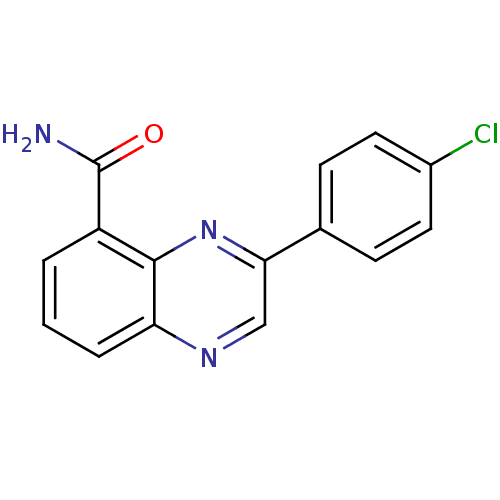
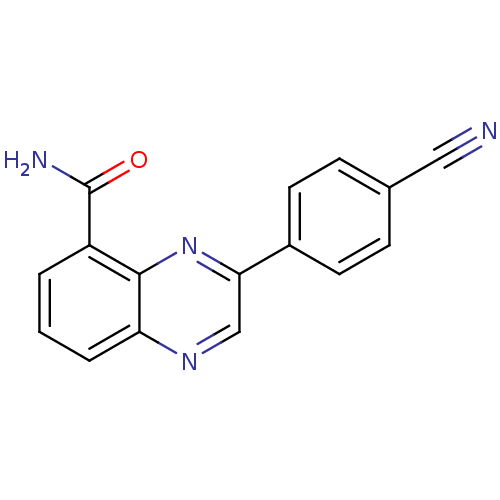
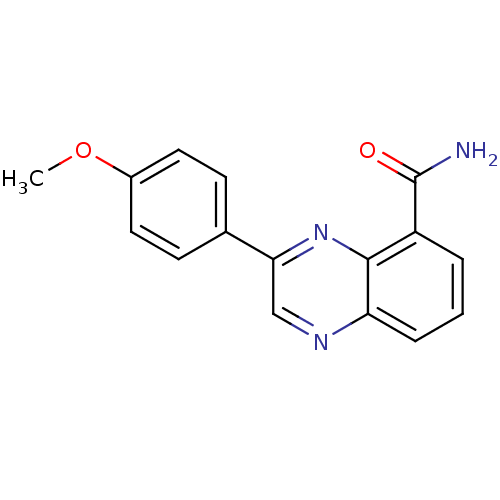
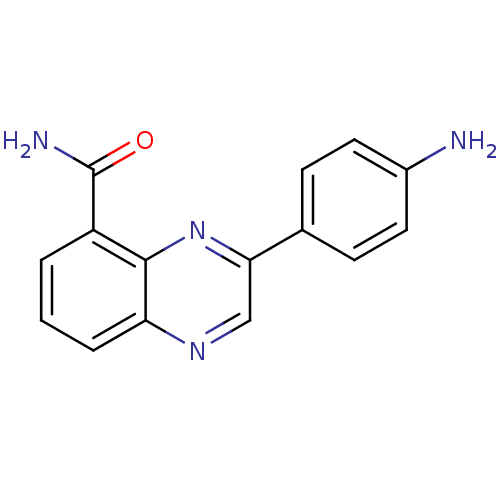
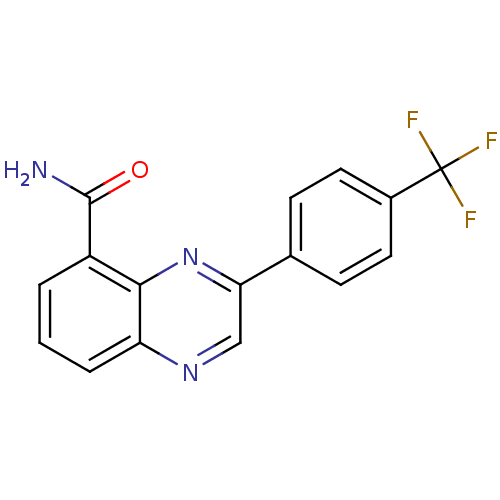
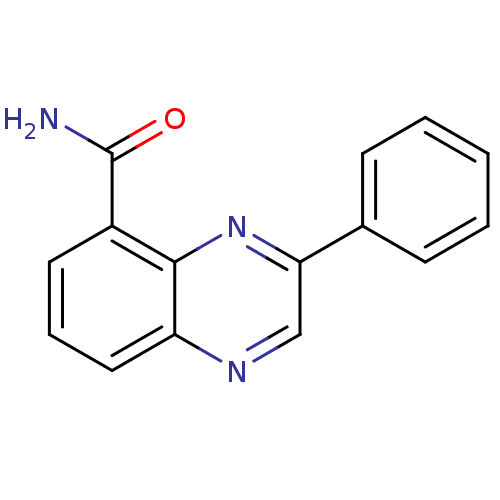
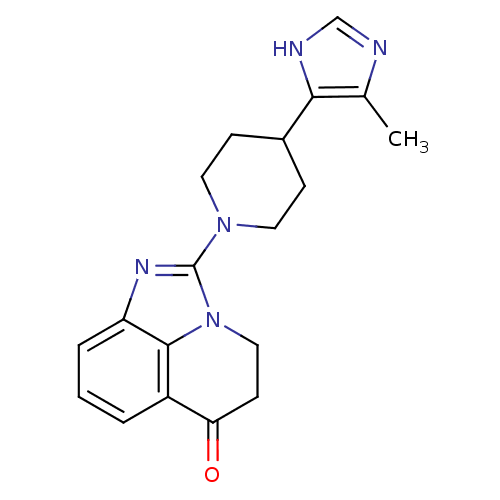
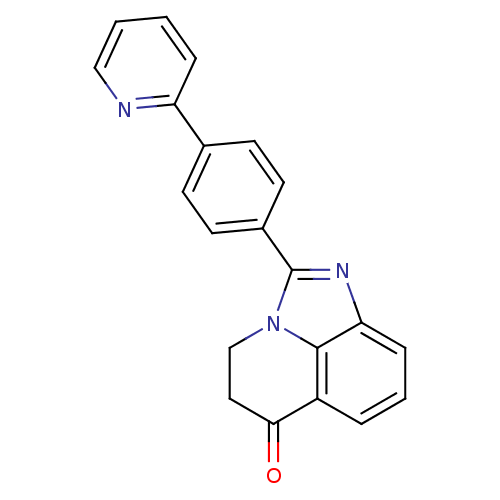
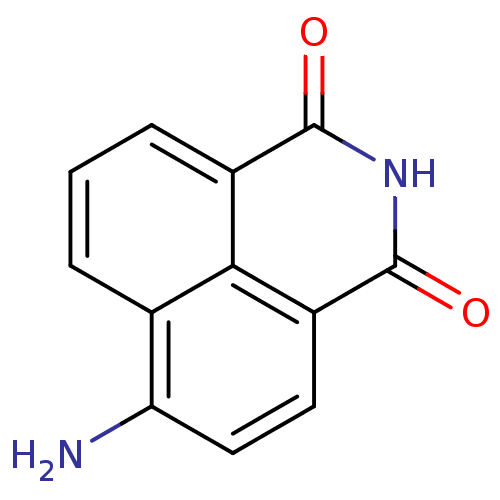
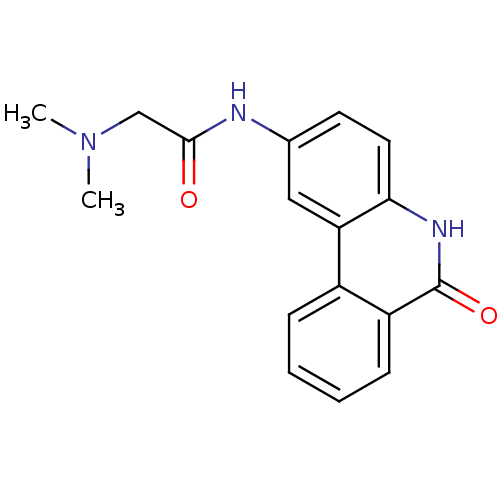
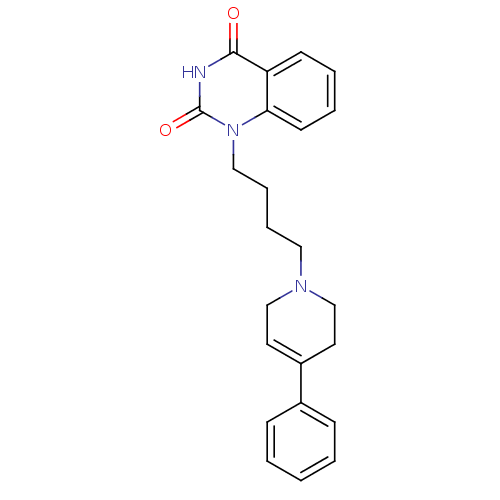
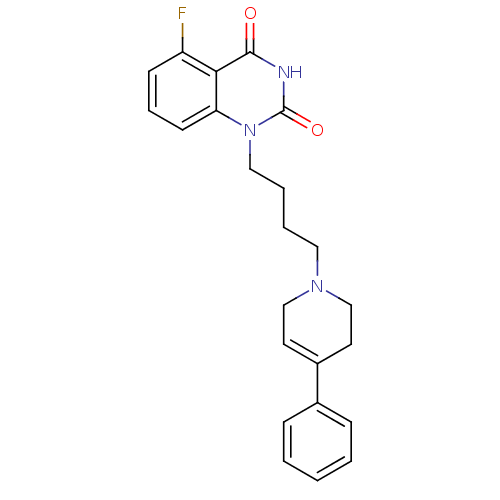
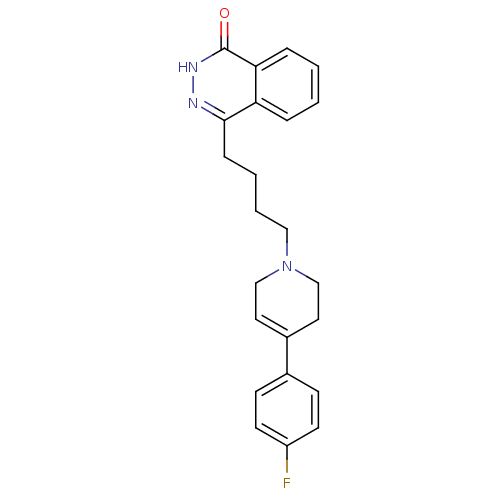
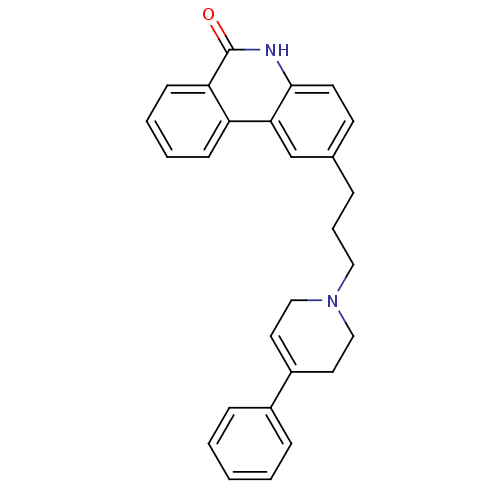
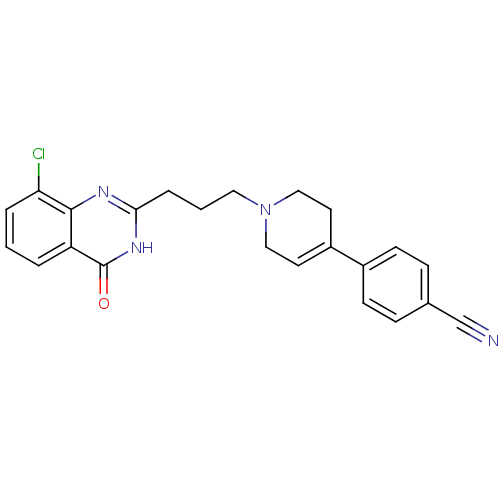
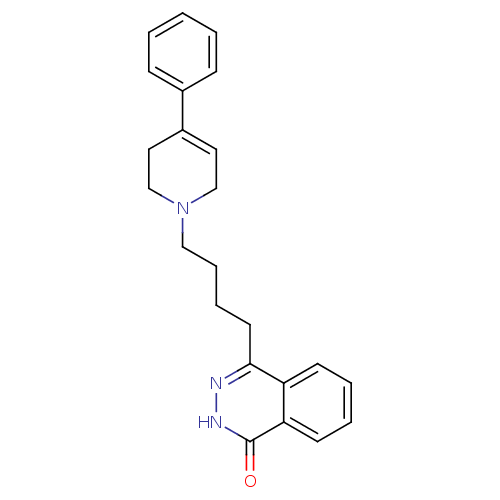
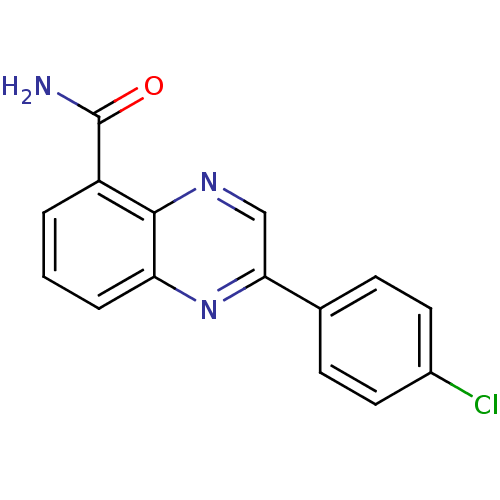
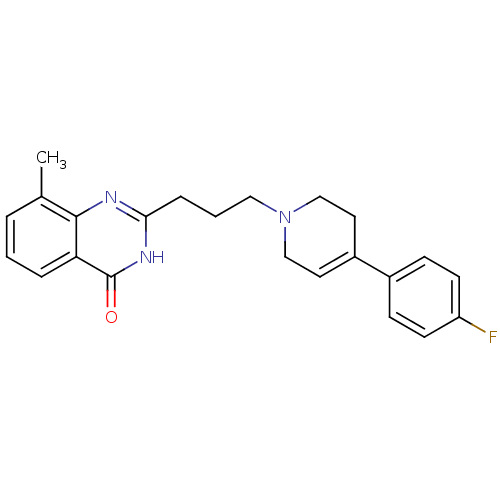
