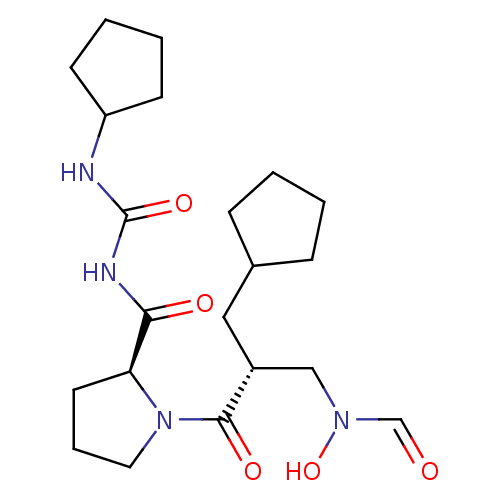
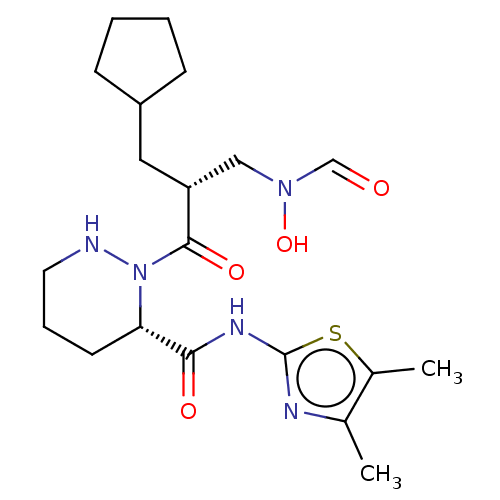
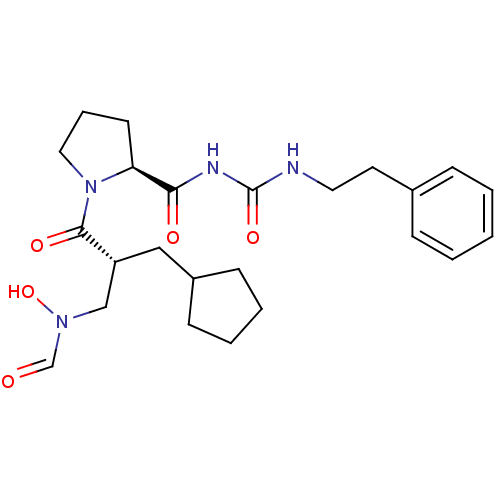
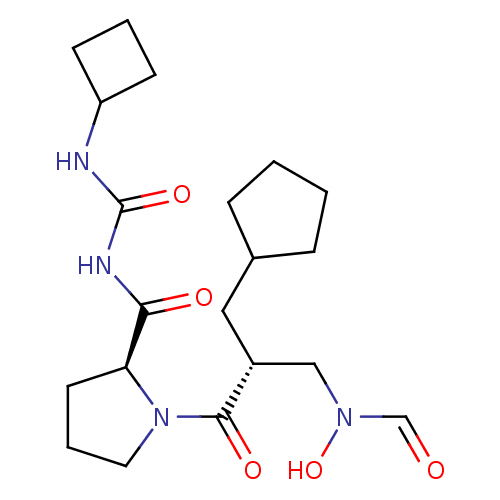
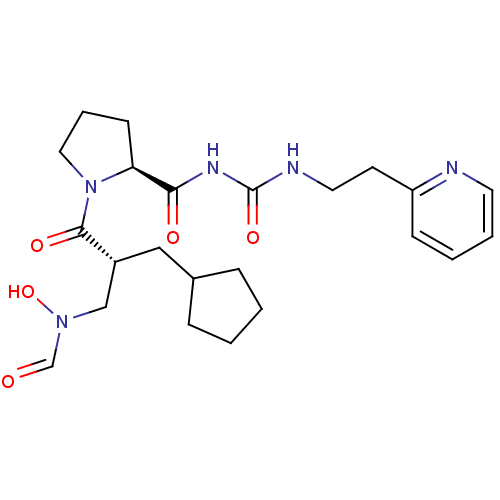
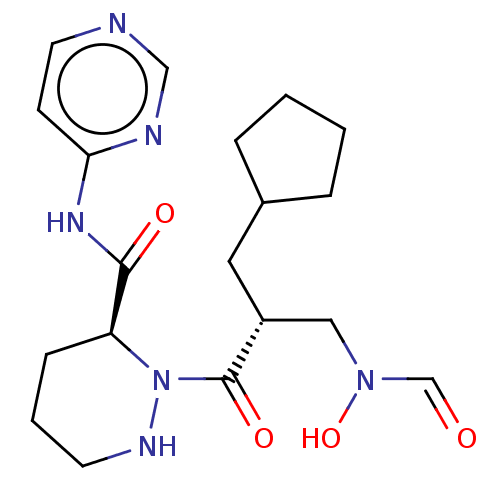
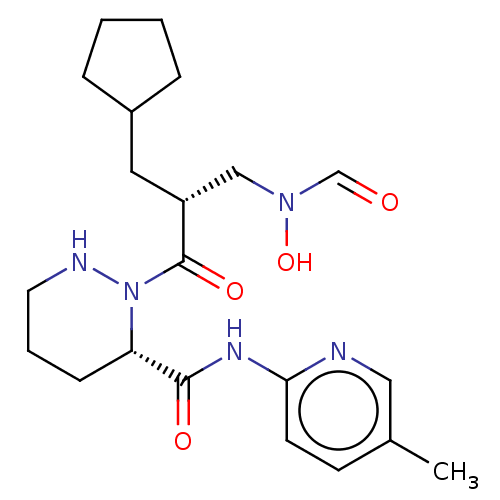
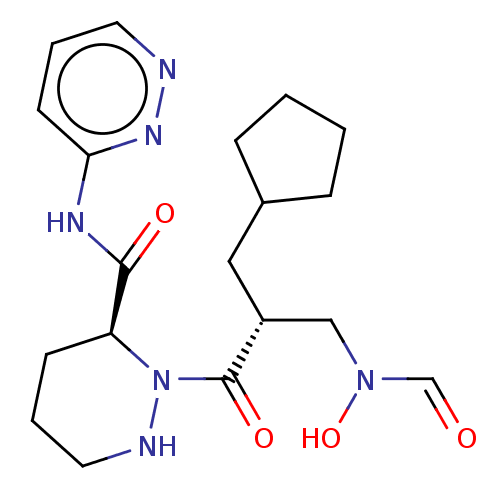
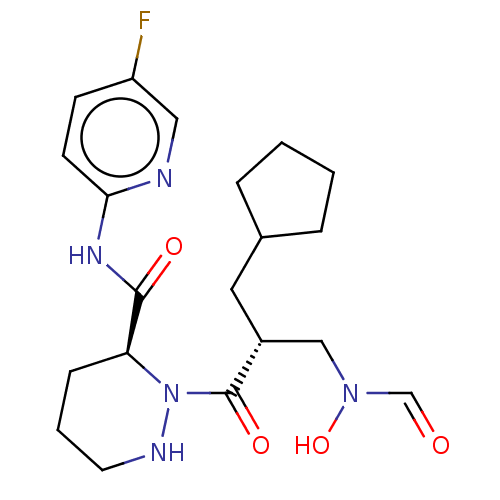
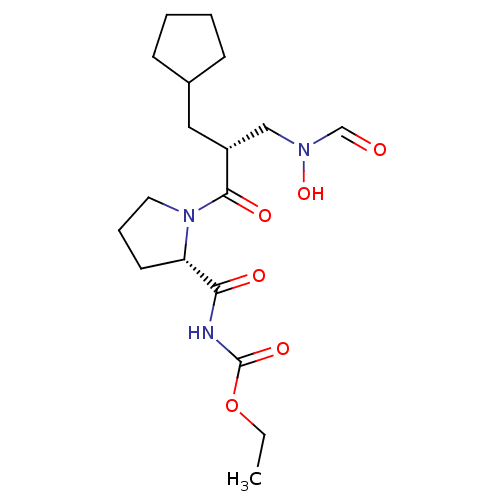
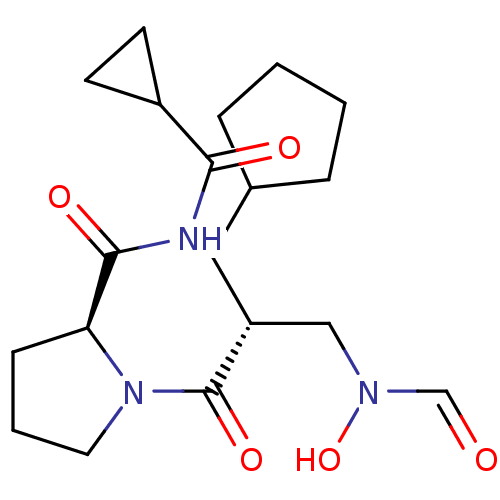
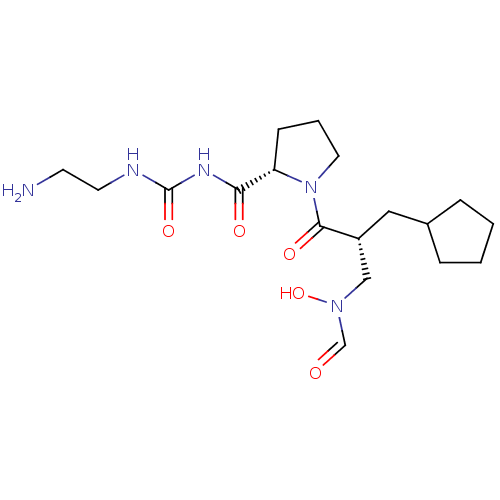
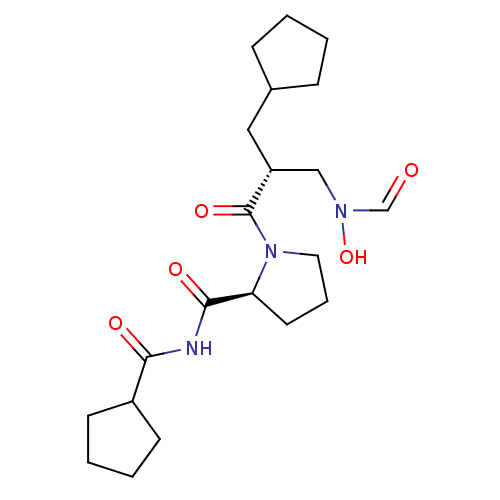
