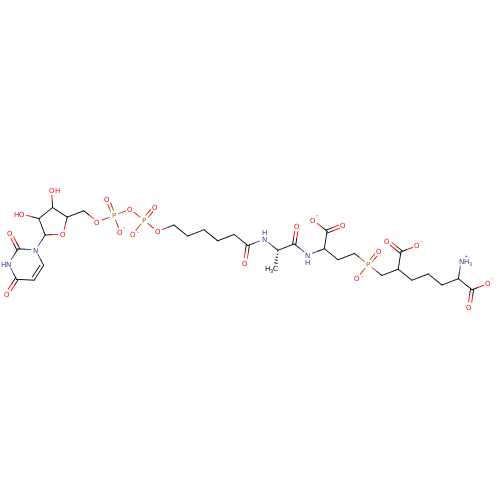
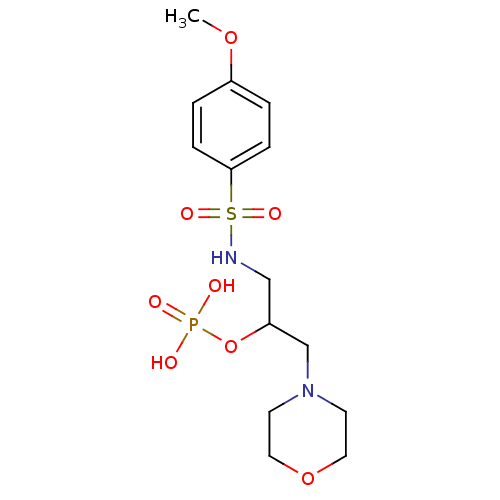
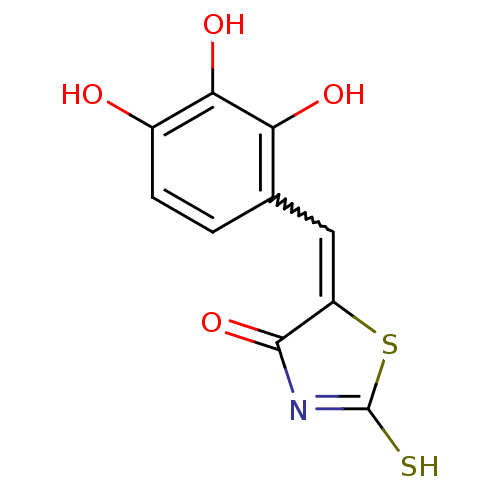
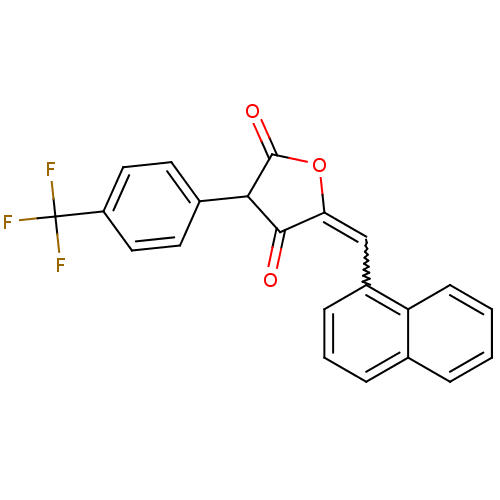
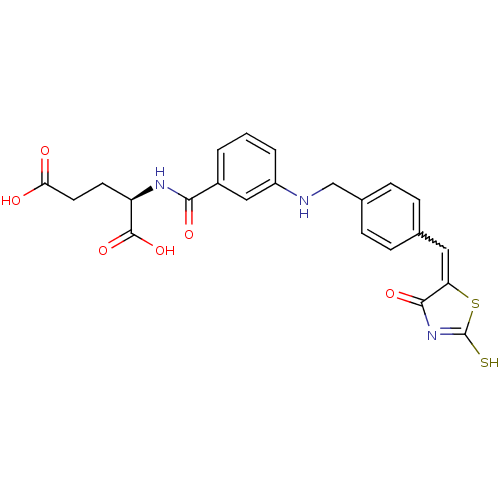
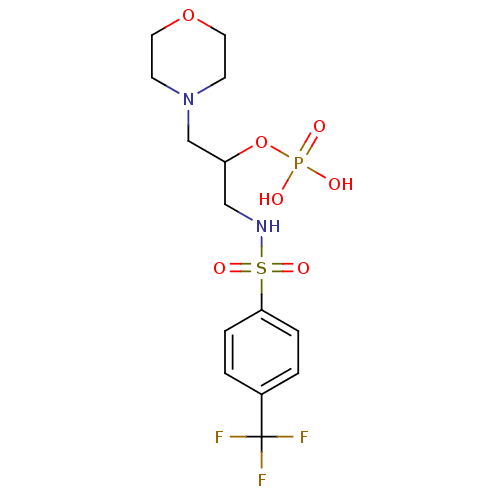
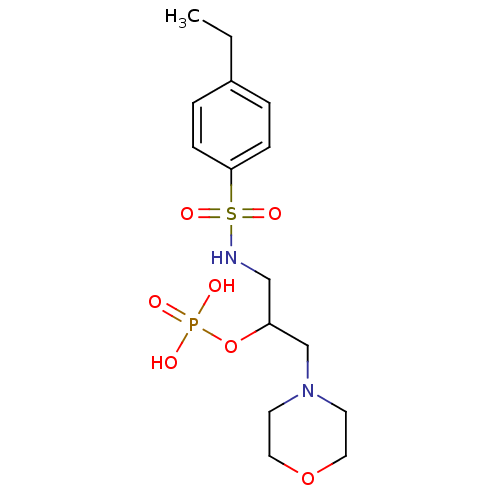
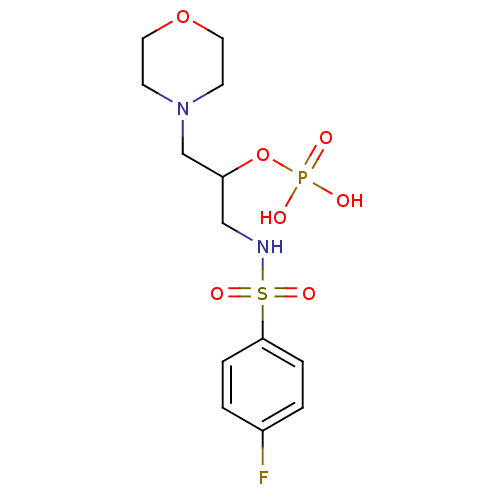
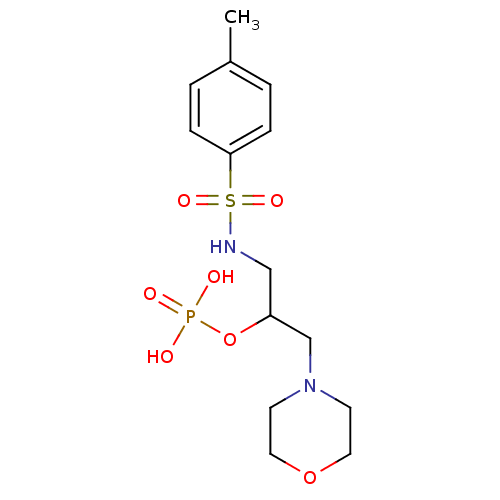
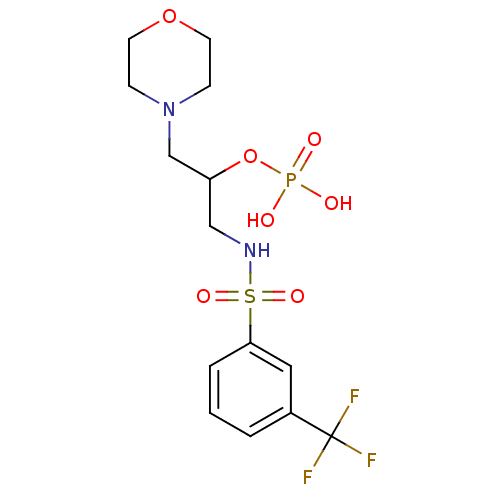
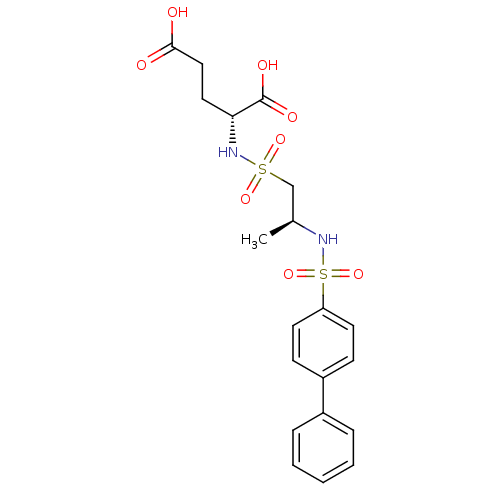
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.00E+3nMpH: 8.0 T: 2°CAssay Description:Compounds were tested for their inhibition of the addition of L-Ala, D-Glu, L-Lys or D-Ala-D-Ala to nucleotide precursors catalyzed by MurC from Esch...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 5.70E+4nMpH: 8.0 T: 2°CAssay Description:Compounds were tested for their inhibition of the addition of L-Ala, D-Glu, L-Lys or D-Ala-D-Ala to nucleotide precursors catalyzed by MurC from Esch...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 6.10E+4nMpH: 8.0 T: 2°CAssay Description:Compounds were tested for their inhibition of the addition of L-Ala, D-Glu, L-Lys or D-Ala-D-Ala to nucleotide precursors catalyzed by MurC from Esch...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 1.20E+5nMpH: 8.0 T: 2°CAssay Description:Compounds were tested for their inhibition of the addition of L-Ala, D-Glu, L-Lys or D-Ala-D-Ala to nucleotide precursors catalyzed by MurC from Esch...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 1.20E+5nMpH: 8.0 T: 2°CAssay Description:Compounds were tested for their inhibition of the addition of L-Ala, D-Glu, L-Lys or D-Ala-D-Ala to nucleotide precursors catalyzed by MurC from Esch...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana
Affinity DataIC50: 1.60E+5nMpH: 8.0 T: 2°CAssay Description:Compounds were tested for their inhibition of the addition of L-Ala, D-Glu, L-Lys or D-Ala-D-Ala to nucleotide precursors catalyzed by MurC from Esch...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoyl-L-alanyl-D-glutamate--L-lysine ligase(Staphylococcus aureus (strain NCTC 8325 / PS 47))
University of Ljubljana
University of Ljubljana