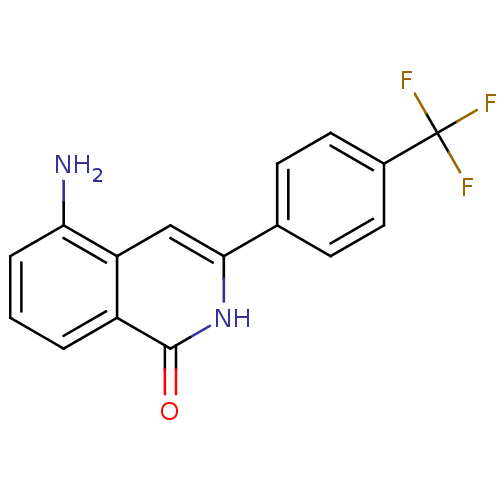
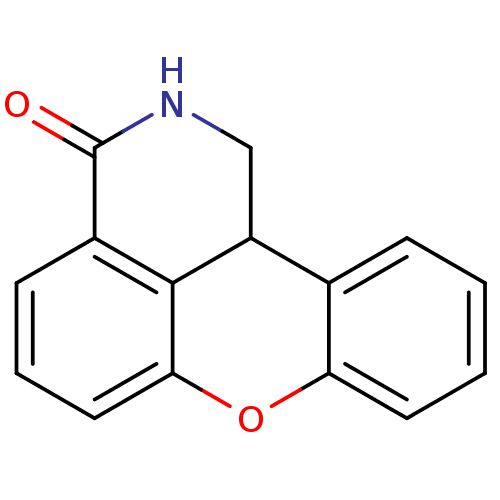
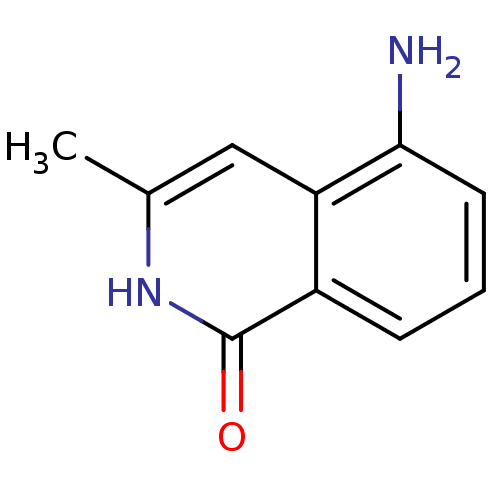
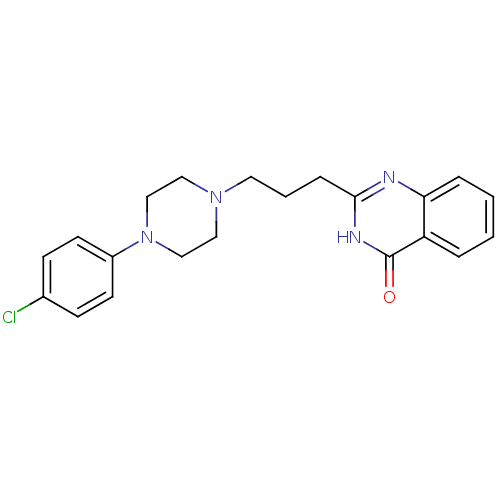
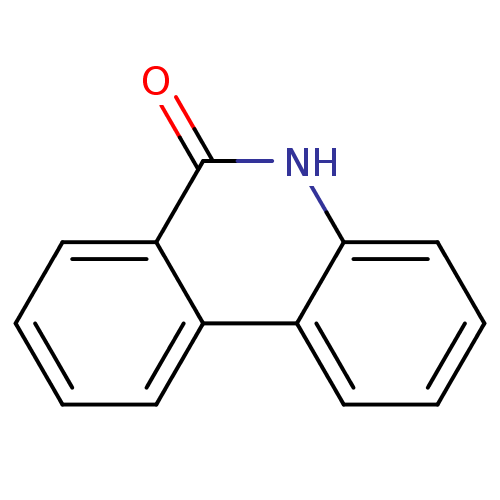
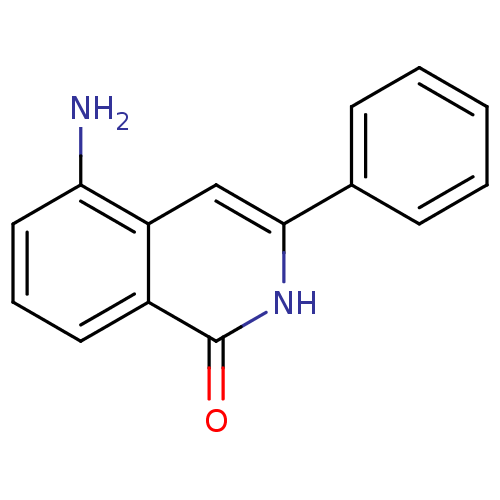
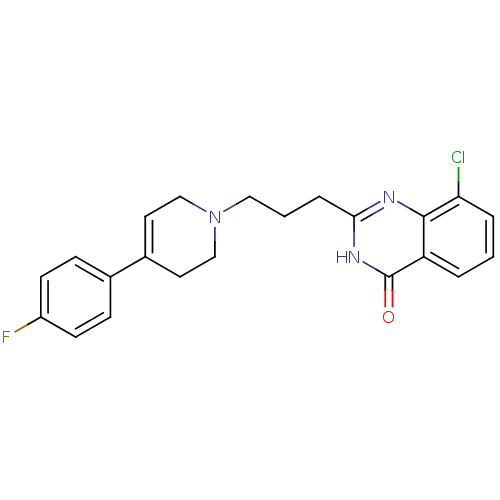
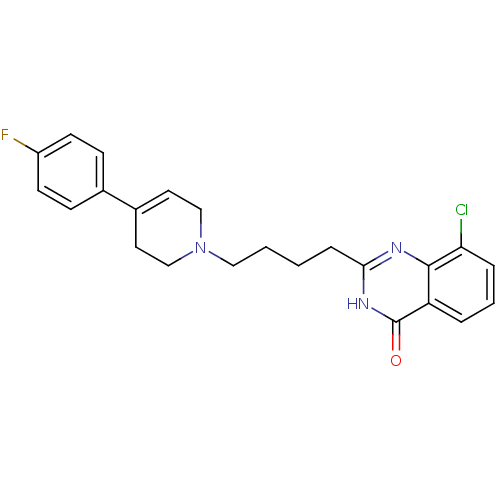
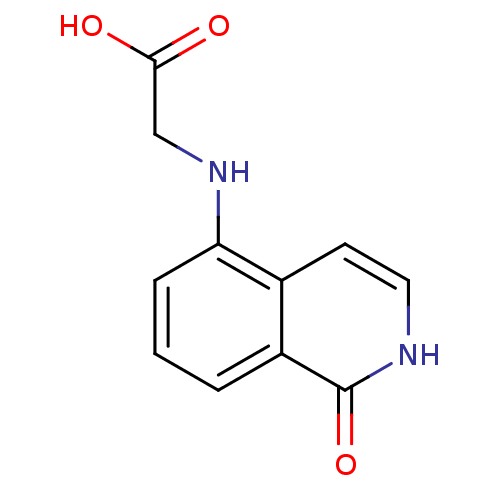
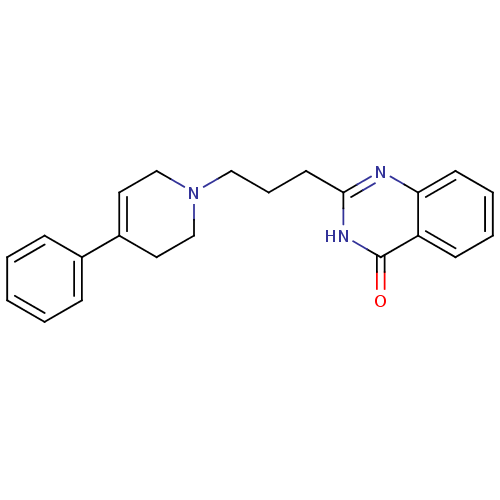
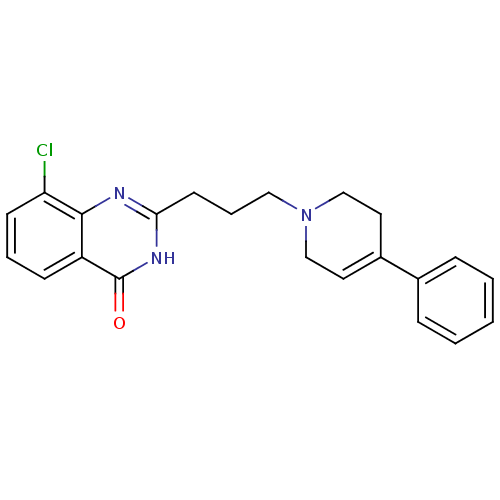
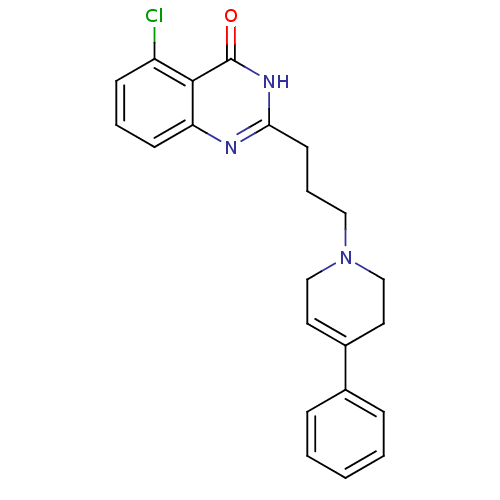
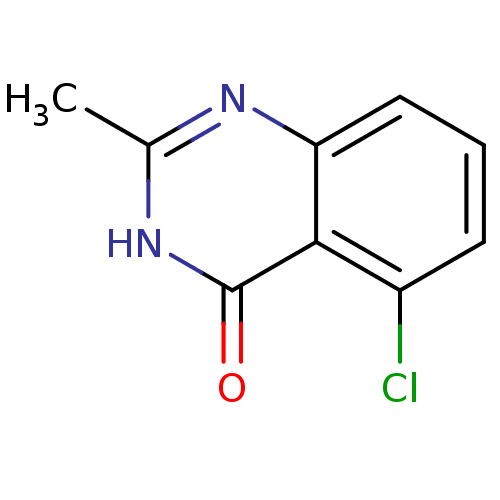
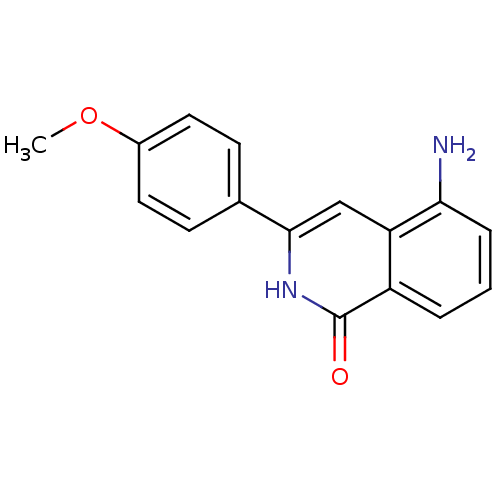
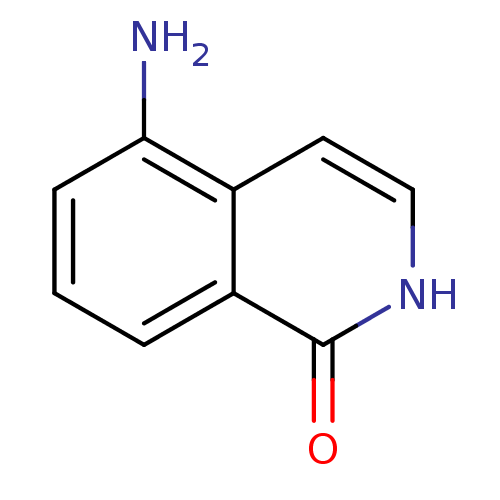
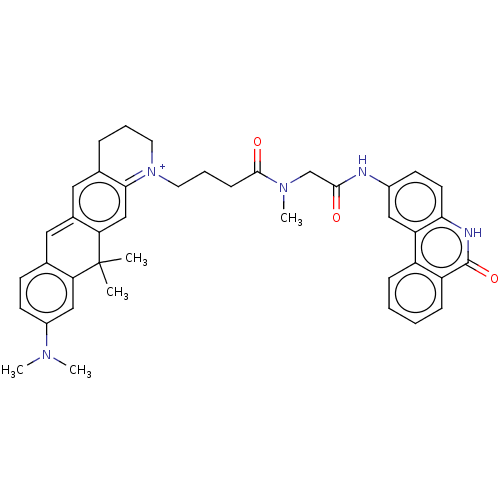
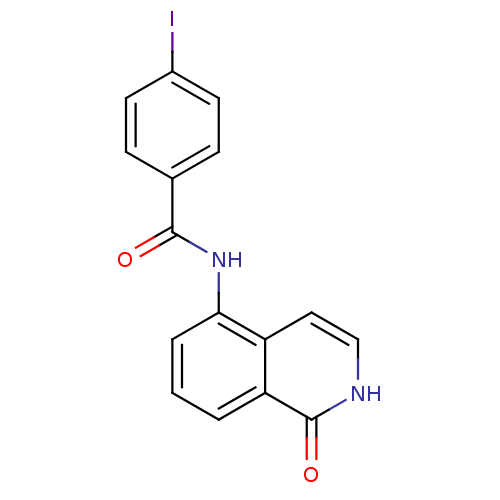
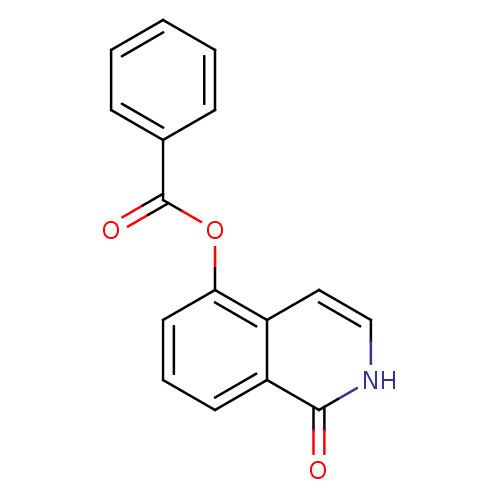
