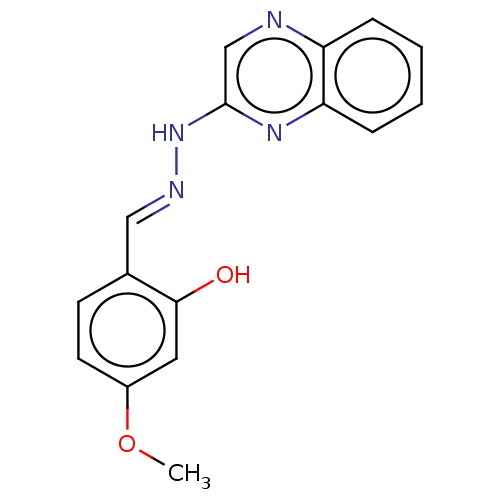
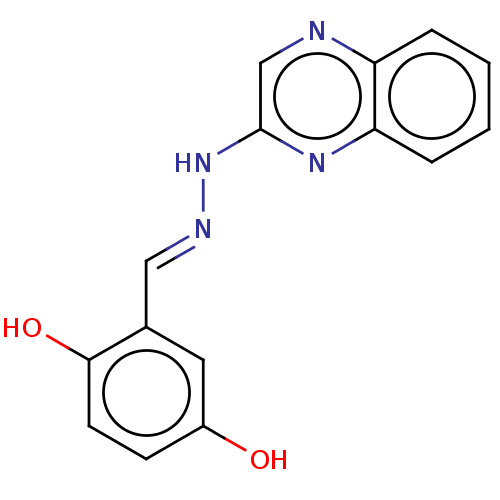
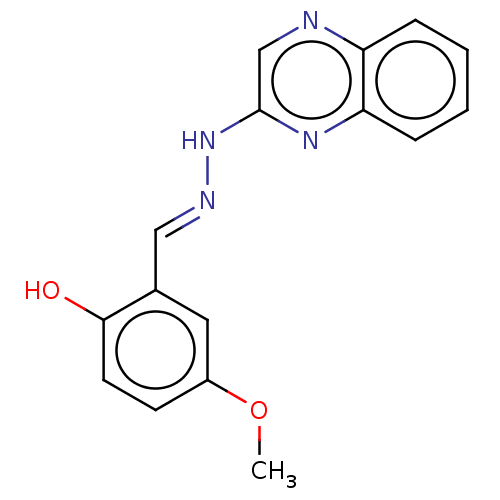
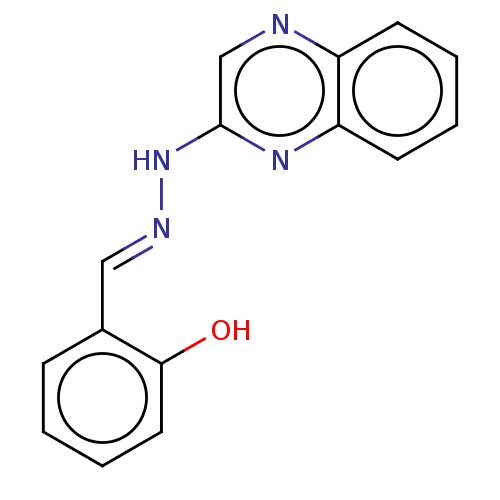
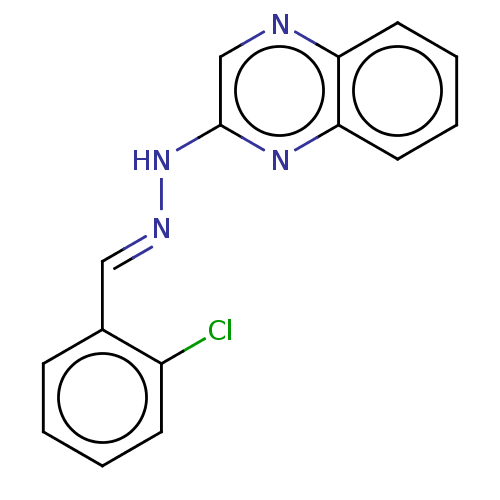
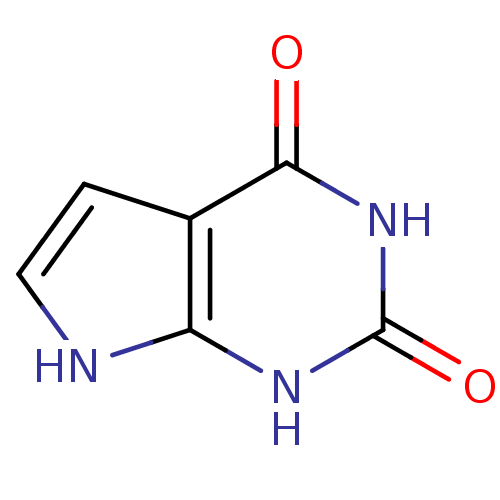
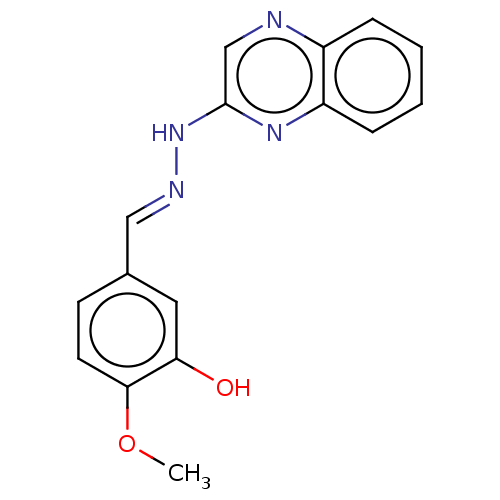
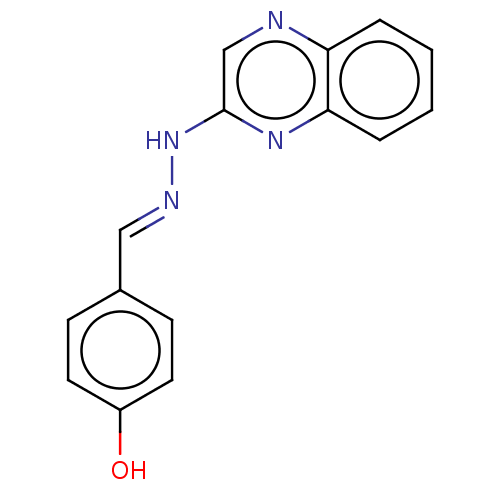
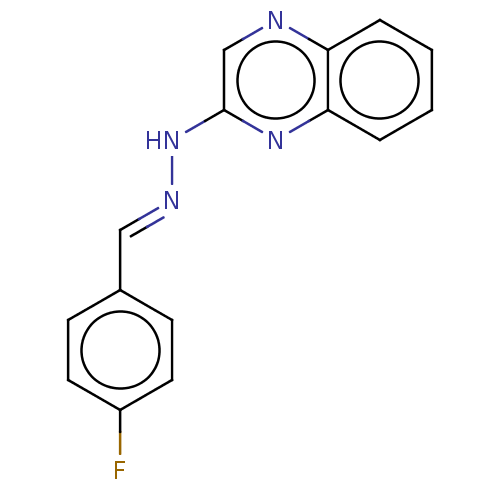
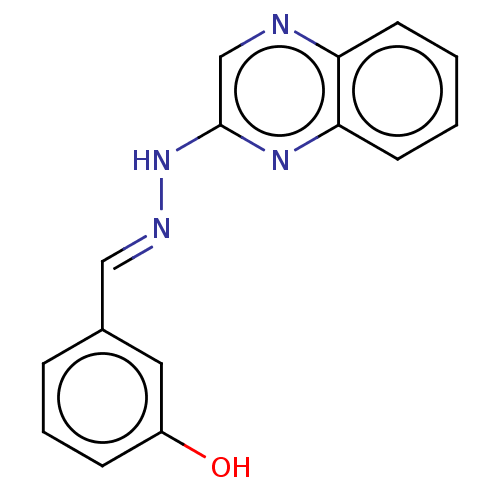
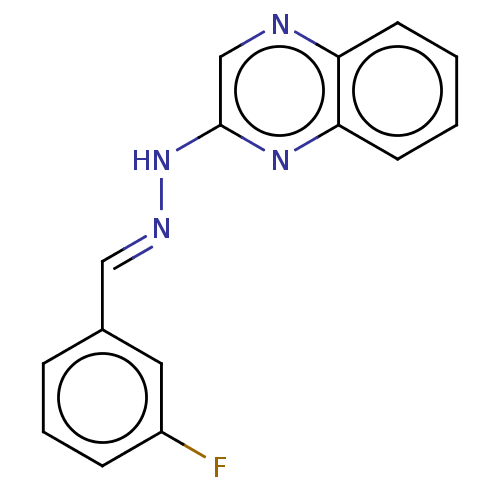
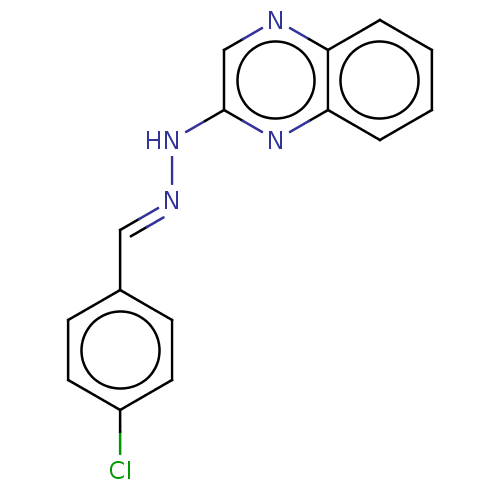
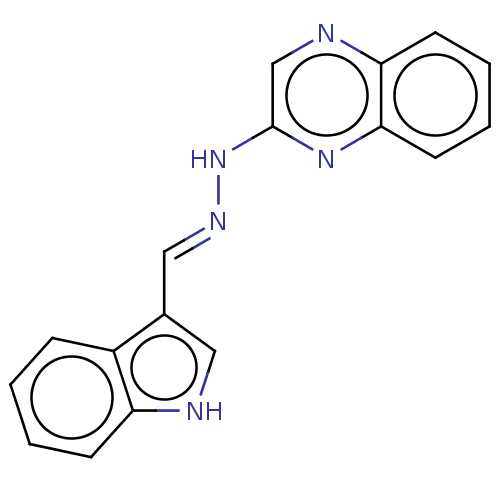
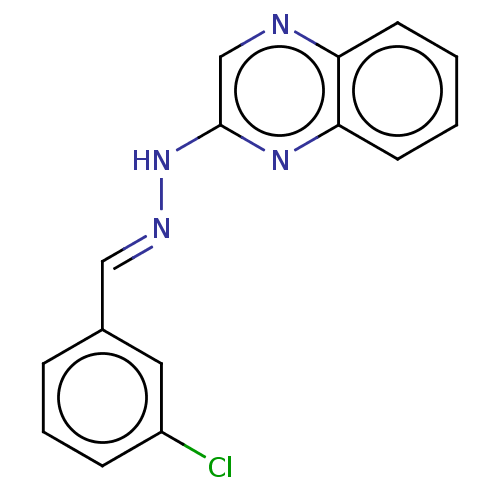
Affinity DataIC50: 3.20E+3nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 8.60E+3nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 9.10E+3nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 1.54E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 3.34E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 3.36E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 3.44E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 3.55E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 3.63E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 3.72E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 3.76E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 3.87E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 4.69E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 4.86E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 5.81E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 6.32E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 6.61E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 7.34E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 8.43E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair
Affinity DataIC50: 9.35E+4nMpH: 7.4 T: 2°CAssay Description:The Thymidine phosphorylase/PD-ECGF (E. coli) activity was determined by measuring the absorbance at 290 nm spectrophotometrically. In brief, total r...More data for this Ligand-Target Pair