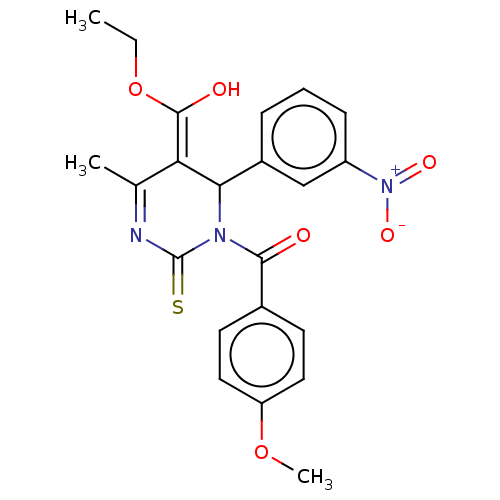
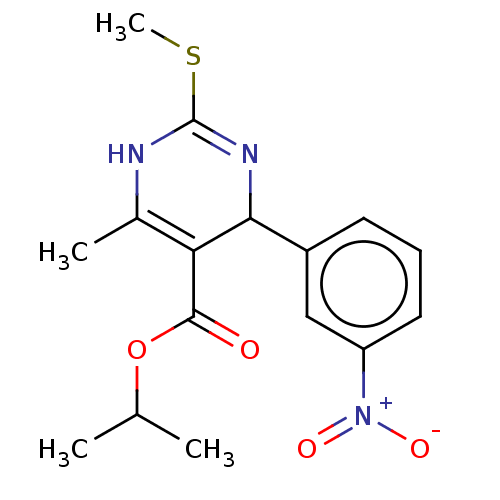
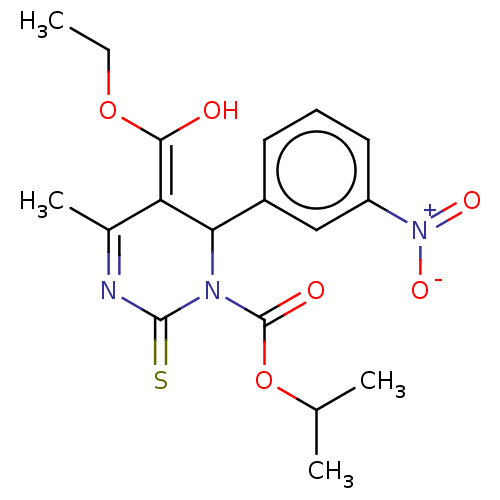
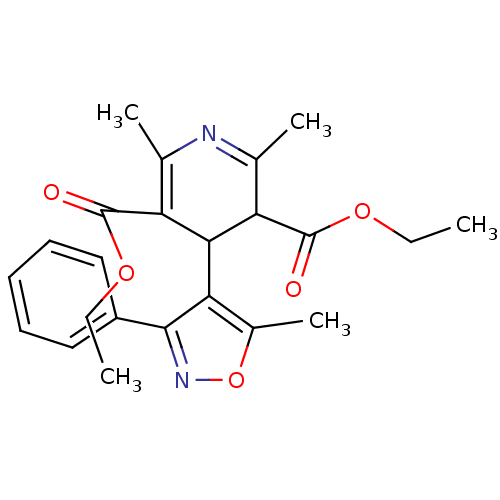
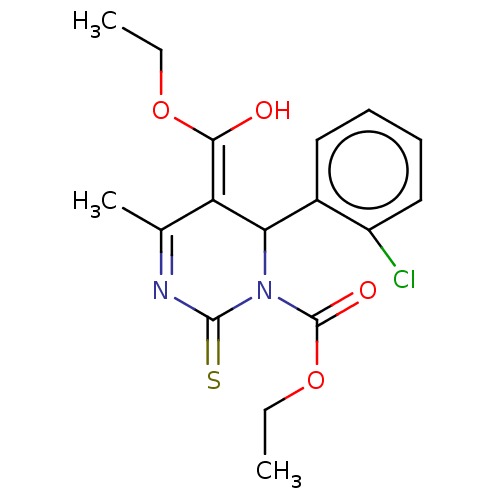
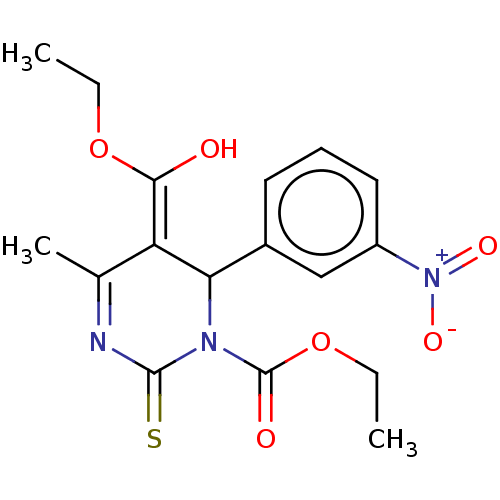
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.00810nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0100nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0130nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0150nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0170nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0220nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0270nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0330nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0450nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0460nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.140nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.150nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel dihydropyridine site of porcine cardiac sarcolemma membrane vesiclesMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.25nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
Affinity DataKd: 0.260nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel in guinea pig ventricular myocardium membranesMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.350nMAssay Description:Inhibition of voltage-gated L-type Ca channel (species unknown)More data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 0.390nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel of guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
Affinity DataKd: 0.400nMAssay Description:Displacement of [3H]nitrendipine from dihydropyridine receptor of guinea pig myocardial membranesMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel dihydropyridine site of porcine cardiac sarcolemma membrane vesiclesMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel dihydropyridine site of porcine cardiac sarcolemma membrane vesiclesMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 1nMAssay Description:In vitro vasorelaxant activity (calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 1.5nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 1.5nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 1.70nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:In vitro vasorelaxant activity (calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/L-type calcium channel subunit beta-3/calcium channel subunit alpha-2/delta-1(Human)
Novassay
US Patent
Novassay
US Patent
Affinity DataIC50: 2.10nMAssay Description:This section describes a scintillation proximity assay (SPA) to measure [3H] gabapentin ([3H]GBP) binding to membranes containing α2δ-1 and...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Inhibition of L-type calcium channel measured using 2-electrode voltage-clamp in human embryonic kidney cells heterologically expressing alpha-1C sub...More data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 2.30nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 2.60nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
Affinity DataKd: 2.60nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel in guinea pig ventricular myocardium membranesMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 2.70nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 2.80nMAssay Description:Inhibition of [3H]nitrendipine binding to guinea pig ileal longitudinal smooth muscleMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of L-type calcium channel measured using 2-electrode voltage-clamp in Chinese hamster ovary cells heterologically expressing alpha-1C subu...More data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 3.10nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 