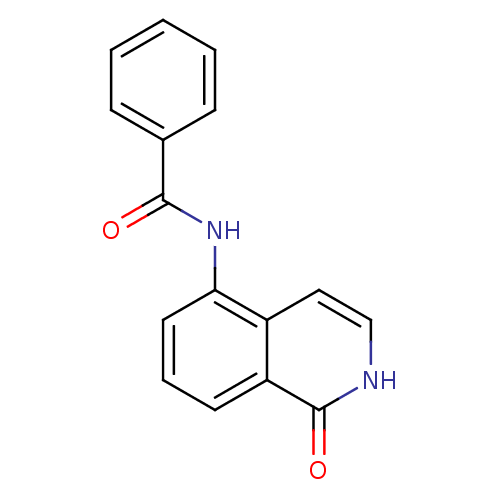
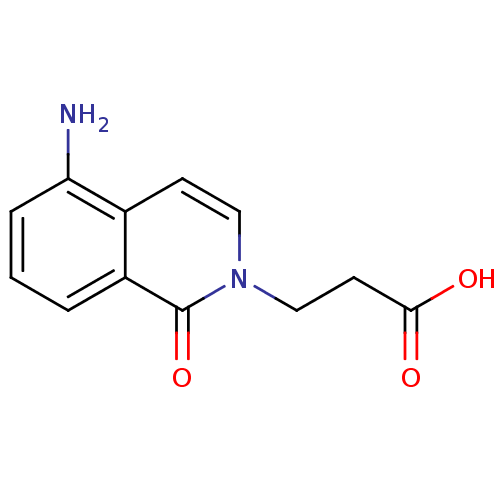
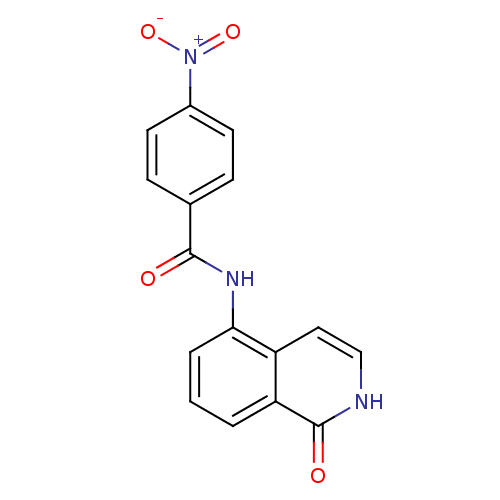
Affinity DataIC50: 0.700nMAssay Description:Inhibition of mouse recombinant PARP2 by chemiluminescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of mouse recombinant PARP2More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of mouse recombinant PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 7nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMpH: 7.8 T: 2°CAssay Description:The enzymatic reaction of the recombinant PARP was quantified by SPA. Radioactivity incorporated from [3H]NAD+ into PAR, and then being captured by P...