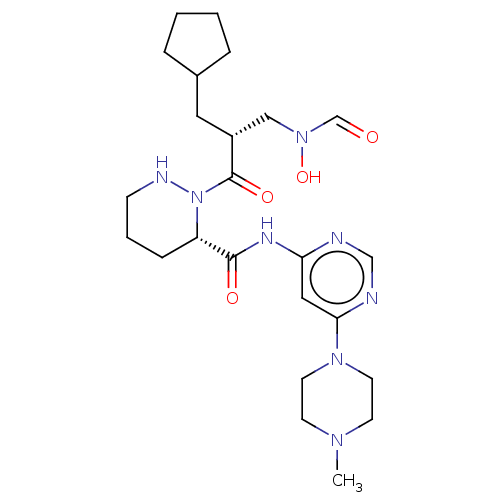
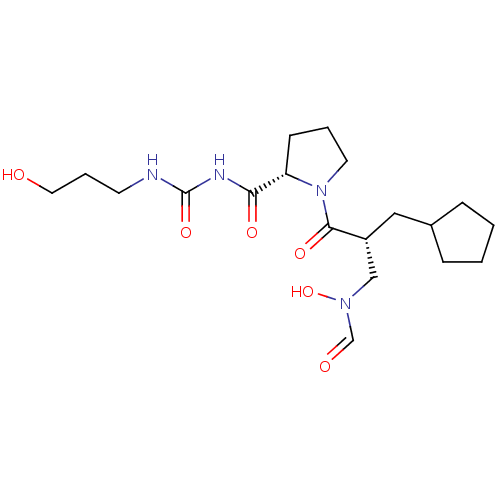
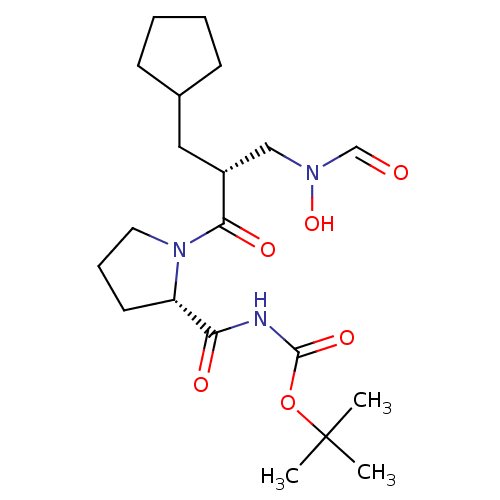
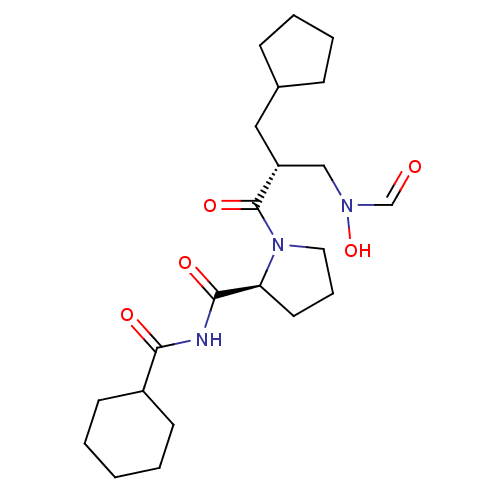
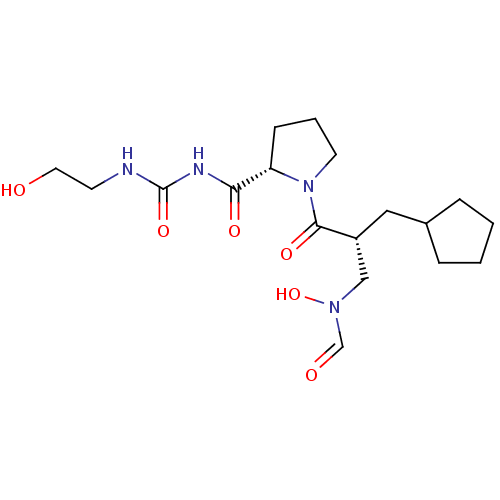
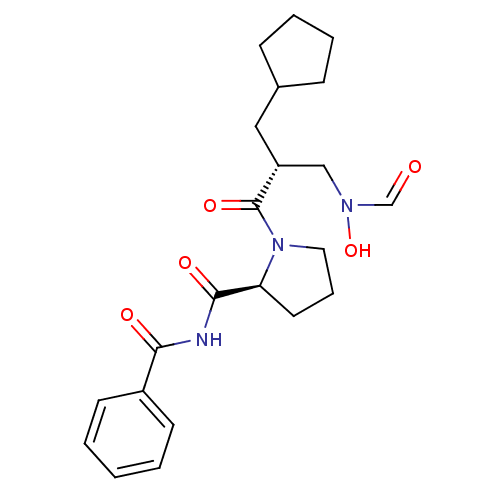
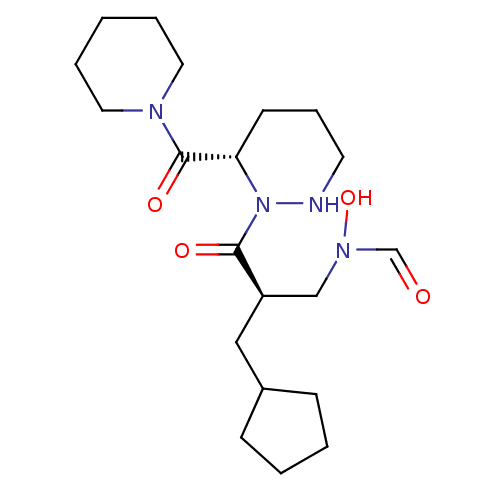
Affinity DataIC50: 0.520nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
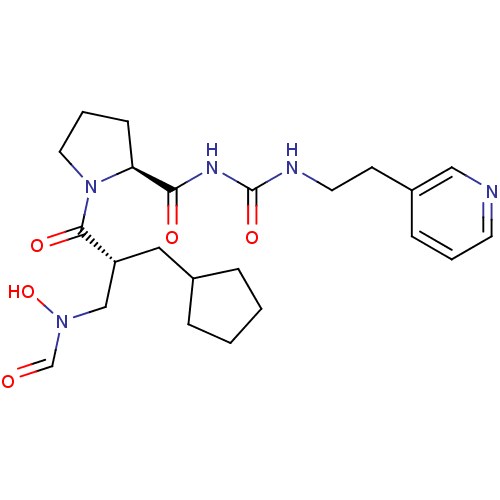
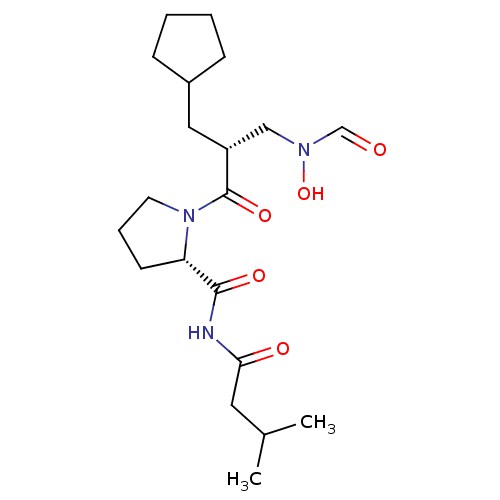
Affinity DataIC50: 0.630nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
