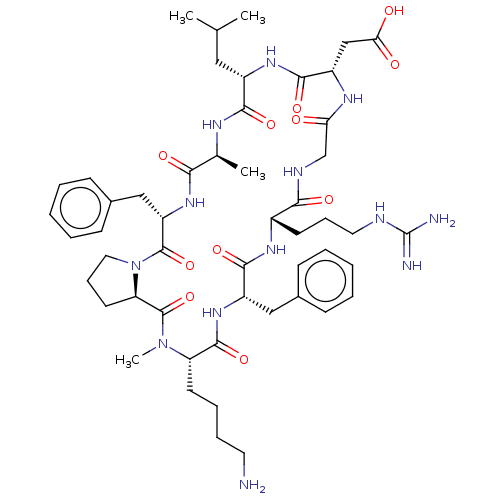
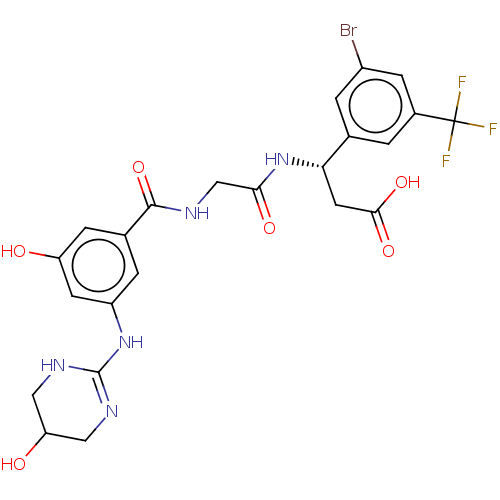
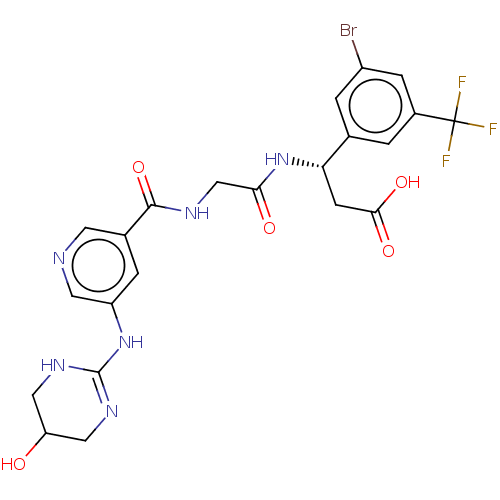
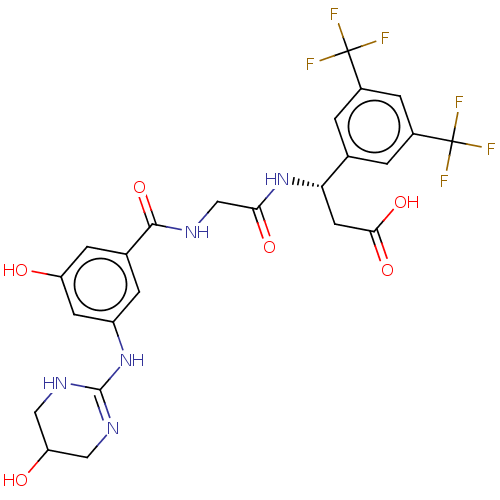
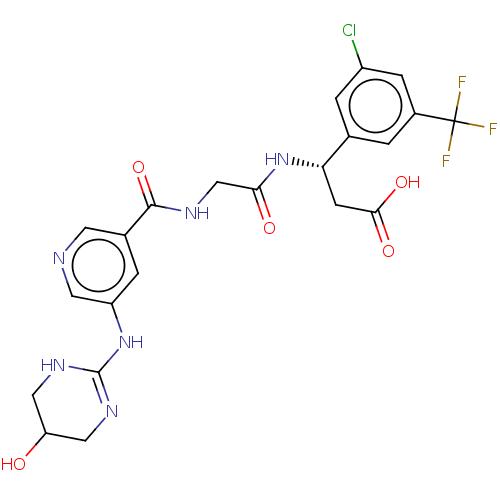
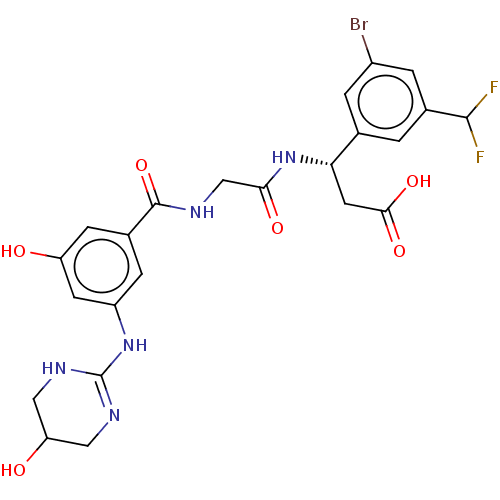
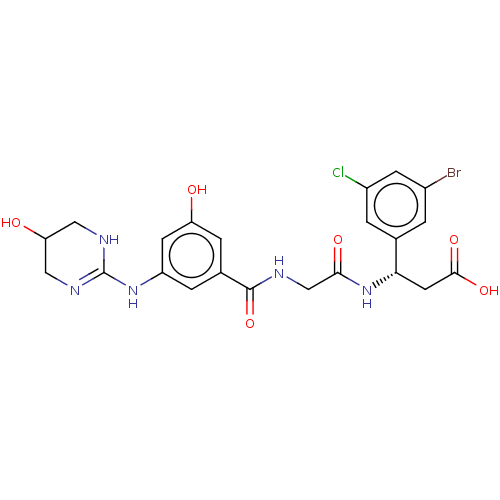
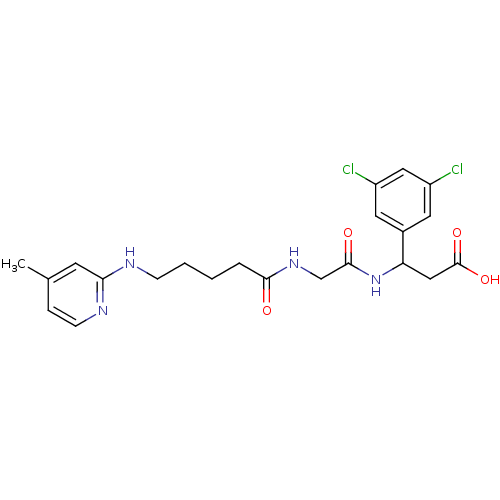
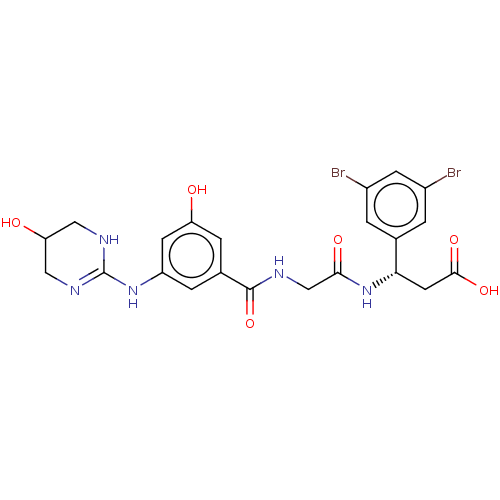
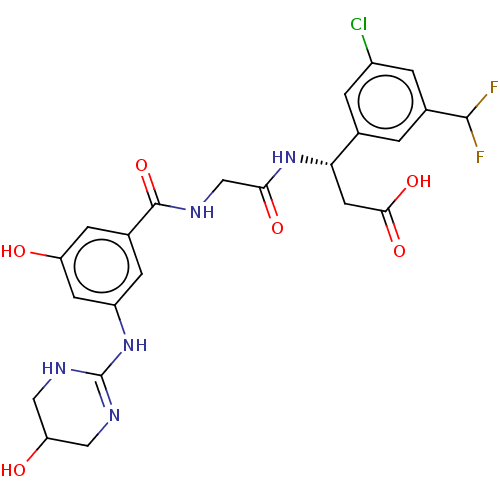
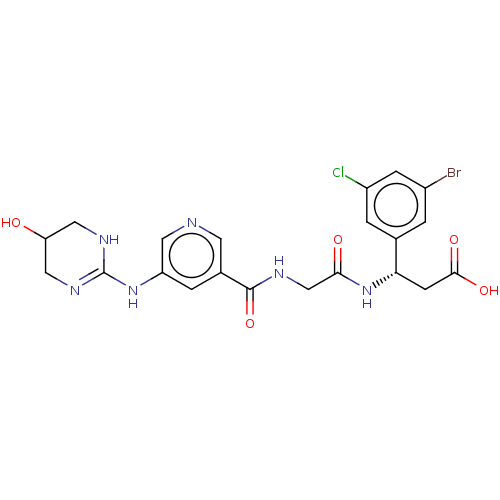
Affinity DataKi: 0.0100nMAssay Description:Binding affinity to human integrin alphaVbeta6 assessed as dissociation constant up to 48 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKd: 0.0158nMAssay Description:Binding affinity to human integrin alphaVbeta6 after 24 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKd: 0.0160nMAssay Description:Binding affinity to human integrin alphaVbeta6 after 24 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0251nMAssay Description:Binding affinity to human integrin alphaVbeta6 assessed as dissociation constant up to 48 hrs by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 0.0316nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0398nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0398nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0398nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0398nMAssay Description:Inhibition of integrin alphavbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0398nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of fibronectin binding to recombinant human alphaV-beta6 integrinMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of integrin alphavbeta6 (unknown origin) by Merck binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0501nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0631nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0631nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0631nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0631nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0794nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0794nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.0794nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Binding