3.60nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:In vitro vasorelaxant activity (calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
Affinity DataKi: 4.10nMAssay Description:Displacement of [3H](+)-PN200-110 binding to calcium channels in guinea pig cardiac membranes.More data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/L-type calcium channel subunit beta-3/calcium channel subunit alpha-2/delta-1(Human)
Novassay
US Patent
Novassay
US Patent
Affinity DataIC50: 4.20nMAssay Description:This section describes a scintillation proximity assay (SPA) to measure [3H] gabapentin ([3H]GBP) binding to membranes containing α2δ-1 and...More data for this Ligand-Target Pair
Affinity DataKd: 5nMAssay Description:Displacement of [3H]nitrendipine from dihydropyridine receptor of guinea pig myocardial membranesMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 5.20nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Displacement of [3H](+)-PN200-110 binding to calcium channels in guinea pig cardiac membranes.More data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Cav1.2 current measured using QPatch automatic path clamp system in CHO cells expressing Cav1.2, beta-2 and alpha-2/delta-1 subunitsMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Inhibition of [3H]nitrendipine binding to L-type calcium channel in guinea pig ventricular myocardium membranesMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 14nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
Affinity DataKi: 14nMAssay Description:Inhibition of [3H]PN-200110 binding to Calcium channel in guinea pig heart membraneMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:In vitro vasorelaxant activity (voltage-gated calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
TargetVoltage-dependent L-type calcium channel subunit alpha-1C/alpha-1D/alpha-1F/alpha-1S(Human)
