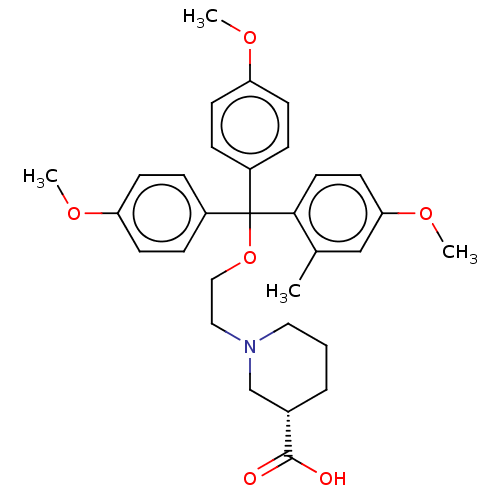
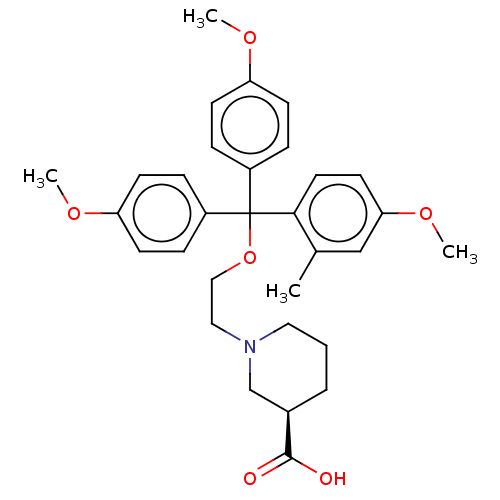
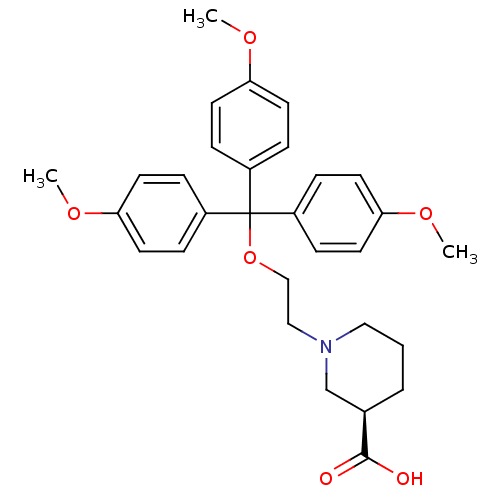
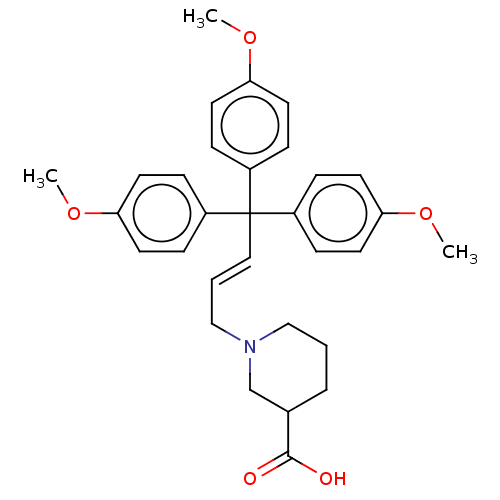
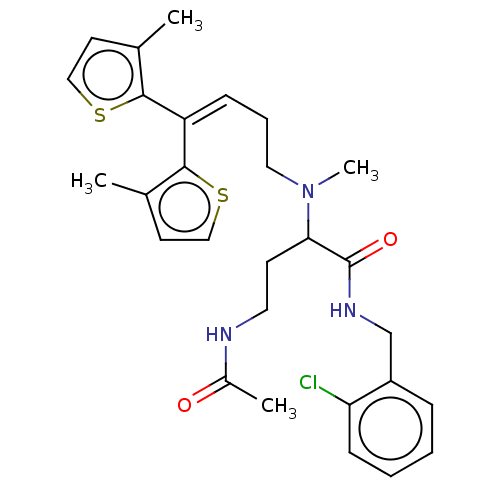
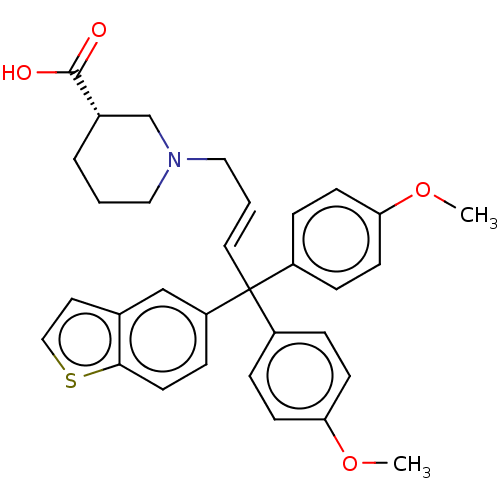
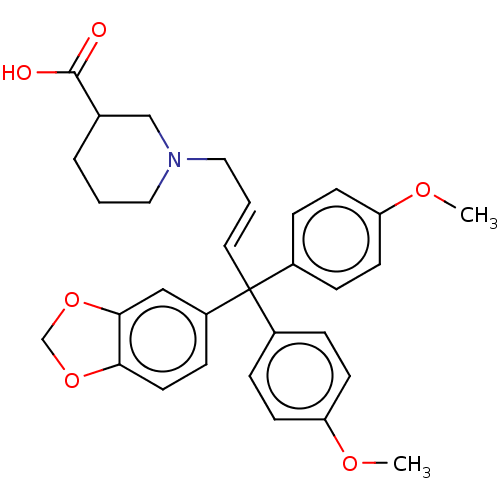
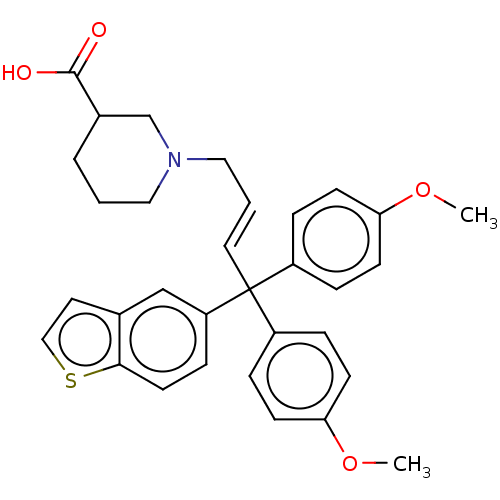
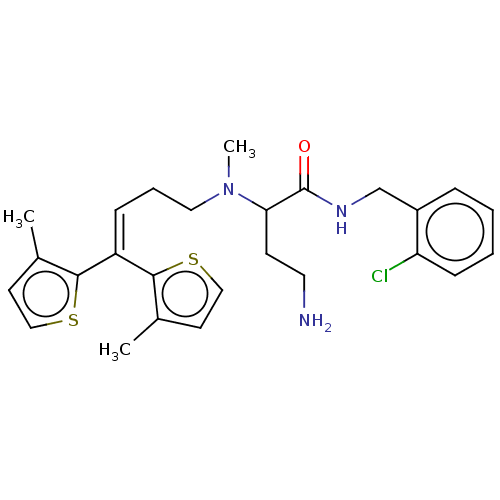
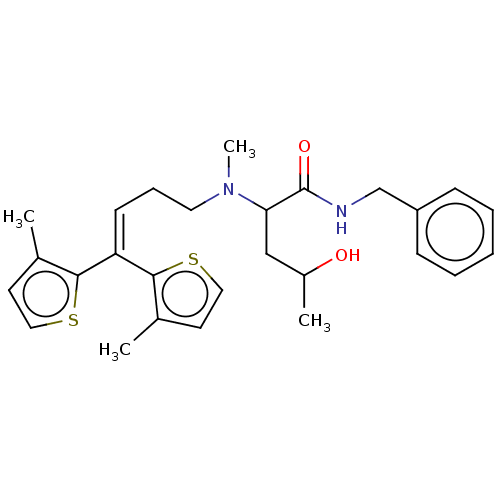
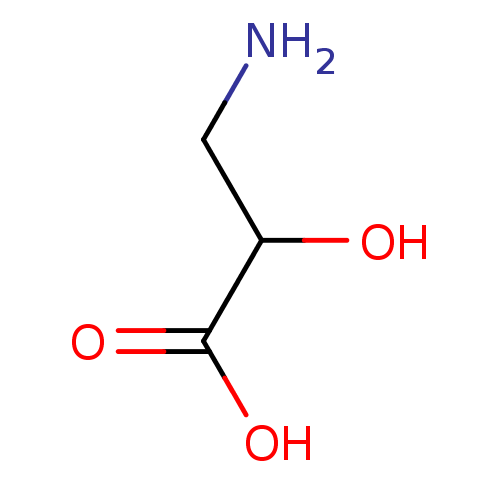
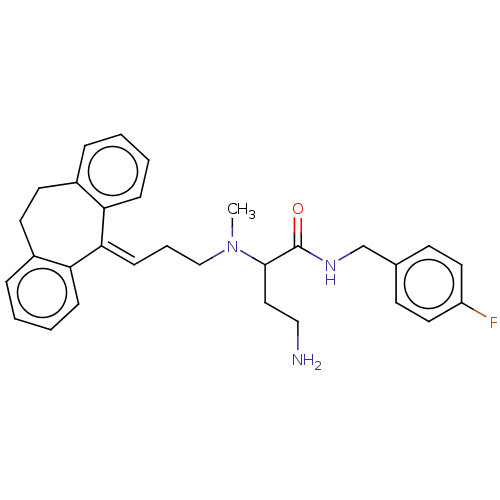
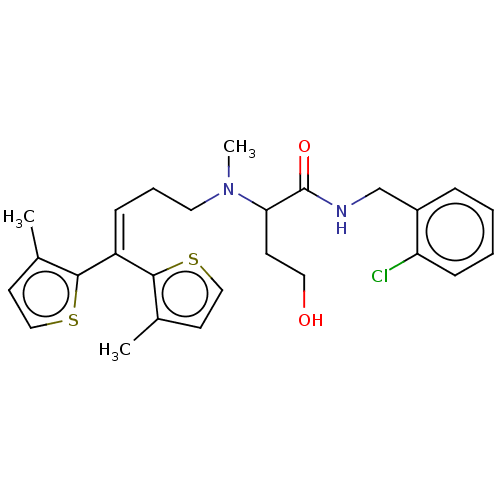
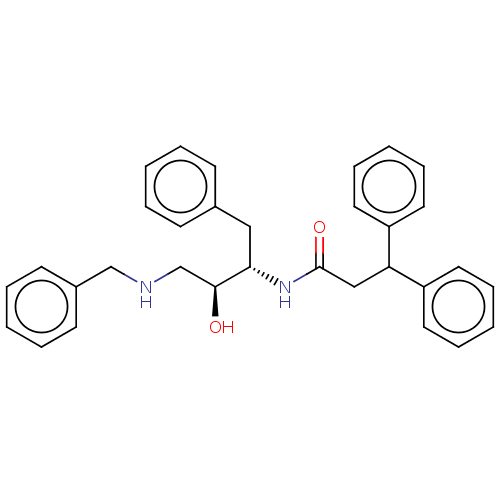
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.29E+3nMAssay Description:Inhibition of mouse GAT4 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.35E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.35E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.35E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as decrease in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition mea...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.35E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cell line assessed as reduction in [3H]GABA uptake incubated for 10 minsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.55E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.55E+3nMAssay Description:Inhibition of mouse GAT4 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.66E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.66E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cell line assessed as reduction in [3H]GABA uptake incubated for 10 minsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.66E+3nMAssay