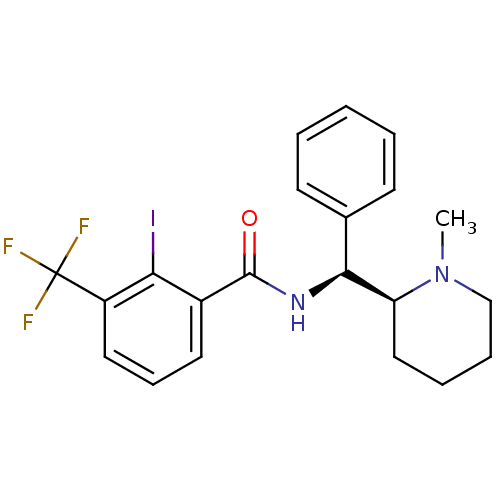
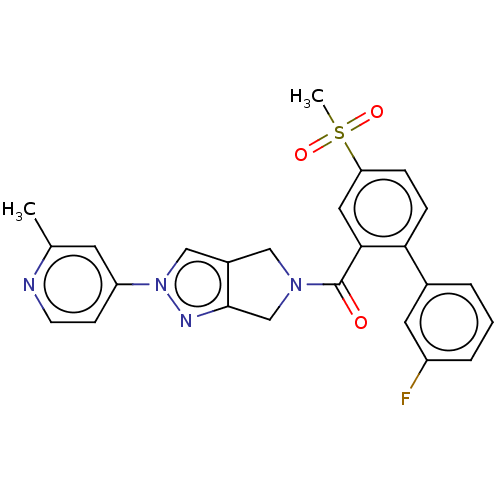
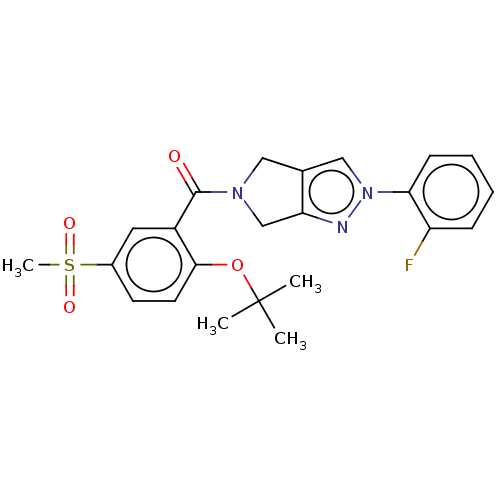
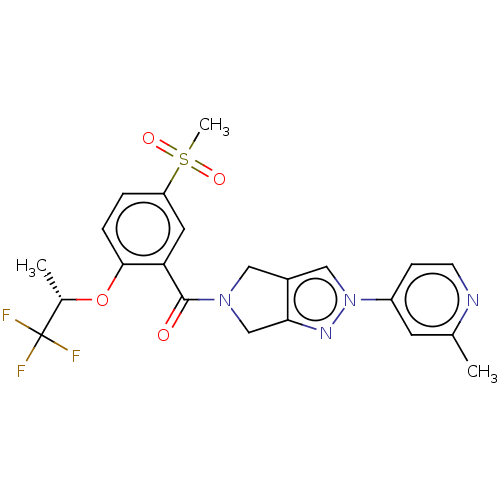
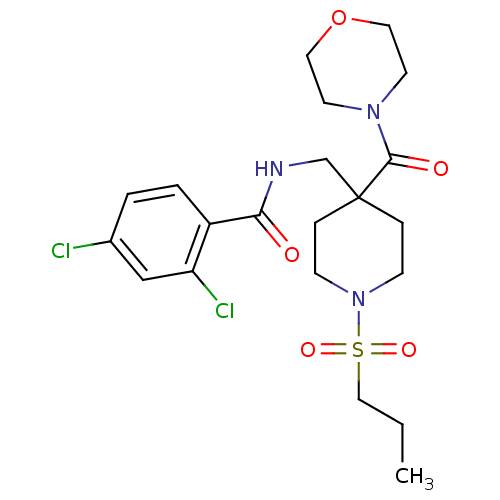
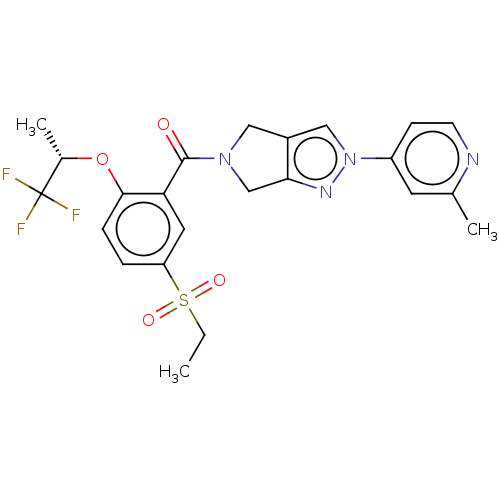
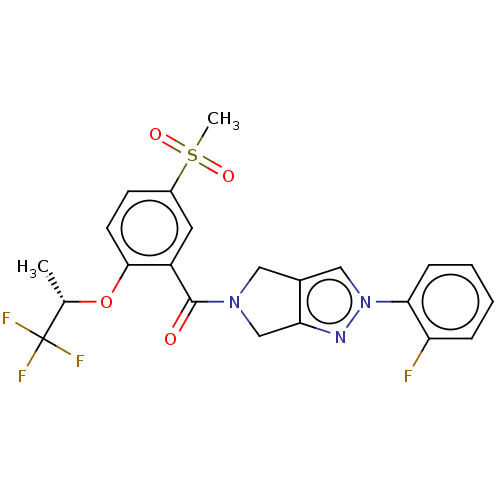
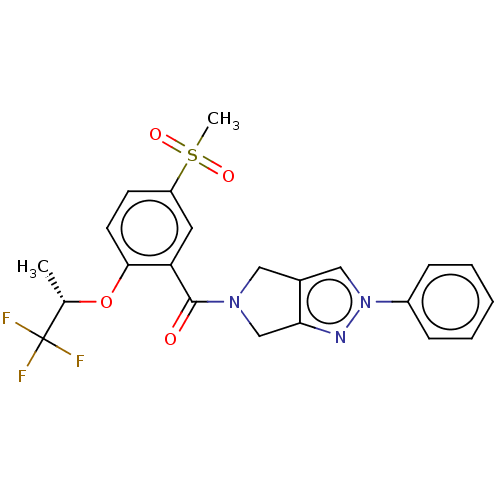
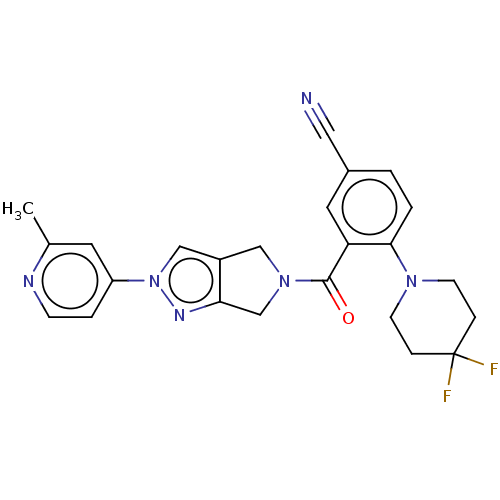
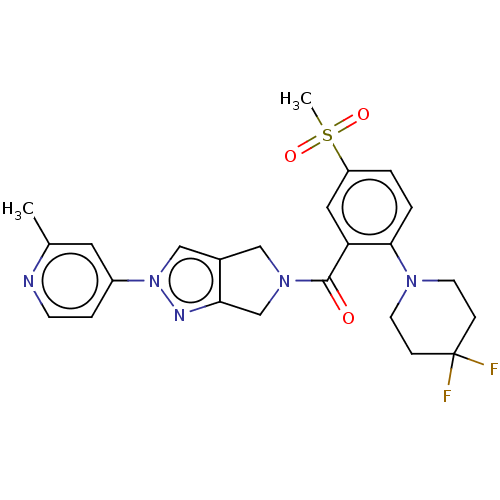
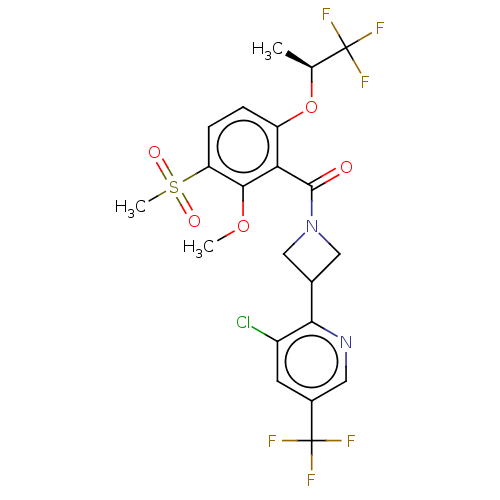
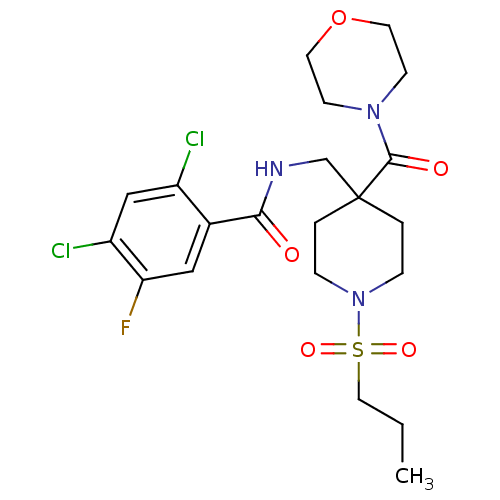