University of Oslo
Curated by ChEMBL
University of Oslo
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:In vitro vasorelaxant activity (calcium channel blocking activity) was determined with potassium-depolarized rabbit thoracic aortaMore data for this Ligand-Target Pair
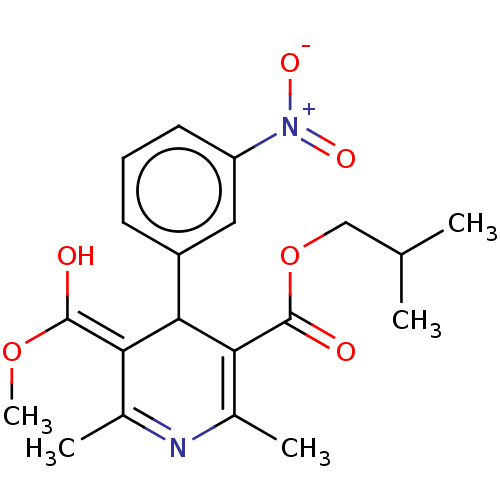
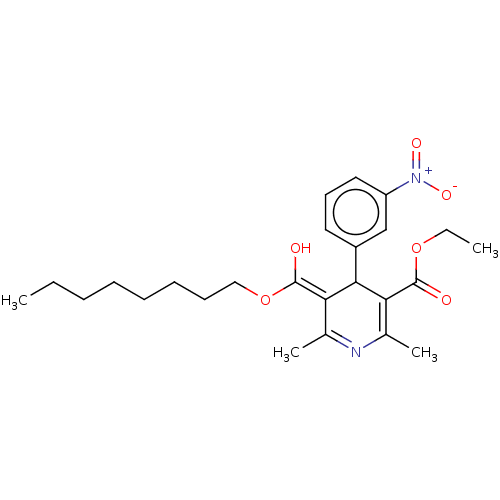
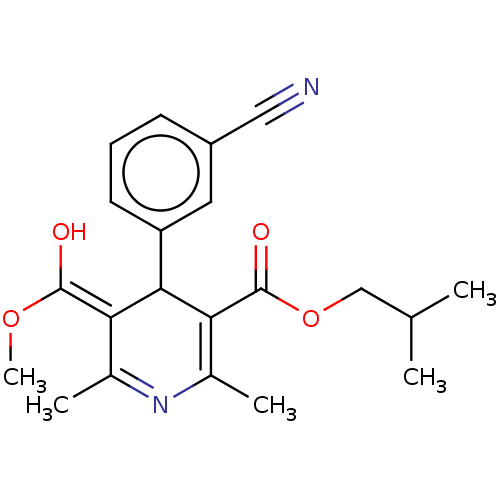
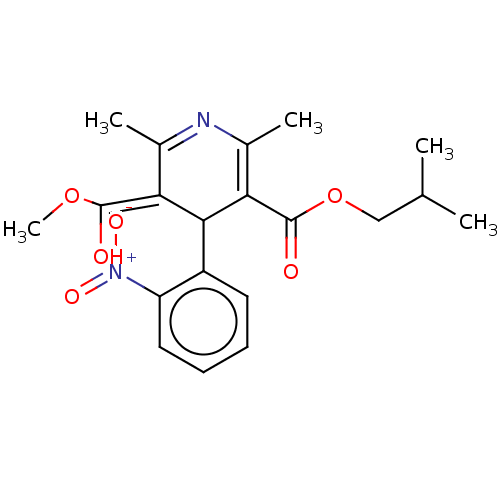
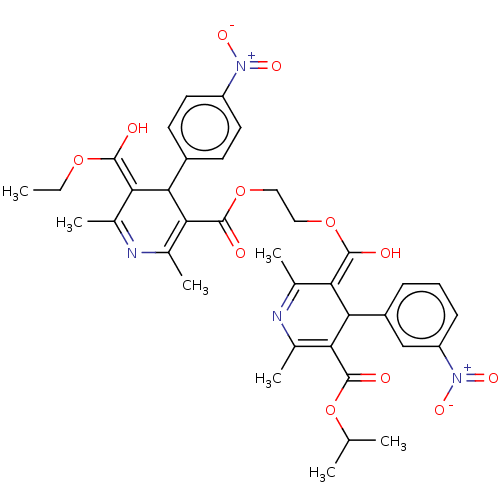
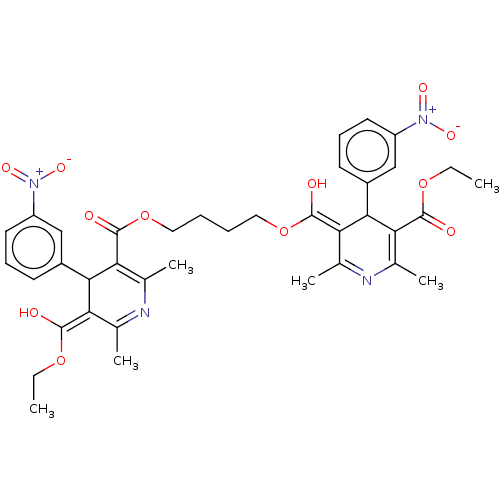
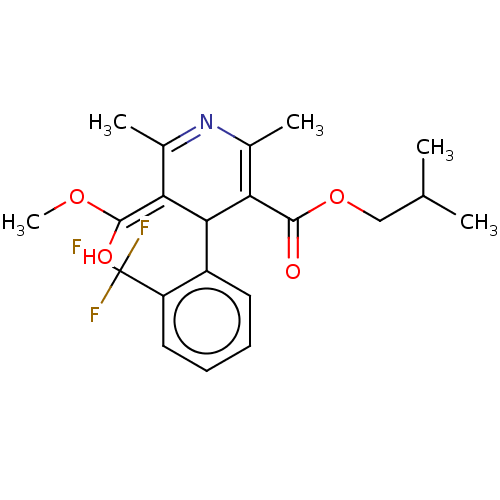
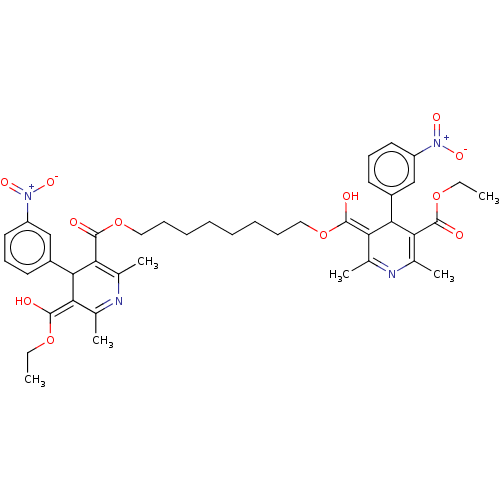
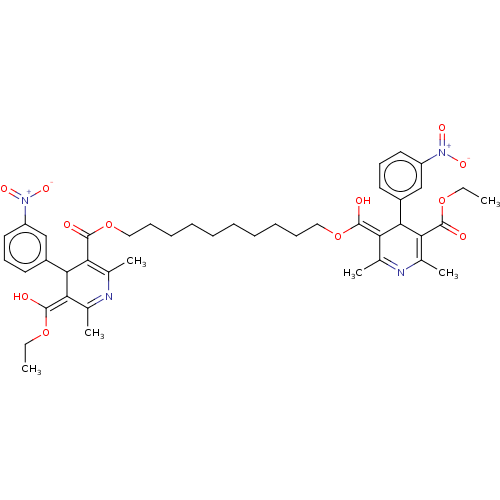
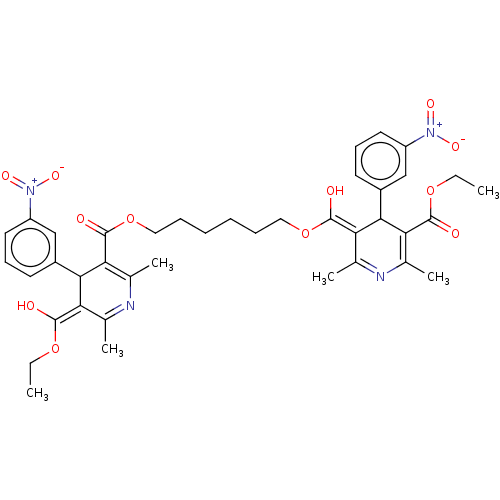
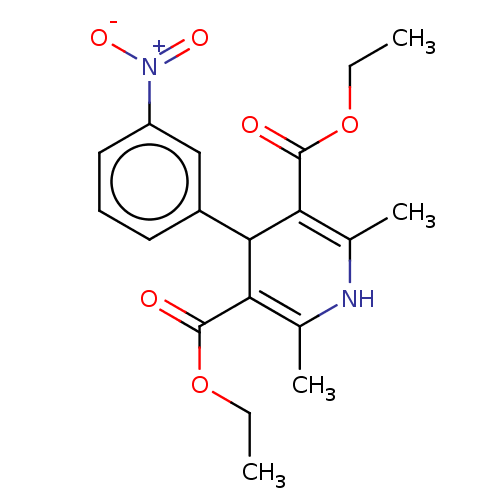
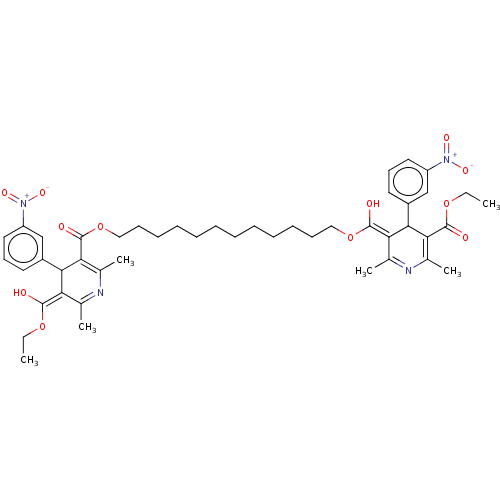
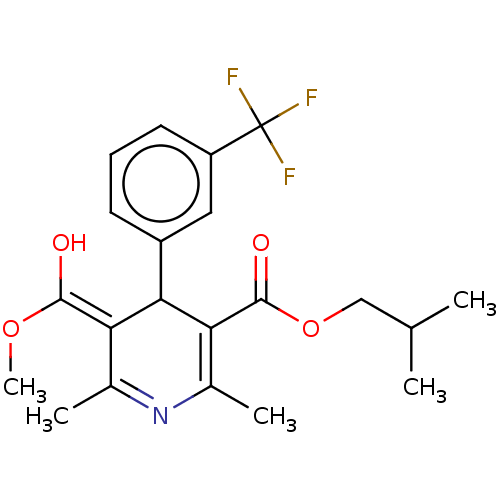
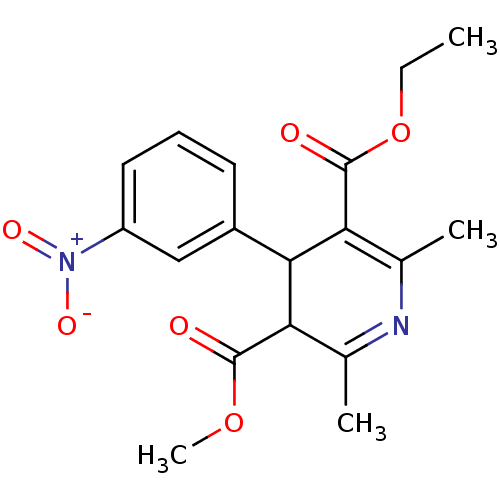
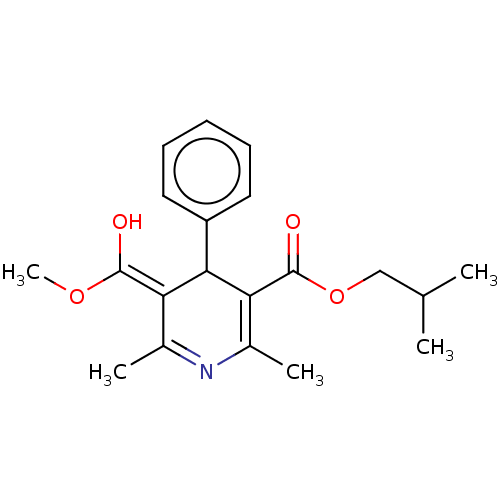
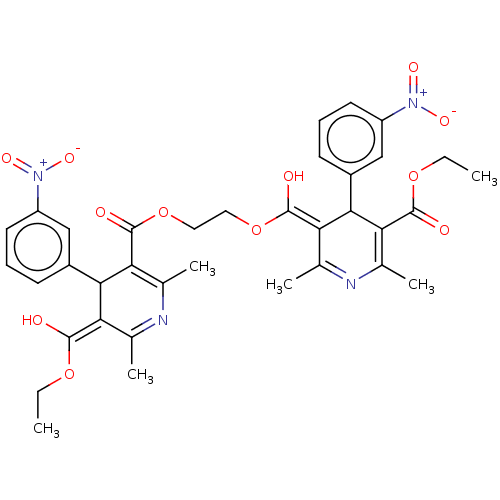