Description:Inhibition of mouse GTA4 receptorMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.66E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as decrease in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition mea...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.95E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.95E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as decrease in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition mea...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.95E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as decrease in [3H]GABA uptakeMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.95E+3nMAssay Description:Inhibition of mouse GTA4 receptorMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 1.95E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as decrease in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition mea...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 2.24E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK cells assessed as reduction in [3H]GABA uptake by measuring remaining [3H]GABA uptake levels preincubated f...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 2.46E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 2.46E+3nMAssay Description:Inhibition of mouse GAT3 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 2.82E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 2.88E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.16E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.39E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.39E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.39E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cells assessed as decrease in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition mea...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.39E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cell line assessed as reduction in [3H]GABA uptake incubated for 10 minsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.55E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.63E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.63E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.63E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.89E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.89E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 3.98E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 4.37E+3nMAssay Description:Inhibition of mouse GAT3 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 4.37E+3nMAssay Description:Inhibition of mouse GAT4 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 4.68E+3nMAssay Description:Inhibition of mouse GAT4 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 4.79E+3nMAssay Description:Inhibition of mouse GAT3 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 4.79E+3nMAssay Description:Inhibition of mouse GAT3 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of mouse GAT4 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 5.01E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cell line assessed as reduction in [3H]GABA uptake incubated for 10 