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMpH: 7.8 T: 2°CAssay Description:The enzymatic reaction of the recombinant PARP was quantified by SPA. Radioactivity incorporated from [3H]NAD+ into PAR, and then being captured by P...More data for this Ligand-Target Pair
Affinity DataIC50: 32nMpH: 7.8 T: 2°CAssay Description:The enzymatic reaction of the recombinant PARP was quantified by SPA. Radioactivity incorporated from [3H]NAD+ into PAR, and then being captured by P...More data for this Ligand-Target Pair
Affinity DataIC50: 62nMpH: 7.8 T: 2°CAssay Description:The enzymatic reaction of the recombinant PARP was quantified by SPA. Radioactivity incorporated from [3H]NAD+ into PAR, and then being captured by P...More data for this Ligand-Target Pair
Affinity DataIC50: 70nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 84nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 84nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 87nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 90nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 90nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of mouse PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of mouse PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 167nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of mouse PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 186nMpH: 7.8 T: 2°CAssay Description:The enzymatic reaction of the recombinant PARP was quantified by SPA. Radioactivity incorporated from [3H]NAD+ into PAR, and then being captured by P...More data for this Ligand-Target Pair
Affinity DataIC50: 260nMAssay Description:Inhibition of mouse PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 300nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 331nMpH: 7.8 T: 2°CAssay Description:The enzymatic reaction of the recombinant PARP was quantified by SPA. Radioactivity incorporated from [3H]NAD+ into PAR, and then being captured by P...More data for this Ligand-Target Pair
Affinity DataIC50: 447nMpH: 7.8 T: 2°CAssay Description:The enzymatic reaction of the recombinant PARP was quantified by SPA. Radioactivity incorporated from [3H]NAD+ into PAR, and then being captured by P...More data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Inhibition of mouse PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 500nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 550nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 608nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 610nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 630nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 660nMT: 2°CAssay Description:To assess the inhibitory activity of novel inhibitors, the PARP enzyme assay was carried out in reaction mixture consisting of activated salmon teste...More data for this Ligand-Target Pair
Affinity DataIC50: 730nMAssay Description:Inhibition of mouse PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 830nMAssay Description:Inhibition of mouse PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+3nMAssay Description:Inhibition of mouse PARP-2More data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+3nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.05E+3nMAssay Description:The assay is based on the use of a probe of formula (IP) that binds to the NAD binding pocket and takes advantage of the significant change in the po...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of mouse recombinant PARP-2 incubated for 60 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of recombinant full length mouse PARP-2 after 10 mins using [3H]NAD+ by solution-phase assayMore data for this Ligand-Target Pair

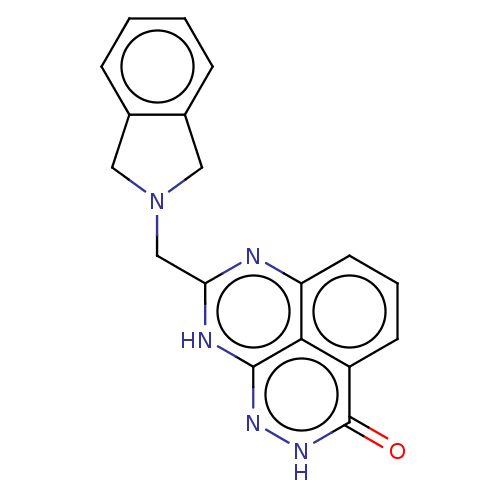
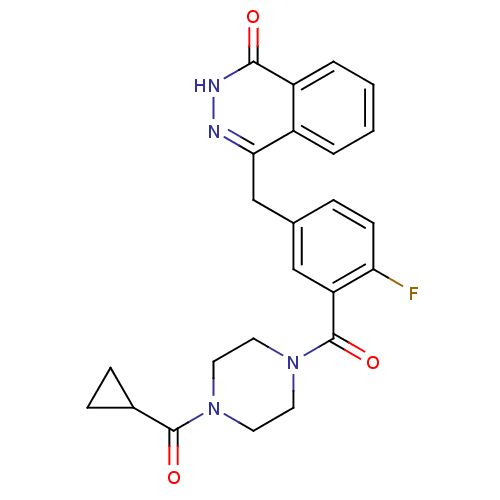
 3D Structure (crystal)
3D Structure (crystal)