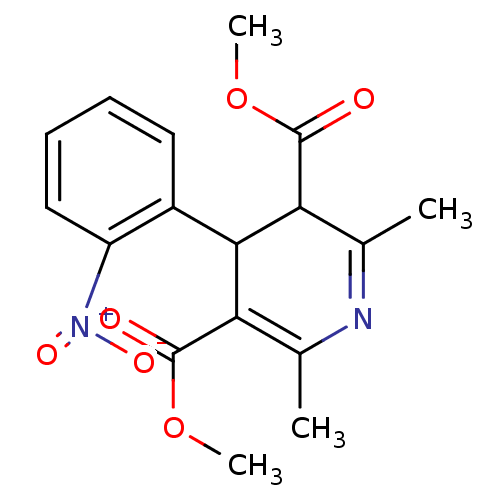
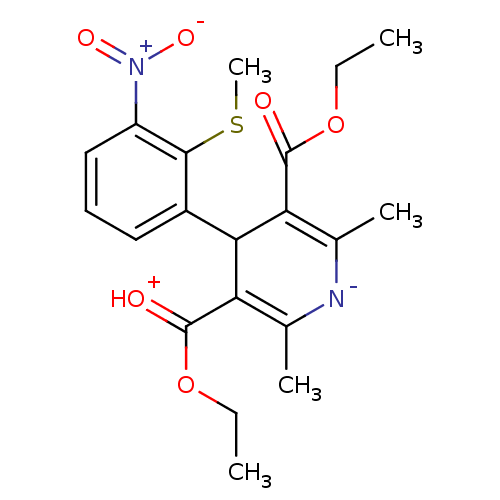
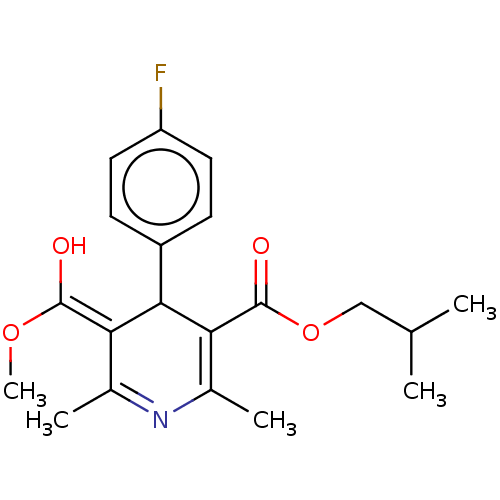
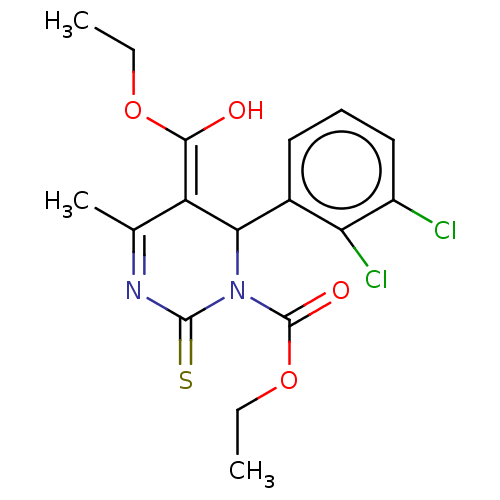

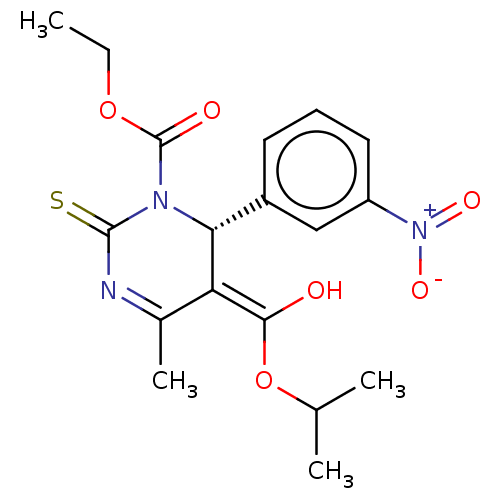
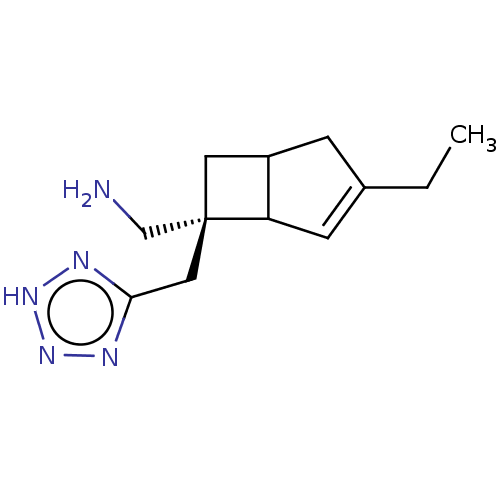
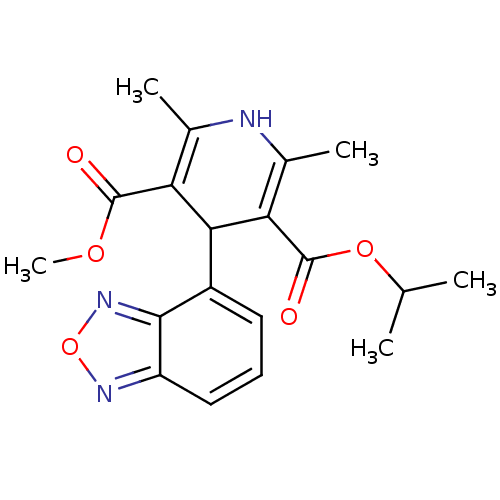
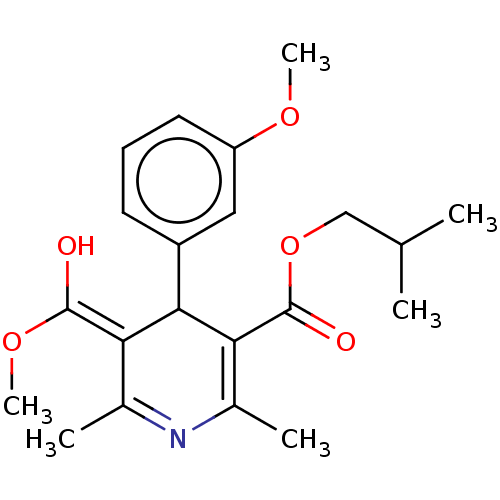
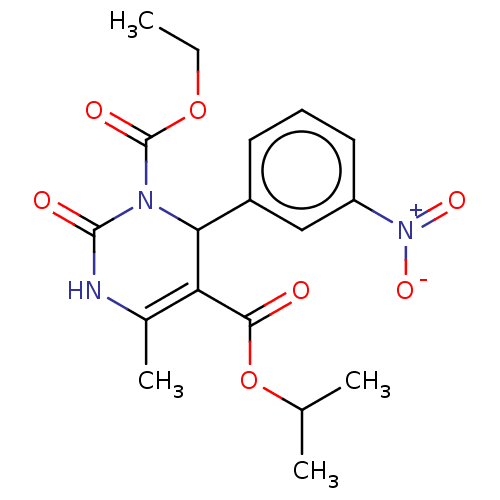
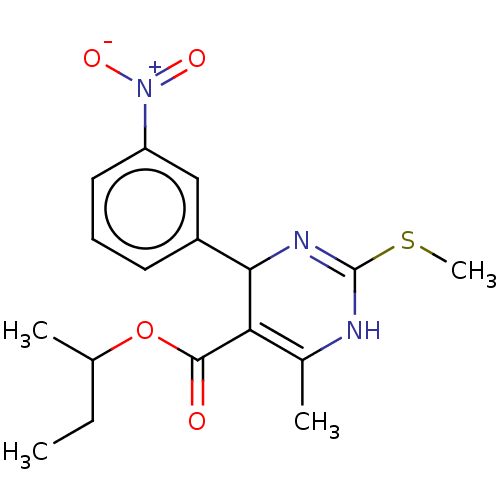
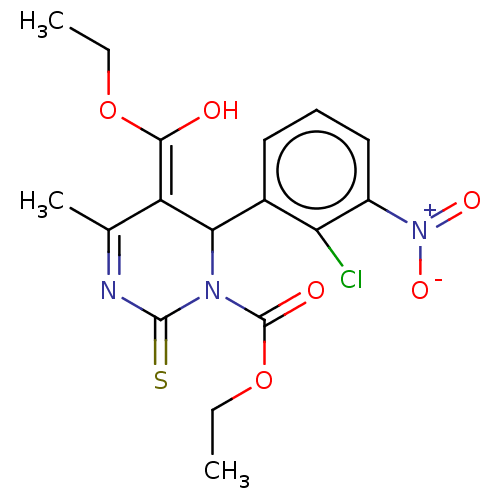
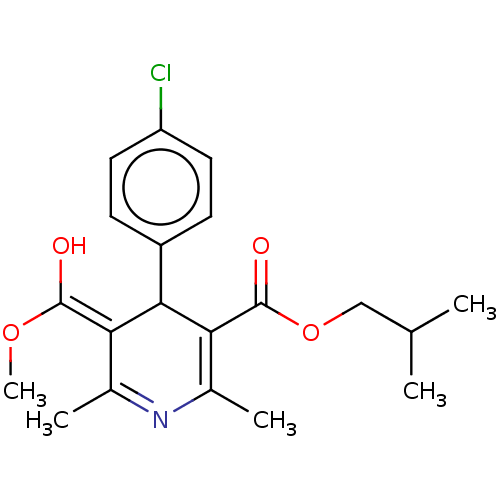
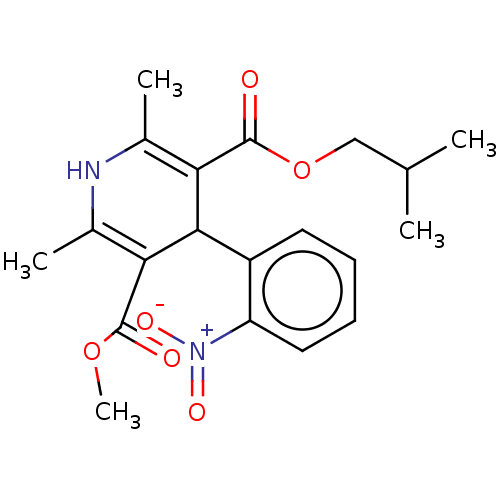
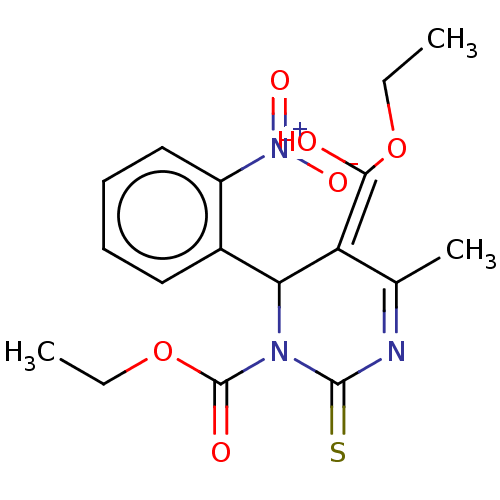
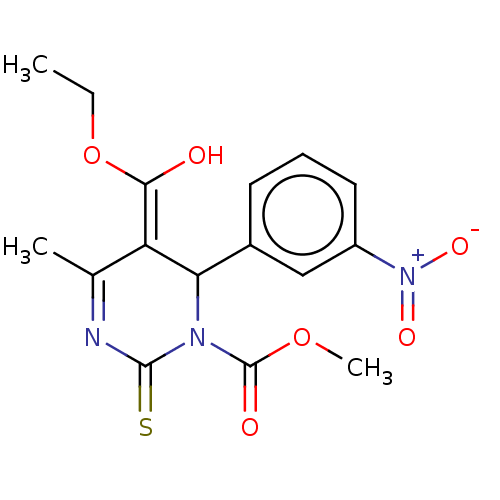
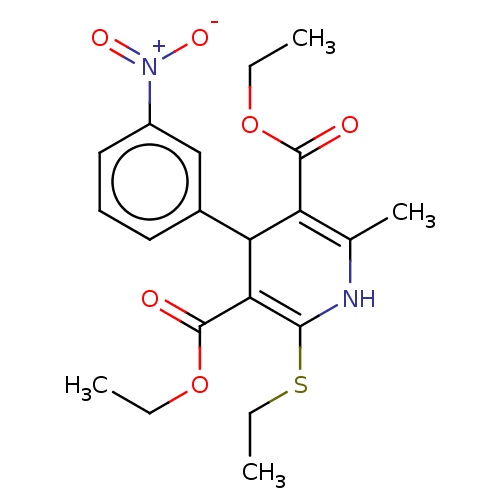
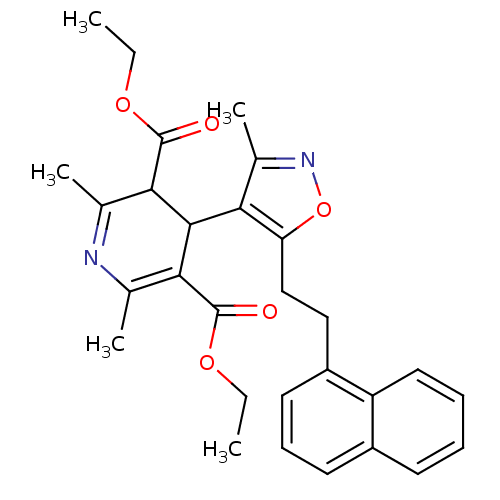
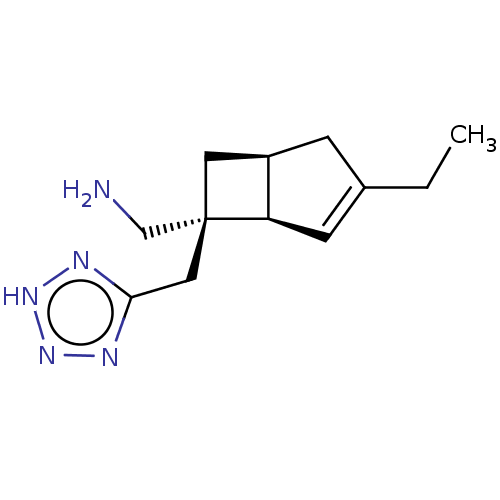
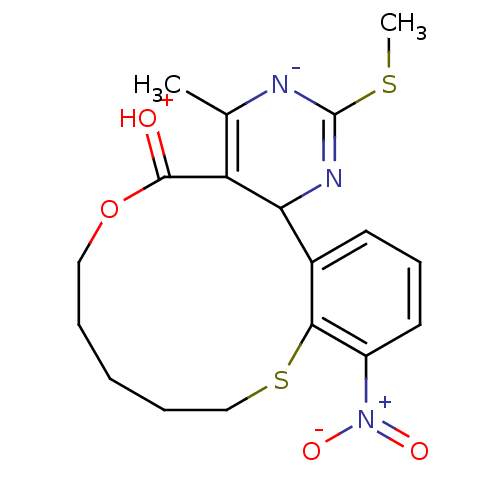
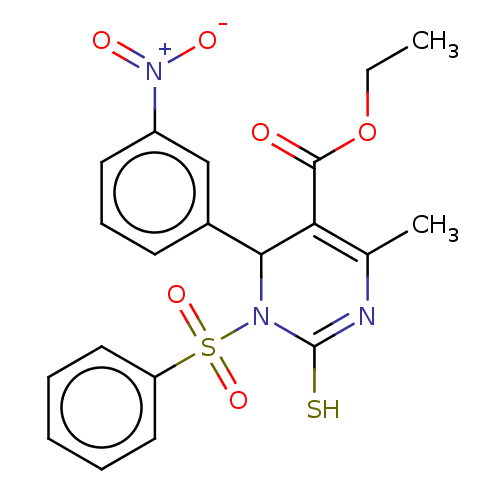
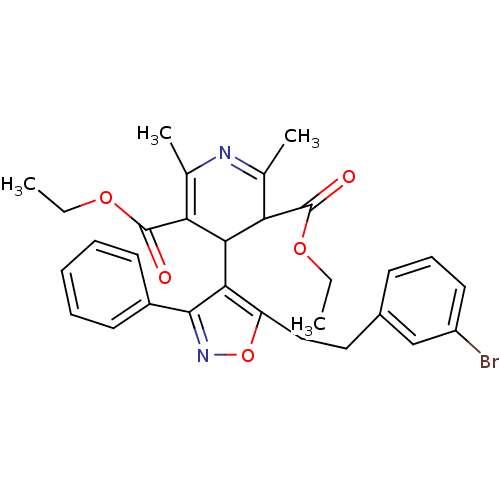
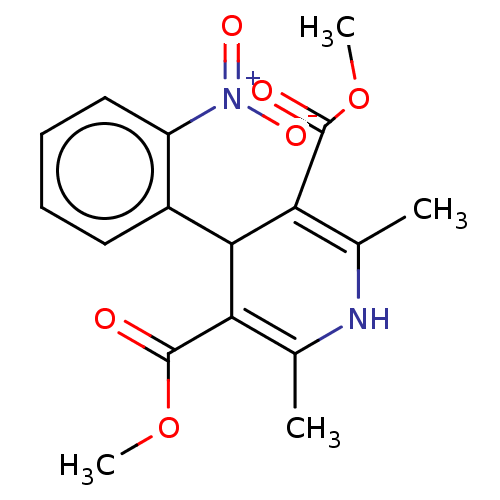
 3D Structure (crystal)
3D Structure (crystal)