minsMore data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 5.01E+3nMAssay Description:Inhibition of mouse GAT4 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 5.13E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cells assessed as decrease in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition mea...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 5.13E+3nMAssay Description:Inhibition of mouse GAT3 expressed in human HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 40 mins by LC-ESI-MS/MS analysi...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 5.13E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cells assessed as reduction in [3H]GABA uptake preincubated for 25 mins followed by [3H]GABA addition an...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 5.13E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK293 cell line assessed as inhibition of [3H]GABA uptake measured after 35 mins by liquid scintillation metho...More data for this Ligand-Target Pair
TargetSodium- and chloride-dependent GABA transporter 3(Mouse)
Jagiellonian University Medical College
Curated by ChEMBL
Jagiellonian University Medical College
Curated by ChEMBL
Affinity DataIC50: 5.13E+3nMAssay Description:Inhibition of mouse GAT3 expressed in HEK cells assessed as reduction in [3H]GABA uptake by measuring remaining [3H]GABA uptake levels preincubated f...More data for this Ligand-Target Pair
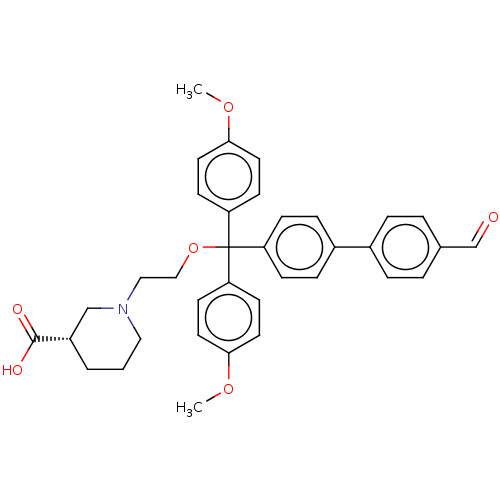
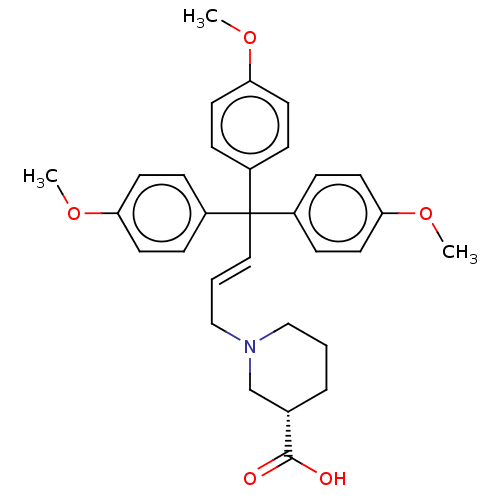
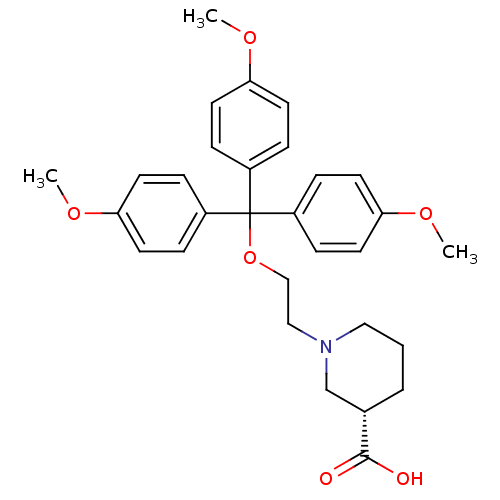
 3D Structure (crystal)
3D Structure (crystal)