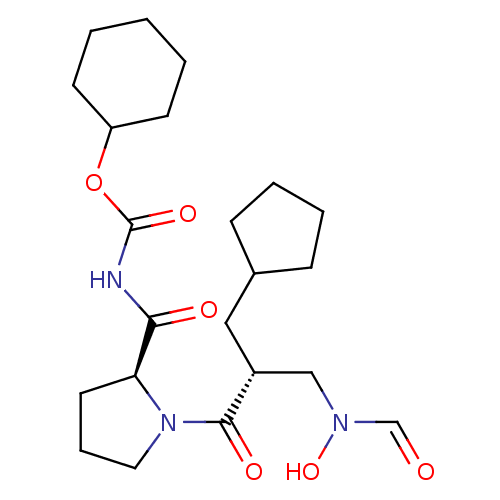
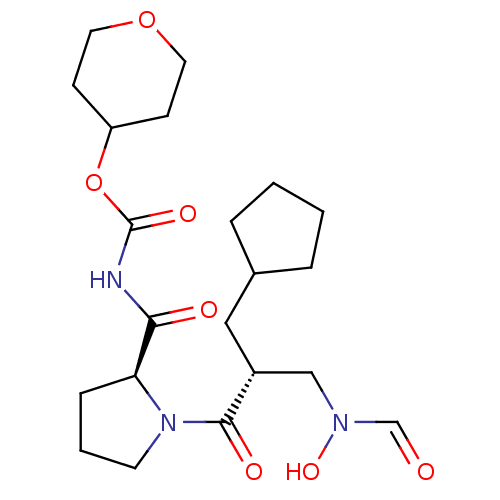
Affinity DataIC50: 0.860nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
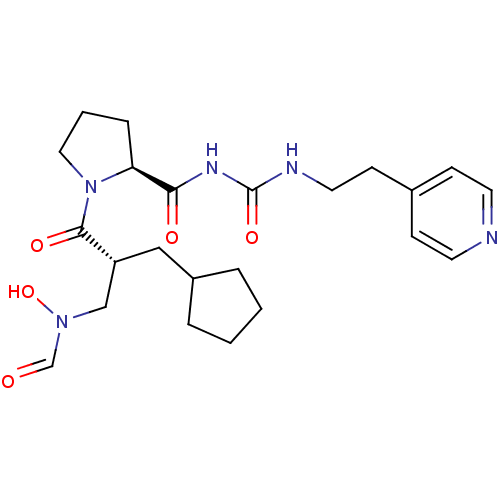
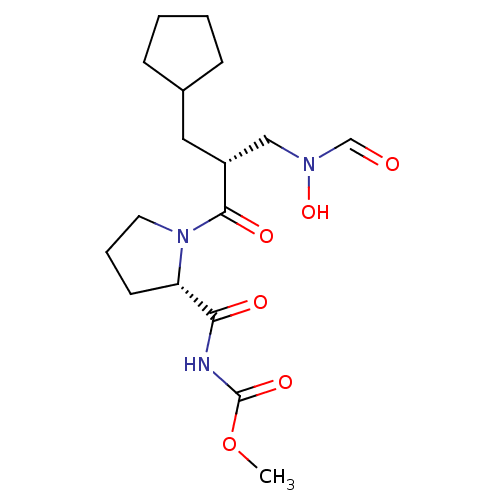
Affinity DataIC50: 0.900nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.930nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of Staphylococcus aureus PDF by fluorescence detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of Staphylococcus aureus PDF by fluorescence detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of Staphylococcus aureus PDF by fluorescence detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.40nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.5nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of Staphylococcus aureus PDF by fluorescence detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.70nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.80nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.40nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.5nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of Staphylococcus aureus PDF by fluorescence detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.20nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.90nMAssay Description:Inhibition of Staphylococcus aureus PDF assessed as formate release from fMAS peptide substrate after 20 mins by formate dehydrogenase coupled assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of Staphylococcus aureus PDF by absorbance detection assayMore data for this Ligand-Target Pair