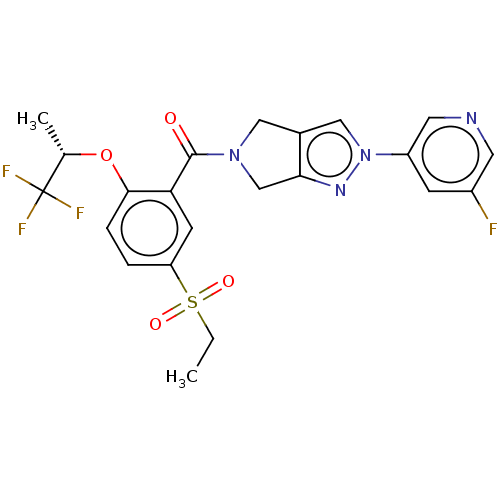
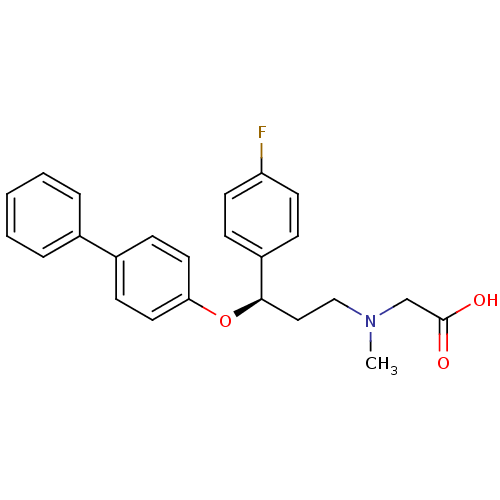
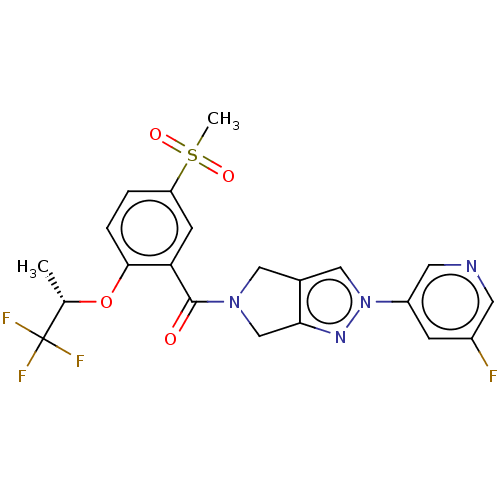
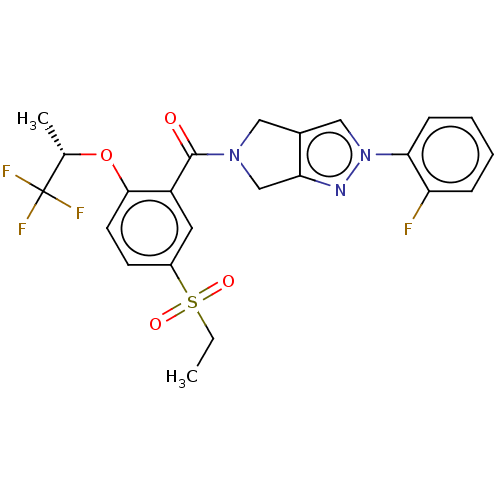
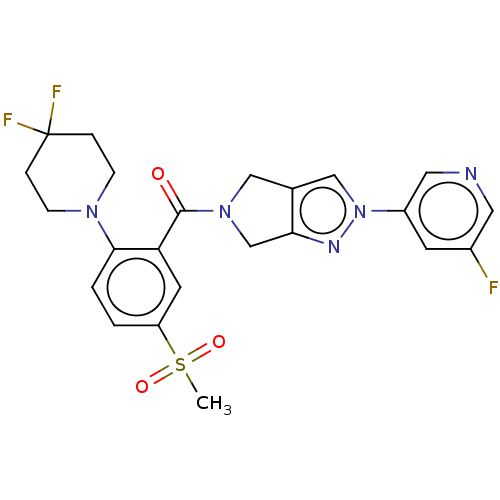
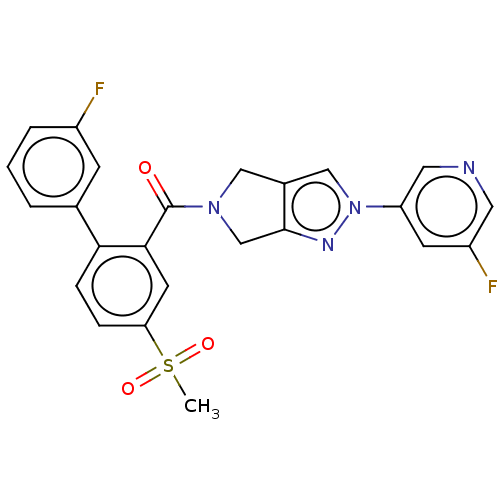
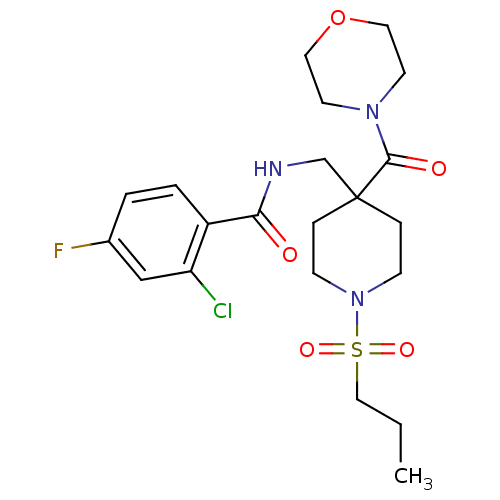
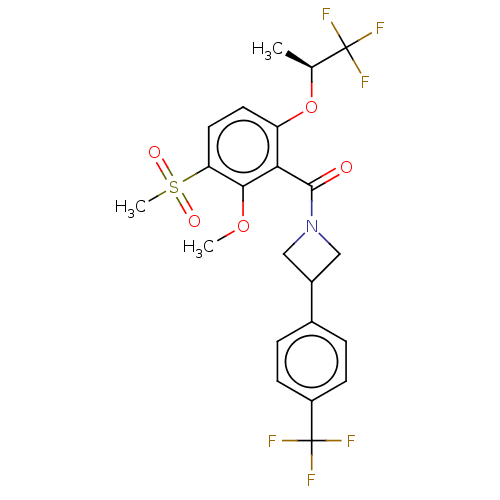
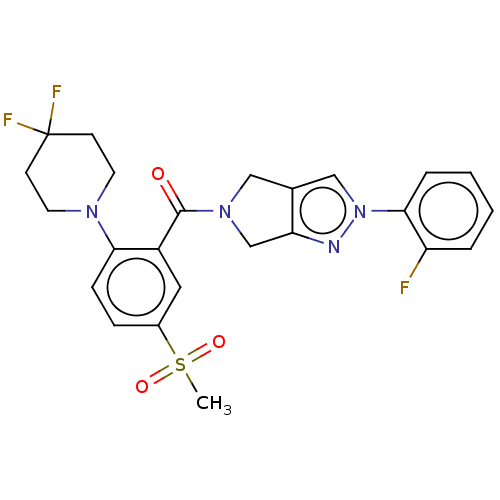
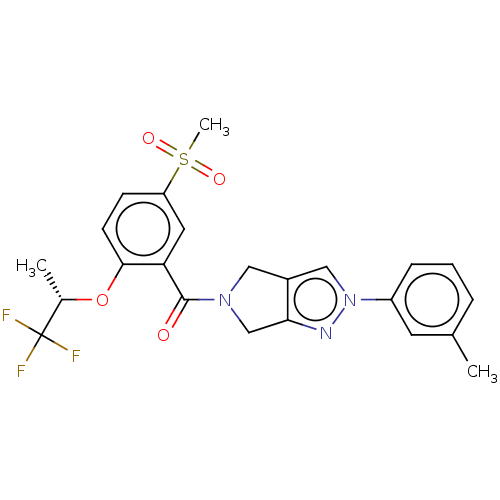
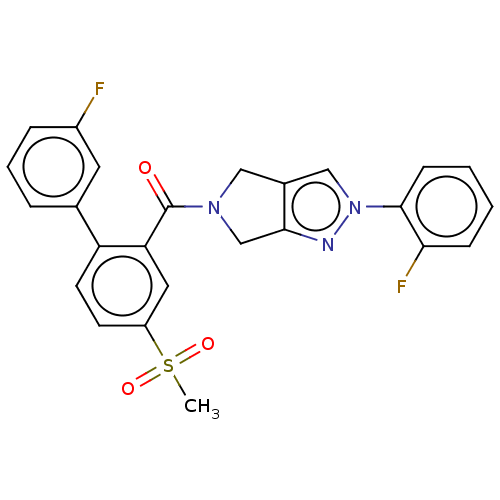
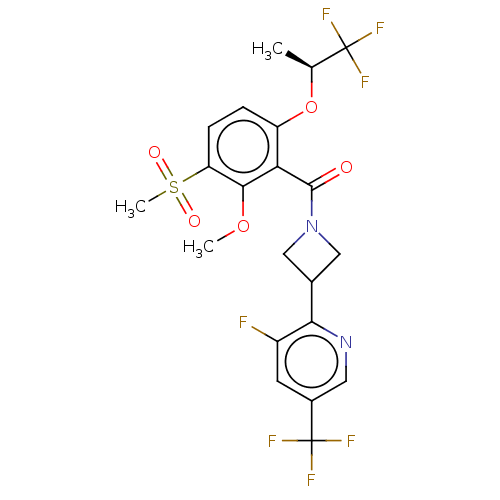