affinity to alphaVbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.107nMAssay Description:Inhibition of integrin alphavbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:The optimized protocol was validated by employing reference compounds such as Cilengitide (+Vβ3/αVβ5−VN interaction) and CWHM12 ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.150nMAssay Description:Inhibition of fibronectin binding to recombinant human alphaV-beta6 integrinMore data for this Ligand-Target Pair
Affinity DataEC50: 0.158nMAssay Description:Induction of integrin alphaVbeta6 receptor internalization in human primary human lung epithelial cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.174nMAssay Description:Inhibition of integrin alphavbeta6 (unknown origin) by radioligand binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.260nMAssay Description:Binding affinity to human integrin alphaVbeta6 incubated for 1 hr by ELISA based solid phase integrin binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.260nMAssay Description:Inhibition of LAP binding to human integrin alphavbeta6 receptor after 1 hr by solid-phase binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.260nMAssay Description:Inhibition of LAP binding to human integrin alphavbeta6 receptor after 1 hr by solid-phase binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.300nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of fibronectin binding to recombinant human alphaV-beta6 integrinMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMpH: 7.4 T: 2°CAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 μg/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM M...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Recombinant human LAP (R&D Systems, 246-LP) diluted to 0.25 g/mL in TBS+ buffer (25 mM Tris pH 7.4, 137 mM NaCl, 2.7 mM KCl, 1 mM CaCl2, 1 mM MgCl2, ...More data for this Ligand-Target Pair