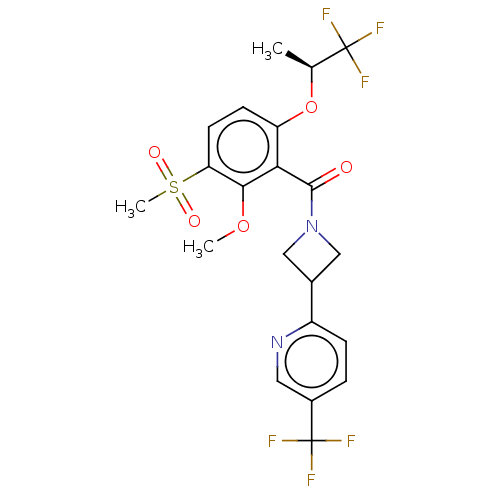
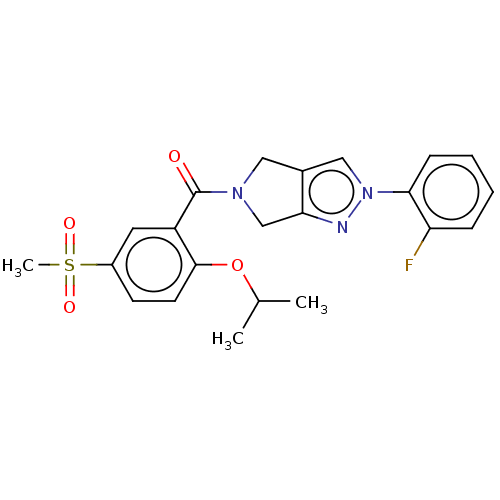
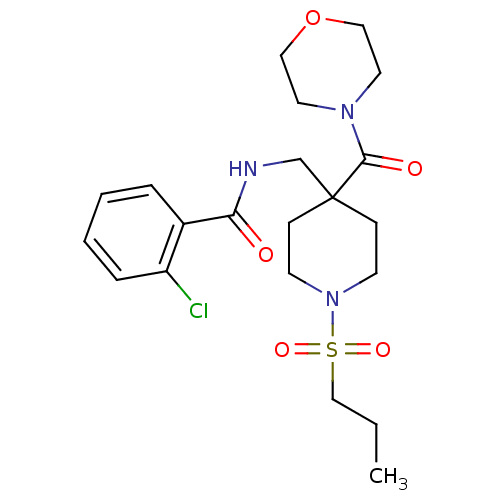
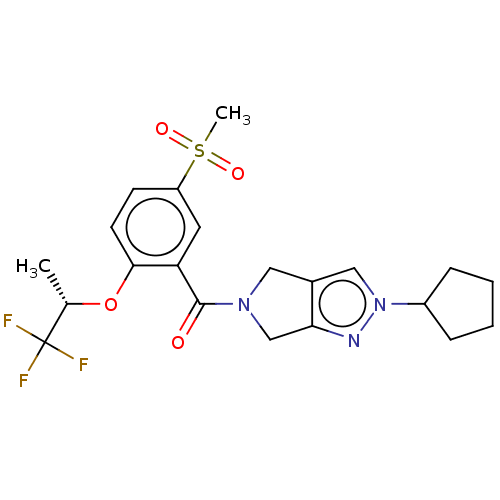
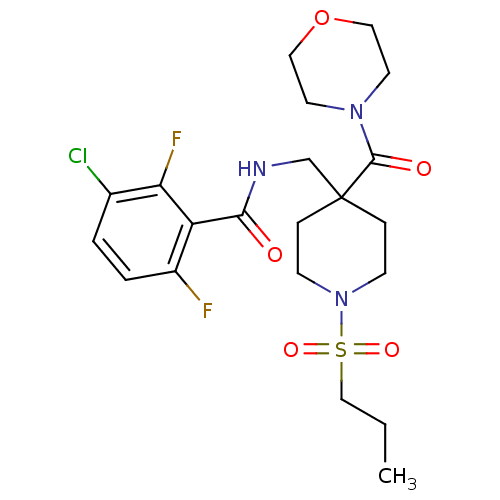
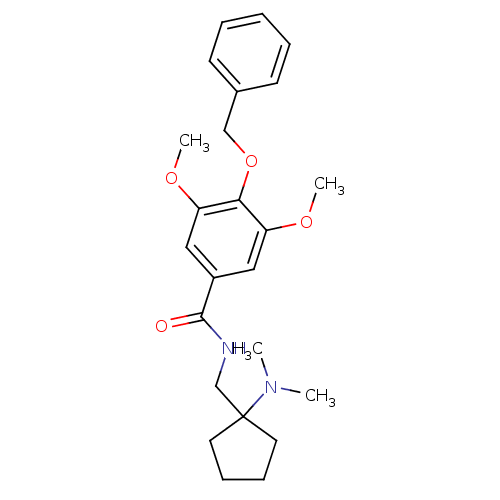
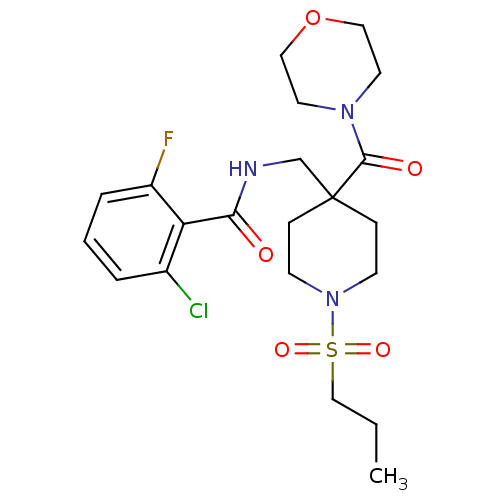
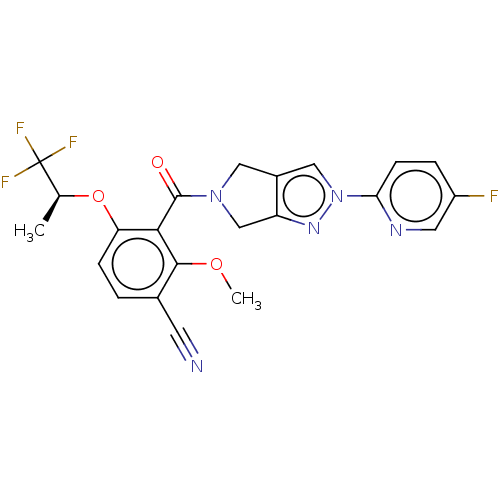
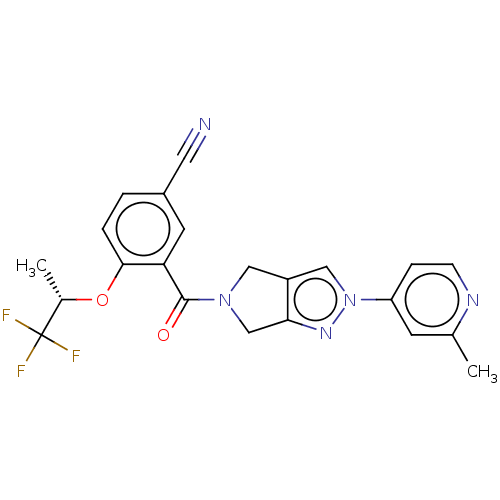
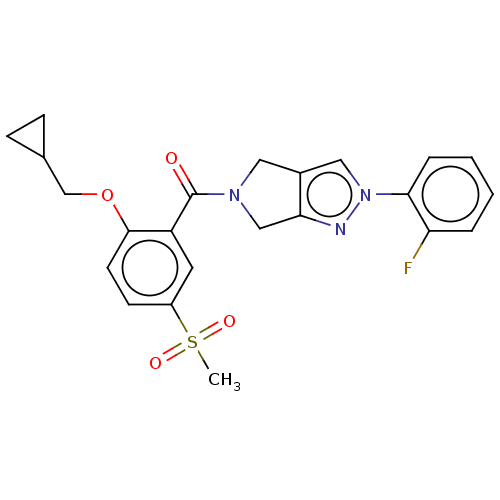
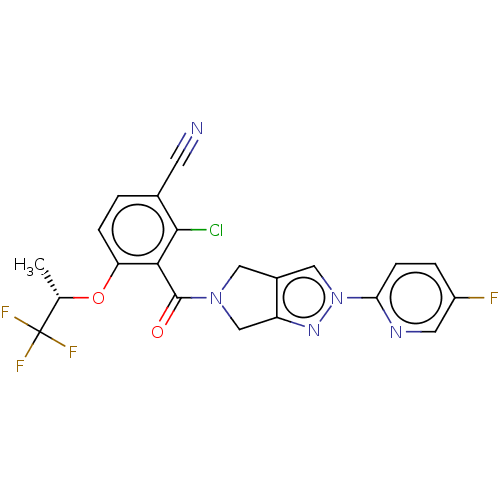
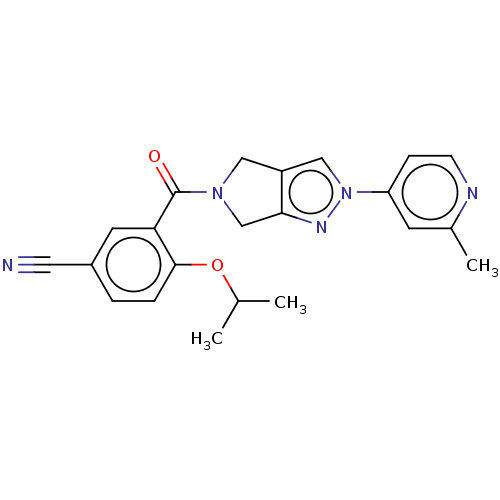
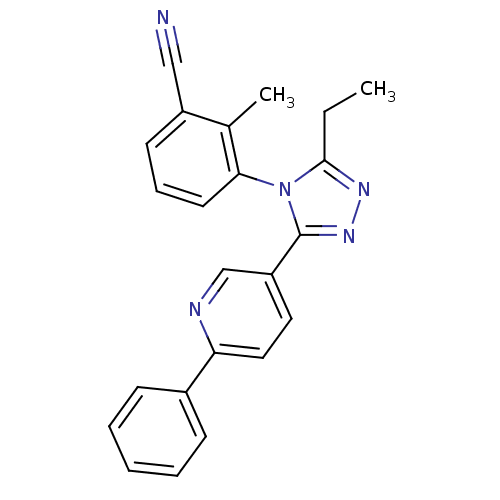
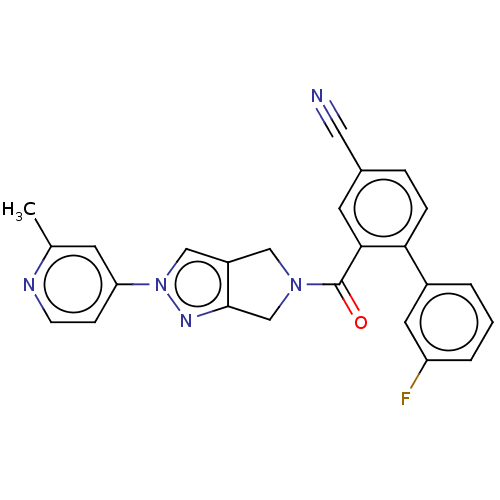
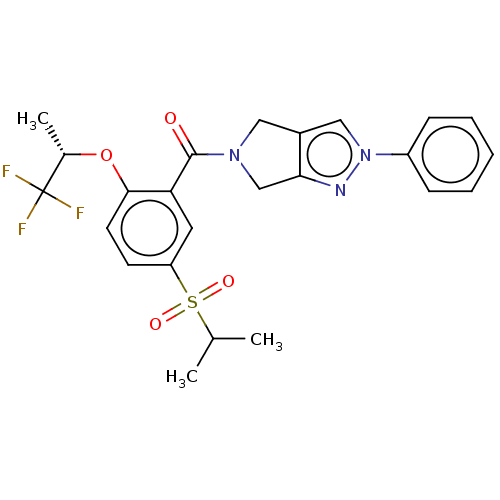
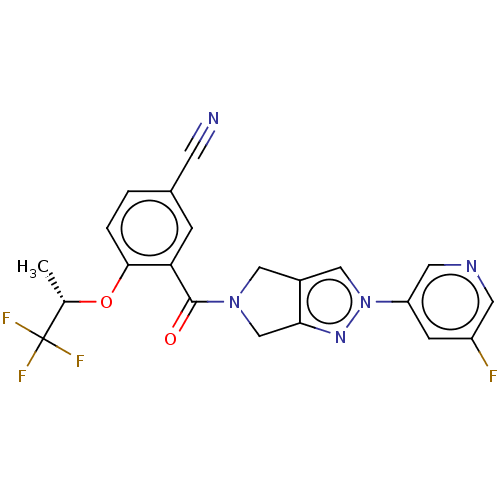
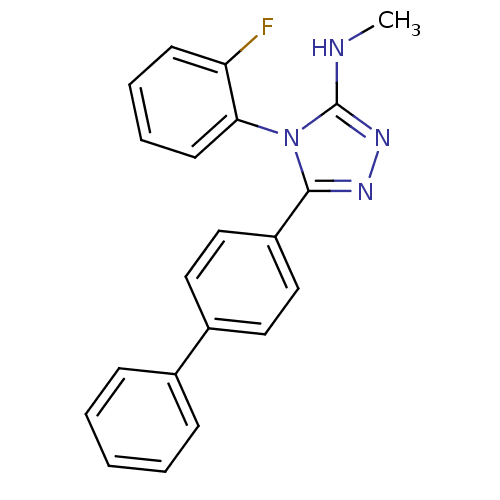
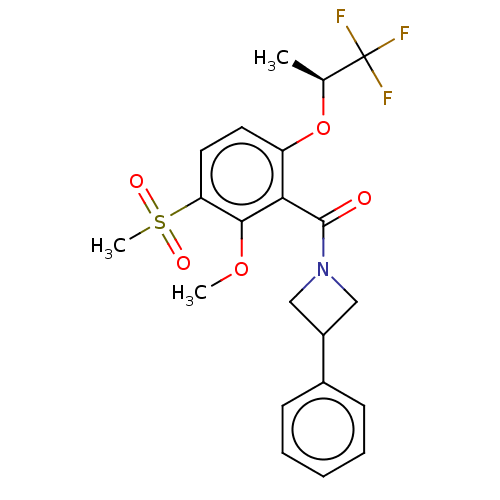
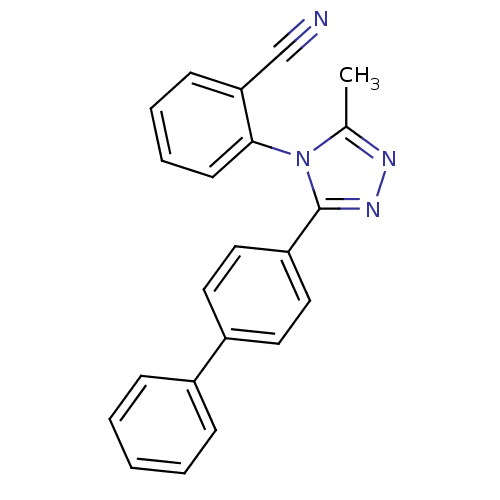
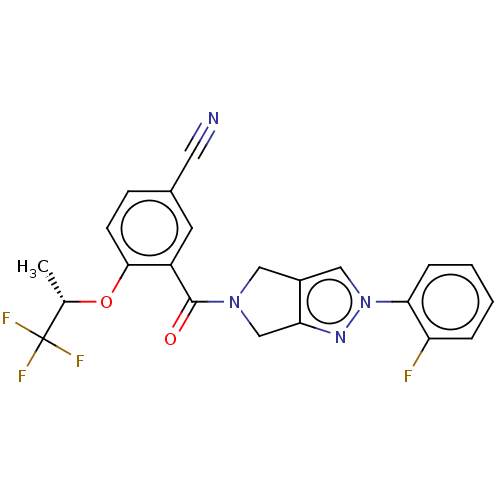
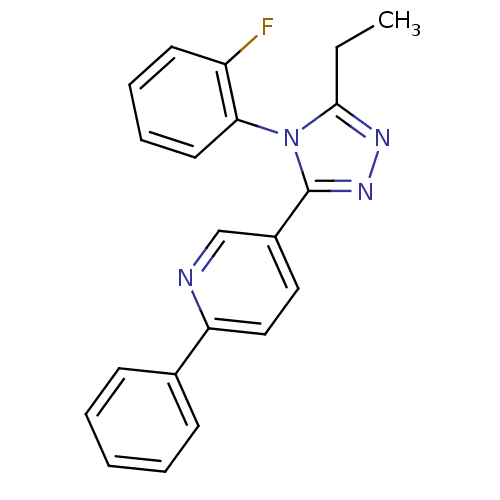
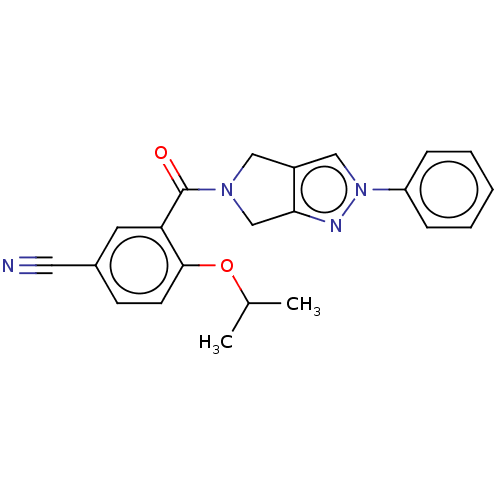
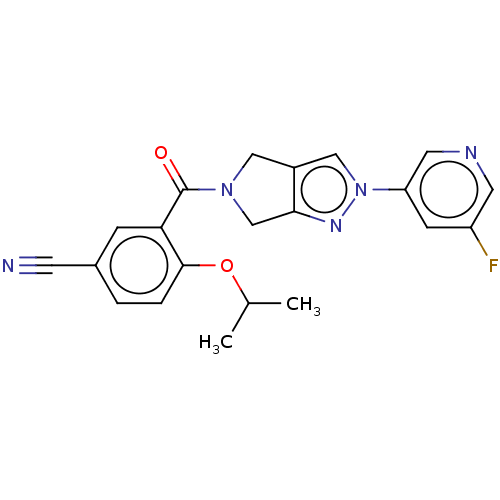
Affinity DataKd: 1.60nMAssay Description:Binding affinity to GlyT1 in rat cortical brain homogenate after 60 mins by gamma countingMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:Inhibition of rat GlyT1More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.70nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 7.40nMAssay Description:Inhibition of rat GlyT1More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 8.90nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of rat GlyT1More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of rat GlyT1More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of rat GlyT1More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of rat GlyT1More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 78nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 79nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 82nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMAssay Description:Inhibition of [3H]glycine uptake at GlyT1 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 96nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of [3H]glycine uptake at GlyT1 in rat C6 cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 108nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair
Affinity DataIC50: 161nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 172nMAssay Description:Inhibition of GlyT1 in rat primary cortical neurons assessed as reduction in radiolabeled glycine uptake by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 183nMAssay Description:Inhibition of GlyT1 in primary rat cortical neurons assessed as reduction in [14C]glycine uptake after 2 hrs by microbeta scintillation counting meth...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:Displacement of [3H]glycine from GlyT1 in rat C6 glioma cells incubated for 30 mins prior to substrate addition measured after 10 mins by scintillati...More data for this Ligand-Target Pair